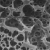
Shape-constrained molecule generation via Equivariant Diffusion and GCN
Project description
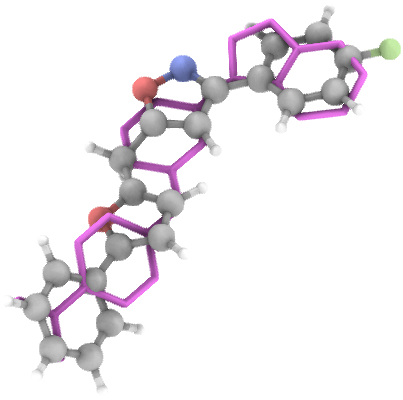
ML Conformer Generator
ML Conformer Generator is a tool for shape-constrained molecule generation with an Equivariant Diffusion Model (EDM) and a Graph Convolutional Network (GCN). It is designed to generate 3D molecular conformations that are both chemically valid and spatially similar to a reference shape.
Supported features
-
Shape-guided molecular generation
Generate novel molecules that conform to arbitrary 3D shapes—such as protein binding pockets or custom-defined spatial regions.
-
Reference-based conformer similarity
Create molecules conformations of which closely resemble a reference structure, supporting scaffold-hopping and ligand-based design workflows.
-
Fragment-based inpainting
Fix specific substructures or fragments within a molecule and complete or grow the rest in a geometrically consistent manner.
-
Inertial Fragment Matching
Generate molecules fragment-by-fragment by leveraging the physical properties of the shape descriptor, improving both shape similarity and chemical validity.
Citation
If you use MLConfGen in your research, please cite:
Denis Sapegin, Fedor Bakharev, Dmitry Krupenya, Azamat Gafurov, Konstantin Pildish, and Joseph C. Bear.
Moment of inertia as a simple shape descriptor for diffusion-based shape-constrained molecular generation.
Digital Discovery, 2025.
DOI: 10.1039/D5DD00318K
Installation
-
Install the package for your preferred backend:
-
pip install mlconfgen[torch]— use the PyTorch-based inference pipeline -
pip install mlconfgen[onnx]— use the torch-free ONNX runtime version
-
-
Load the weights from Huggingface
edm_moi_chembl_15_39.pt
adj_mat_seer_chembl_15_39.pt
🐍 Python API
See interactive examples: ./python_api_demo.ipynb
from rdkit import Chem
from mlconfgen import MLConformerGenerator, evaluate_samples
model = MLConformerGenerator(
edm_weights="./edm_moi_chembl_15_39.pt",
adj_mat_seer_weights="./adj_mat_seer_chembl_15_39.pt",
diffusion_steps=100,
)
reference = Chem.MolFromMolFile('./assets/demo_files/ceyyag.mol')
samples = model.generate_conformers(reference_conformer=reference, n_samples=20, variance=2)
aligned_reference, std_samples = evaluate_samples(reference, samples)
🚀 Overview
This solution employs:
- Equivariant Diffusion Model (EDM) [1]: For generating atom coordinates and types under a shape constraint.
- Graph Convolutional Network (GCN) [2]: For predicting atom adjacency matrices.
- Deterministic Standardization Pipeline: For refining and validating generated molecules.
🧠 Model Training
- Trained on 1.6 million compounds from the ChEMBL database.
- Filtered to molecules with 15–39 heavy atoms.
- Supported elements:
H, C, N, O, F, P, S, Cl, Br.
🧪 Standardization Pipeline
The generated molecules are post-processed through the following steps:
- Largest Fragment picker
- Valence check
- Kekulization
- RDKit sanitization
- Constrained Geometry optimization via MMFF94 Molecular Dynamics
📏 Evaluation Pipeline
Aligns and Evaluates shape similarity between generated molecules and a reference using Shape Tanimoto Similarity [3] via Gaussian Molecular Volume overlap.
Hydrogens are ignored in both reference and generated samples for this metric.
📊 Performance (100 Denoising Steps)
Tested on 100,000 samples using 1,000 CCDC Virtual Screening [4] reference compounds.
