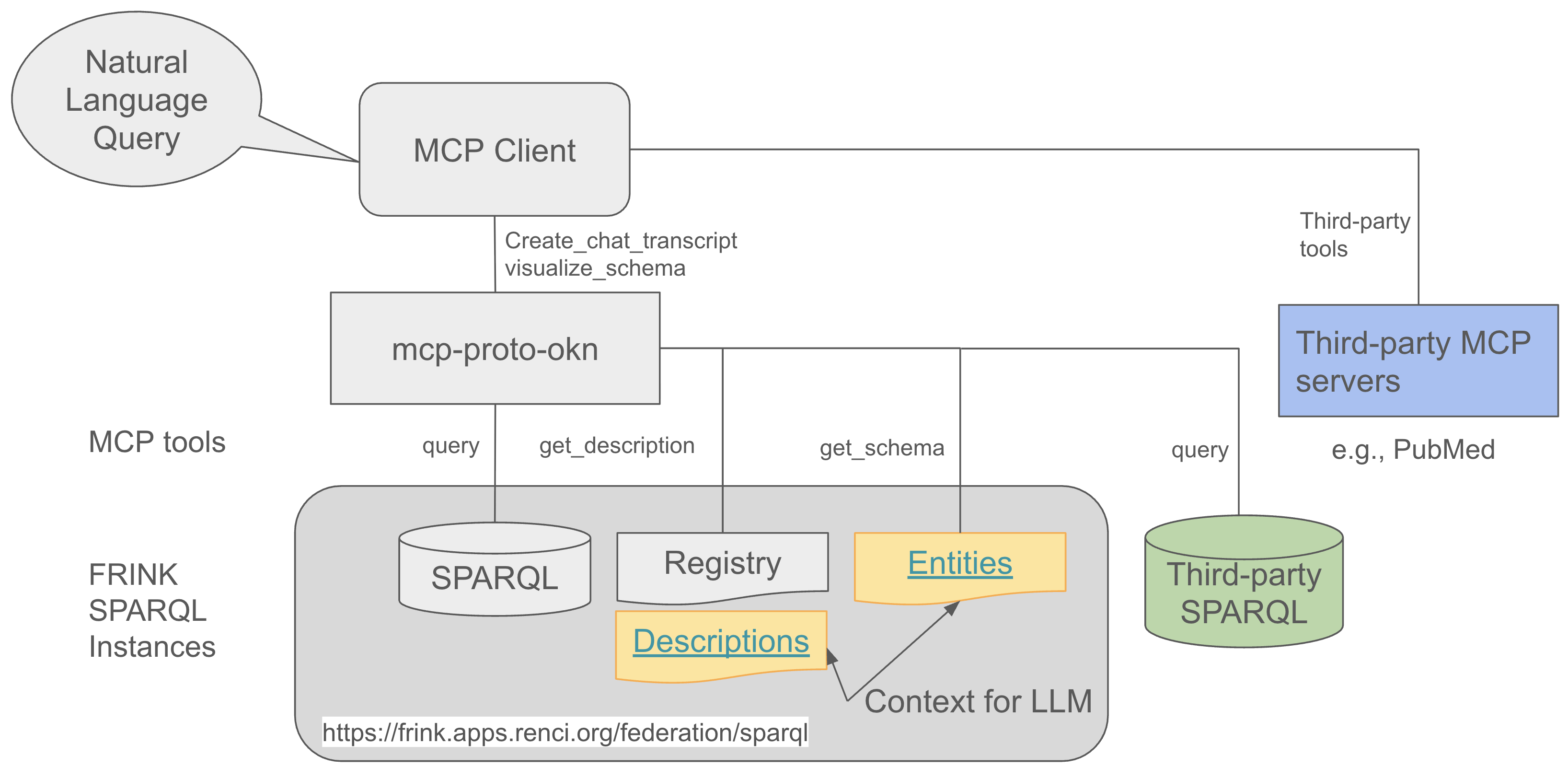
MCP server for querying FRINK SPARKQL endpoints, a project by the NSF Prototype Open Knowledge Network (Proto-OKN) program.
Project description
MCP Proto-OKN Server
A Model Context Protocol (MCP) server providing seamless access to SPARQL endpoints with specialized support for the NSF-funded Proto-OKN Project (Prototype Open Knowledge Network). This server enables querying the scientific knowledge graphs hosted on the FRINK platform. In addition, third-party SPARQL endpoints can be queried.
Features
- 🔗 FRINK Integration: Automatic detection and documentation linking for FRINK-hosted knowledge graphs
- 🧬 Proto-OKN Ecosystem: Optimized support for biomedical and scientific knowledge graphs, including:
- SPOKE - Scalable Precision Medicine Open Knowledge Engine
- BioBricks ICE - Chemical safety and cheminformatics data
- SAWGraph - Safe Agricultural Products and Water monitoring
- Additional Proto-OKN knowledge graphs - Expanding ecosystem of scientific data
- ⚙️ Flexible Configuration: Support for both FRINK and custom SPARQL endpoints
- 📚 Automatic Documentation: Registry links and metadata for Proto-OKN knowledge graphs
- 🔗 Federated Query: Prompts can query multiple endpoints
Architecture
The MCP Server Proto-OKN acts as a bridge between AI assistants (like Claude) and SPARQL knowledge graphs, enabling natural language queries to be converted into structured SPARQL queries and executed against scientific databases.
Prerequisites
Before installing the MCP Server Proto-OKN, ensure you have:
- Operating System: macOS, Linux, or Windows
- Client Application: One of the following:
- Claude Desktop with Pro or Max subscription
- VS Code Insiders with GitHub Copilot subscription
Installation
Prerequisites
The MCP Proto-OKN server requires the uv package manager to be installed on your system. If you don't have it installed run:
# macOS/Linux
curl -LsSf https://astral.sh/uv/install.sh | sh
# Windows
powershell -c "irm https://astral.sh/uv/install.ps1 | iex"
Note: Once
uvis installed, theuvxcommand in the configuration below will automatically download and run the latest version from PyPI when needed.
Claude Desktop Setup
Recommended for most users
-
Download and Install Claude Desktop
Visit https://claude.ai/download and install Claude Desktop for your operating system.
Requirements: Claude Pro or Max subscription is required for MCP server functionality.
-
Configure MCP Server
Option A: Download Pre-configured File (Recommended)
Download the pre-configured
claude_desktop_config.jsonfile with FRINK endpoints from the repository and copy it to the appropriate location:macOS:
# Download the config file curl -o /tmp/claude_desktop_config.json https://raw.githubusercontent.com/sbl-sdsc/mcp-proto-okn/main/config/claude_desktop_config.json # Copy to Claude Desktop configuration directory cp /tmp/claude_desktop_config.json "$HOME/Library/Application Support/Claude/"
Windows PowerShell:
# Download the config file Invoke-WebRequest -Uri "https://raw.githubusercontent.com/sbl-sdsc/mcp-proto-okn/main/config/claude_desktop_config.json" -OutFile "$env:TEMP\claude_desktop_config.json" # Copy to Claude Desktop configuration directory Copy-Item "$env:TEMP\claude_desktop_config.json" "$env:APPDATA\Claude\"
Option B: Manual Configuration
Alternatively, you can manually edit the configuration file in Claude Desktop. Navigate to
Claude->Settings->Developer->Edit Configto edit it.Below is an example of how to configure FRINK endpoints. Third-party SPARQL endpoints with a custom description can be added (see uniprot-sparql example below).
{ "mcpServers": { "spoke-sparql": { "command": "uvx", "args": [ "mcp-proto-okn", "--endpoint", "https://frink.apps.renci.org/spoke/sparql" ] }, "biobricks-sparql": { "command": "uvx", "args": [ "mcp-proto-okn", "--endpoint", "https://frink.apps.renci.org/biobricks-ice/sparql" ] }, "uniprot-sparql": { "command": "uvx", "args": [ "mcp-proto-okn", "--endpoint", "https://sparql.uniprot.org/sparql", "--description", "Resource for protein sequence and function information. For details: https://purl.uniprot.org/html/index-en.htm" ] } } }
Important: If you have existing MCP server configurations, do not use Option A as it will overwrite your existing configuration. Instead, use Option B and manually merge the Proto-OKN endpoints with your existing
mcpServersconfiguration. -
Restart Claude Desktop
After saving the configuration file, quit Claude Desktop completely and restart it. The application needs to restart to load the new configuration and start the MCP servers.
-
Verify Installation
- Launch Claude Desktop
- Navigate to
Claude->Settings->Connectors - Verify that the configured Proto-OKN endpoints appear in the connector list
- You can configure each service to always ask for permission or to run it unsupervised (recommended)
VS Code Setup
For advanced users and developers
-
Install VS Code Insiders
Download and install VS Code Insiders from https://code.visualstudio.com/insiders/
Note: VS Code Insiders is required as it includes the latest MCP (Model Context Protocol) features.
-
Install GitHub Copilot Extension
- Open VS Code Insiders
- Sign in with your GitHub account
- Install the GitHub Copilot extension
Requirements: GitHub Copilot subscription is required for MCP integration.
-
Configure Workspace
Create or edit
.vscode/mcp.jsonin your workspace:{ "servers": { "mcp-spoke-sparql": { "command": "uvx", "args": [ "mcp-proto-okn", "--endpoint", "https://frink.apps.renci.org/spoke/sparql" ] } } }
Note: For the VS Code Insiders configuration, change "mcpServers" to "servers".
-
Use the MCP Server
- Open a new chat window in VS Code
- Select Agent mode
- Choose Claude Sonnet 4 or later model for optimal performance
- The MCP servers will automatically connect and provide knowledge graph access
Quick Start
Once configured, you can immediately start querying knowledge graphs through natural language prompts in Claude Desktop or VS Code chat interface.
Example Queries
-
Knowledge Graph Overview
Provide a concise overview of the SPOKE knowledge graph, including its main purpose, data sources, and key features. -
Multi-Entity Analysis
Antibiotic contamination can contribute to antimicrobial resistance. Find locations with antibiotic contamination. -
Cross-Knowledge Graph Comparison
What type of data is available for perfluorooctanoic acid in SPOKE, BioBricks, and SAWGraph?