General Overview
- ⏱ Avg time to generate 50 valid samples: 11.46 sec (NVIDIA H100) (100 samples batch)
- ⚡️ Generation speed: 4.18 valid molecules/sec (100 samples batch)
- 💾 GPU memory (per generation thread): Up to 14.0 GB (
float1639 atoms 100 samples) - 📐 Avg Shape Tanimoto Similarity: 53.32% (Basic generation) - 69.97% (Inertial Fragment Matching)
- 🎯 Max Shape Tanimoto Similarity: 99.69%
- 🔬 Avg Chemical Tanimoto Similarity (2-hop 2048-bit Morgan Fingerprints): 10.87%
- 🧬 % Chemically novel (vs. training set): 99.84%
- ✔️ % Valid molecules (post-standardization): 48% (ML Bond Prediction) - 93% (OpenBabel bond prediction)
- 🔁 % Unique molecules in generated set: 99.94%
- 📎 Fréchet Fingerprint Distance (2-hop 2048-bit Morgan Fingerprints):
- To ChEMBL: 4.13
- To PubChem: 2.64
- To ZINC (250k): 4.95
PoseBusters [5] validity check results:
Overall stats:
- PB-valid molecules: 91.33 %
Detailed Problems:
- position: 0.01 %
- mol_pred_loaded: 0.0 %