The AI assistant will automatically convert your natural language queries into appropriate SPARQL queries, execute them against the configured endpoints, and return structured, interpretable results.
Development
Installing from Source
If you want to run a development version:
# Clone the repository
git clone https://github.com/sbl-sdsc/mcp-proto-okn.git
cd mcp-proto-okn
# Install dependencies
uv sync
Building and Publishing
# Increment version number (patch, minor, major)
uv version --bump minor
# Build the package
uv build
# Publish to PyPI (maintainers only)
uv publish
API Reference
Available Tools
query
Executes SPARQL queries against the configured endpoint.
Parameters:
query_string(string, required): A valid SPARQL query
Returns:
- JSON object containing query results
get_description
Retrieves endpoint metadata and documentation.
Parameters:
- None
Returns:
- String containing endpoint description, PI information, funding details, and related documentation links
Command Line Interface
Required Parameters:
--endpoint: SPARQL endpoint URL (e.g.,https://frink.apps.renci.org/spoke/sparql)
Optional Parameters:
--description: Custom description for the SPARQL endpoint (auto-generated for FRINK endpoints)
Example Usage:
uvx mcp-proto-okn --endpoint https://frink.apps.renci.org/spoke/sparql
Troubleshooting
Common Issues
MCP server not appearing in Claude Desktop:
- Ensure you've completely quit and restarted Claude Desktop (not just closed the window)
- Check that your JSON configuration is valid (use a JSON validator)
- Verify that
uvxis installed and accessible in your PATH
Connection errors:
- Check your internet connection
- Verify the SPARQL endpoint URL is correct and accessible
- Some endpoints may have rate limits or temporary downtime
Performance issues:
- Complex SPARQL queries may take time to execute
- Consider breaking down complex queries into smaller parts
- Check the endpoint's documentation for query best practices
License
This project is licensed under the BSD 3-Clause License. See the LICENSE file for details.
Citation
If you use MCP Server Proto-OKN in your research, please cite the following works:
@software{rose2025mcp-proto-okn,
title={MCP Server Proto-OKN},
author={Rose, P.W. and Nelson, C.A. and Shi, Y. and Baranzini, S.E.},
year={2025},
url={https://github.com/sbl-sdsc/mcp-proto-okn}
}
@software{rose2025spoke-genelab,
title={NASA SPOKE-GeneLab Knowledge Graph},
author={Rose, P.W. and Nelson, C.A. and Gebre, S.G. and Soman, K. and Grigorev, K.A. and Sanders, L.M. and Costes, S.V. and Baranzini, S.E.},
year={2025},
url={https://github.com/BaranziniLab/spoke_genelab}
}
Related Publications
-
Nelson, C.A., Rose, P.W., Soman, K., Sanders, L.M., Gebre, S.G., Costes, S.V., Baranzini, S.E. (2025). "Nasa Genelab-Knowledge Graph Fabric Enables Deep Biomedical Analysis of Multi-Omics Datasets." NASA Technical Reports, 20250000723. Link
-
Sanders, L., Costes, S., Soman, K., Rose, P., Nelson, C., Sawyer, A., Gebre, S., Baranzini, S. (2024). "Biomedical Knowledge Graph Capability for Space Biology Knowledge Gain." 45th COSPAR Scientific Assembly, July 13-21, 2024. Link
Acknowledgments
Funding
This work is supported by:
- National Science Foundation Award #2333819: "Proto-OKN Theme 1: Connecting Biomedical information on Earth and in Space via the SPOKE knowledge graph"
Related Projects
- Proto-OKN Project - Prototype Open Knowledge Network initiative
- FRINK Platform - Knowledge graph hosting infrastructure
- Knowledge Graph Registry - Catalog of available knowledge graphs
- Model Context Protocol - AI assistant integration standard
- Original MCP Server SPARQL - Base implementation reference
For questions, issues, or contributions, please visit our GitHub repository.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file mcp_proto_okn-0.2.0.tar.gz.
File metadata
- Download URL: mcp_proto_okn-0.2.0.tar.gz
- Upload date:
- Size: 13.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.7.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e4a6438c4e68b7f4e44317517a35309ab7f89605a91db29b7fcd128fe0ad8fa2
|
|
| MD5 |
9dd2092dd501eb5de7eb6ce4f05f6e2e
|
|
| BLAKE2b-256 |
1b3f501dd988e3f36e18f9fec8a733680092d5c66a456df958cc4c324464c492
|
File details
Details for the file mcp_proto_okn-0.2.0-py3-none-any.whl.
File metadata
- Download URL: mcp_proto_okn-0.2.0-py3-none-any.whl
- Upload date:
- Size: 10.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.7.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ba983cc6bb8ada8af110ba4695f06073a9586a25de0c1348bd0566beb384236c
|
|
| MD5 |
340ecbf82dd2e6df377ba9a350f57ca5
|
|
| BLAKE2b-256 |
5aedee8e39f280b0fd708e8849a0d3a7be6e2c009e586c9008aa99c5920103fd
|