- sanitization: 0.01 %
- inchi_convertible: 0.01 %
- all_atoms_connected: 0.0 %
- bond_lengths: 0.24 %
- bond_angles: 0.70 %
- internal_steric_clash: 2.31 %
- aromatic_ring_flatness: 3.34 %
- non-aromatic_ring_non-flatness: 0.27 %
Synthesizability of the generated compounds
SA Score [6]
1 (easy to make) - 10 (very difficult to make)
Average SA Score: 3.18
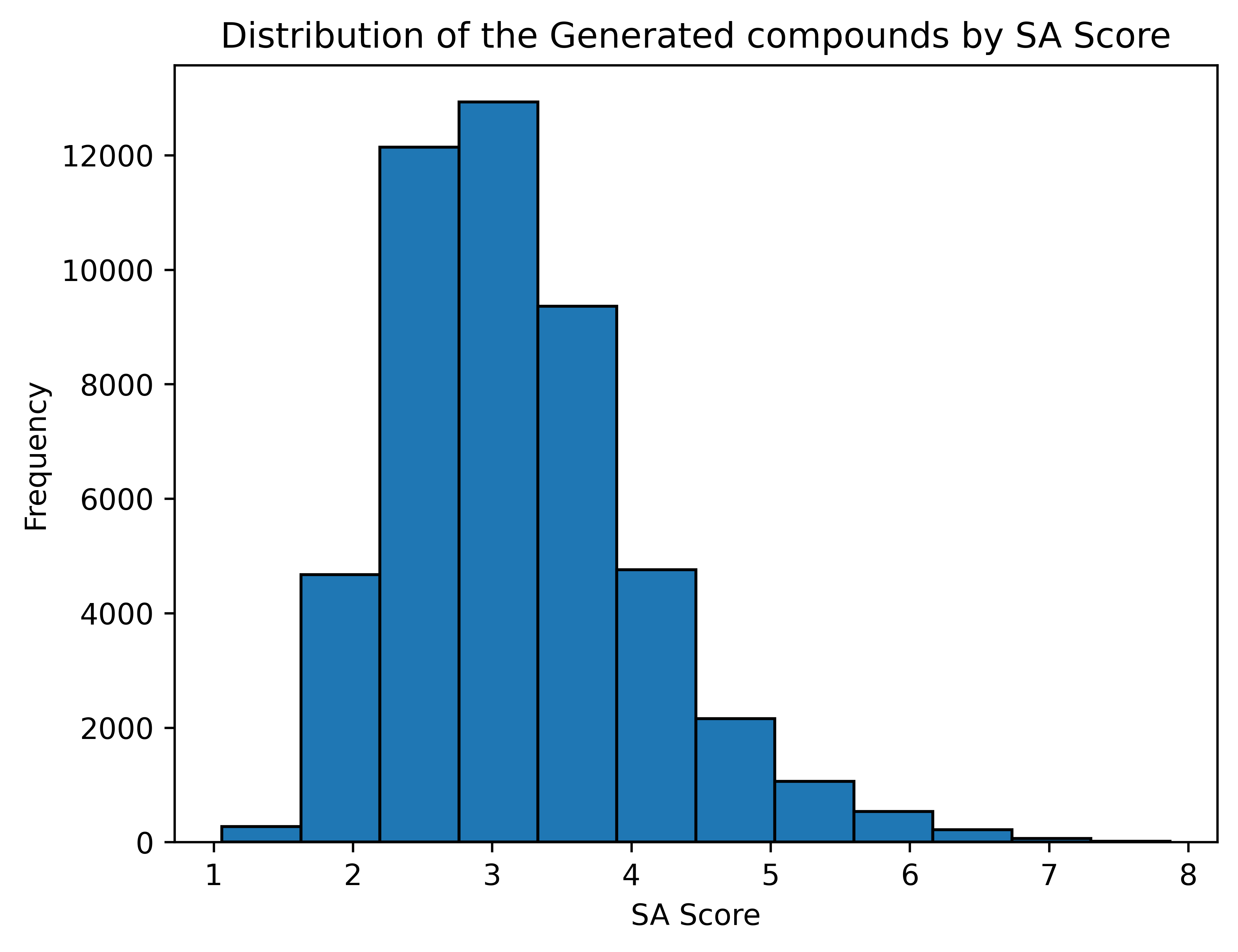
Generation Examples
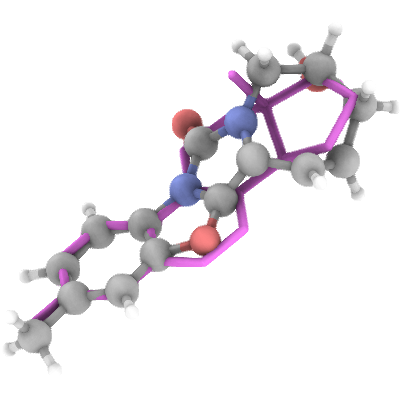
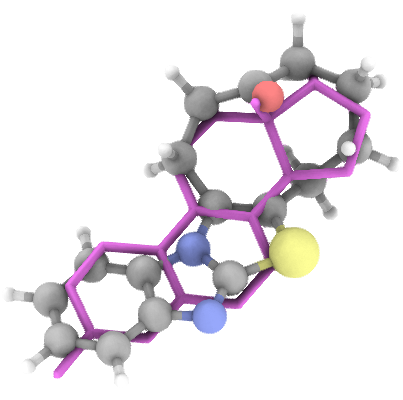
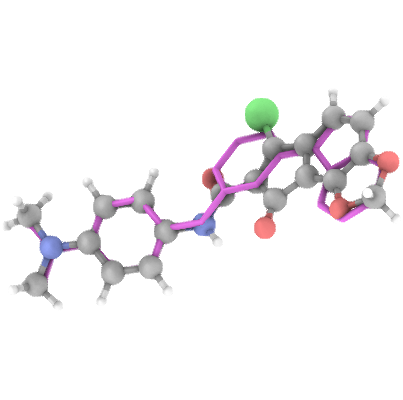
💾 Access & Licensing
The Python package and inference code are available on GitHub under Apache 2.0 License
The trained model Weights are available at
And are licensed under CC BY-NC-ND 4.0
The usage of the trained weights for any profit-generating activity is restricted.
For commercial licensing and inference-as-a-service, contact: Denis Sapegin
ONNX Inference:
For torch Free inference an ONNX version of the model is present.
Weights of the model in ONNX format are available at:
egnn_chembl_15_39.onnx
adj_mat_seer_chembl_15_39.onnx
from mlconfgen import MLConformerGeneratorONNX
from rdkit import Chem
model = MLConformerGeneratorONNX(
egnn_onnx="./egnn_chembl_15_39.onnx",
adj_mat_seer_onnx="./adj_mat_seer_chembl_15_39.onnx",
diffusion_steps=100,
)
reference = Chem.MolFromMolFile('./assets/demo_files/yibfeu.mol')
samples = model.generate_conformers(reference_conformer=reference, n_samples=20, variance=2)
Install ONNX GPU runtime (if needed):
pip install onnxruntime-gpu
Export to ONNX
An option to compile the model to ONNX is provided
requires onnxscript==0.2.2
pip install onnxscript
from mlconfgen import MLConformerGenerator
from onnx_export import export_to_onnx
model = MLConformerGenerator()
export_to_onnx(model)
This compiles and saves the ONNX files to: ./
Testing
To execute all tests (including slow generation ones)
pytest -v tests
To bypass generation tests
pytest -v tests -m "not slow"
Streamlit App
Running
- Move the trained PyTorch weights into
./streamlit_app
./streamlit_app/edm_moi_chembl_15_39.pt
./streamlit_app/adj_mat_seer_chembl_15_39.pt
-
Install the dependencies
pip install -r ./streamlit_app/requirements.txt -
Bring the app UI up:
cd ./streamlit_app streamlit run app.py
Streamlit App Development
-
To enable development mode for the 3D viewer (
stspeck), set_RELEASE = Falsein./streamlit/stspeck/__init__.py. -
Navigate to the 3D viewer frontend and start the development server:
cd ./frontend/speck/frontend npm run start
This will launch the dev server at
http://localhost:3001 -
In a separate terminal, run the Streamlit app from the root frontend directory:
cd ./streamlit_app streamlit run app.py
-
To build the production version of the 3D viewer, run:
cd ./streamlit_app/stspeck/frontend npm run build
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file mlconfgen-0.3.0.tar.gz.
File metadata
- Download URL: mlconfgen-0.3.0.tar.gz
- Upload date:
- Size: 66.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7f66a299e7f31277a1e817e33db5a491ae07bddcc272b56e0e1360ae7989f567
|
|
| MD5 |
650f333436609a07558f65b51c460cca
|
|
| BLAKE2b-256 |
a999db6892b2f9175ba9565e7d9224f6b72fcb77a11c8c9cd20df21801896f30
|
Provenance
The following attestation bundles were made for mlconfgen-0.3.0.tar.gz:
Publisher:
release.yaml on Membrizard/ml_conformer_generator
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
mlconfgen-0.3.0.tar.gz -
Subject digest:
7f66a299e7f31277a1e817e33db5a491ae07bddcc272b56e0e1360ae7989f567 - Sigstore transparency entry: 1109795611
- Sigstore integration time:
-
Permalink:
Membrizard/ml_conformer_generator@91b6de2b8a809c83a93cfb854dae6446804922bc -
Branch / Tag:
refs/tags/v0.3.0 - Owner: https://github.com/Membrizard
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yaml@91b6de2b8a809c83a93cfb854dae6446804922bc -
Trigger Event:
push
-
Statement type:
File details
Details for the file mlconfgen-0.3.0-py3-none-any.whl.
File metadata
- Download URL: mlconfgen-0.3.0-py3-none-any.whl
- Upload date:
- Size: 71.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b7e0da8640843578b587bbbf706e710b593892768058f171f0ac9a7ea46bf6c6
|
|
| MD5 |
17f5d648205409a106a1ba42cd471fa8
|
|
| BLAKE2b-256 |
44e412fe011645758d93401fc315206e08ddfb1dbbfd0279a749b0f78fc45a4c
|
Provenance
The following attestation bundles were made for mlconfgen-0.3.0-py3-none-any.whl:
Publisher:
release.yaml on Membrizard/ml_conformer_generator
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
mlconfgen-0.3.0-py3-none-any.whl -
Subject digest:
b7e0da8640843578b587bbbf706e710b593892768058f171f0ac9a7ea46bf6c6 - Sigstore transparency entry: 1109795618
- Sigstore integration time:
-
Permalink:
Membrizard/ml_conformer_generator@91b6de2b8a809c83a93cfb854dae6446804922bc -
Branch / Tag:
refs/tags/v0.3.0 - Owner: https://github.com/Membrizard
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yaml@91b6de2b8a809c83a93cfb854dae6446804922bc -
Trigger Event:
push
-
Statement type: