MultiMin: Multivariate Gaussian fitting
Project description
Introducing MultiMin
MultiMin is a Python package designed to provide numerical tools for fitting composed multivariate distributions to data. It is particularly useful for modelling complex multimodal distributions in N-dimensions.
These are the main features of MultiMin:
Multivariate Fitting: Tools for fitting composed multivariate normal distributions (CMND).
Visualization: Density plots and specific visualization utilities.
Statistical Analysis: Tools for handling covariance matrices and correlations.
Resources
Documentation including examples and full API documentation: https://multimin.readthedocs.io.
PyPI project page: https://pypi.org/project/multimin/.
Github repo: https://github.com/seap-udea/multimin
Installation
From PyPI
MultiMin is available on PyPI at https://pypi.org/project/multimin/. You can install it with:
pip install -U multiminIf you prefer, you can install the latest version of the developers taking it from the github repo:
pip install -U git+https://github.com/seap-udea/multiminFrom Sources
You can also install from the GitHub repository:
git clone https://github.com/seap-udea/multimin
cd multimin
pip install .For development, use an editable installation:
cd multimin
pip install -e .In Google Colab
If you use Google Colab, you can install MultiMin by executing:
!pip install -Uq multiminor
pip install -Uq git+https://github.com/seap-udea/multiminTheoretical Background
The core of MultiMin is the Composed Multivariate Normal Distribution (CMND). The theory behind it posits that any multivariate distribution function \(p(\tilde U):\Re^{N}\rightarrow\Re\), where \(\tilde U:(u_1,u_2,u_3,\ldots,u_N)\) are random variables, can be approximated with arbitrary precision by a normalized linear combination of \(M\) Multivariate Normal Distributions (MND):
where the multivariate normal \(\mathcal{N}(\tilde U; \tilde \mu, \Sigma)\) with mean vector \(\tilde \mu\) and covariance matrix \(\Sigma\) is given by:
The covariance matrix \(\Sigma\) elements are defined as \(\Sigma_{ij} = \rho_{ij}\sigma_{i}\sigma_{j}\), where \(\sigma_i\) is the standard deviation of \(u_i\) and \(\rho_{ij}\) is the correlation coefficient between variable \(u_i\) and \(u_j\) (\(-1<\rho_{ij}<1\), \(\rho_{ii}=1\)).
The normalization condition on \(p(\tilde U)\) implies that the set of weights \(\{w_k\}_M\) are also normalized, i.e., \(\sum_i w_i=1\).
Fitting procedure
To estimate the parameters of the CMND that best describe a given dataset , we use the Likelihood Statistics method.
Given a dataset of \(S\) objects with state vectors \(\{\tilde U_k\}_{k=1}^S\), the likelihood \(\mathcal{L}\) of the CMND parameters is defined as the product of the probability densities evaluated at each data point:
The goal is to find the set of parameters (weights, means, and covariances) that maximize this likelihood. In practice, it is numerically more stable to minimize the negative normalized log-likelihood:
This approach allows us to fit the distribution without making strong assumptions about the underlying normality of the data, effectively treating the CMND as a series expansion of the true probability density function.
In MultiMin, we use the scipy.optimize.minimize function to find the set of parameters that minimize the negative normalized log-likelihood.
Quickstart
Getting started with MultiMin is straightforward. Import the package:
import multimin as mnNOTE: If you are working in Google Colab, load the matplotlib backend before producing plots:
%matplotlib inline
Here is a basic example of how to use MultiMin to fit a 3D distribution composed of 2 Multivariate Normals.
1. Define a true distribution
First, we define a distribution from which we will generate synthetic data. We use a Composed Multivariate Normal Distribution (CMND) with 2 Gaussian components (ngauss=2) in 3 dimensions (nvars=3).
import numpy as np
import multimin as mn
# Define parameters for 2 Gaussian components
weights = [0.5, 0.5]
mus = [[1.0, 0.5, -0.5], [1.0, -0.5, +0.5]]
sigmas = [[1, 1.2, 2.3], [0.8, 0.2, 3.3]]
deg = np.pi/180
angles = [
[10*deg, 30*deg, 20*deg],
[-20*deg, 0*deg, 30*deg],
]
# Calculate covariance matrices from rotation angles
Sigmas = mn.Stats.calc_covariance_from_rotation(sigmas, angles)
# Create the CMND object
CMND = mn.ComposedMultiVariateNormal(mus=mus, weights=weights, Sigmas=Sigmas)2. Generate sample data
We generate 5000 random samples from this distribution to serve as our “observed” data.
np.random.seed(1)
sample = CMND.rvs(5000)3. Visualize the data
We can check the distribution of the generated data using DensityPlot.
import matplotlib.pyplot as plt
# Define properties labels
properties = dict(
x=dict(label=r"$x$", range=None),
y=dict(label=r"$y$", range=None),
z=dict(label=r"$z$", range=None),
)
# Plot the density plot
G = mn.DensityPlot(properties, figsize=3)
hargs=dict(bins=30,cmap='Spectral_r')
histogram=G.plot_hist(sample,**hargs)
sargs=dict(s=0.5,edgecolor='None',color='r')
scatter=G.scatter_plot(sample,**sargs)
Data Scatter Plot
4. Initialize the Fitter and Run the Fit
We initialize the FitCMND handler with the expected number of Gaussians (2) and variables (3). We then run the fitting procedure.
# Initialize the fitter
F = mn.FitCMND(ngauss=2, nvars=3)
# Run the fit (using advance=True for better convergence on complex models)
F.fit_data(sample, advance=True)5. Check and Plot Results
Finally, we visualize the fitted distribution compared to the data.
# Plot the fit result
G = F.plot_fit(
props=["x", "y", "z"],
hargs=dict(bins=30, cmap='YlGn'),
sargs=dict(s=0.2, edgecolor='None', color='r'),
figsize=3
)
Fit Result
6. Inspect Parameters and Get Explicit PDF Function
You can tabulate the fitted parameters and obtain an explicit Python function that evaluates the fitted PDF. Below, each step is shown with its output.
Stage 1: Tabulate the fitted CMND
F.cmnd.tabulate(sort_by='weight')Output:
w mu_1 mu_2 mu_3 sigma_1 sigma_2 sigma_3 rho_12 rho_13 rho_23 component 2 0.509108 1.019245 -0.480997 0.618821 0.794906 0.245786 3.327537 0.539417 -0.008936 -0.017769 1 0.490892 0.957687 0.517584 -0.463392 1.039489 1.538029 2.116544 -0.209695 0.121184 -0.527142
Stage 2: Get the source code and a callable function
code, cmnd = F.cmnd.get_function()Output (the printed code, which you can copy):
from multimin import nmd
def cmnd(X):
mu1_1 = 0.957687
mu1_2 = 0.517584
mu1_3 = -0.463392
mu1 = [mu1_1, mu1_2, mu1_3]
Sigma1 = [[1.080538, -0.335252, 0.266619], [-0.335252, 2.365532, -1.716008], [0.266619, -1.716008, 4.479757]]
n1 = nmd(X, mu1, Sigma1)
mu2_1 = 1.019245
mu2_2 = -0.480997
mu2_3 = 0.618821
mu2 = [mu2_1, mu2_2, mu2_3]
Sigma2 = [[0.631876, 0.10539, -0.023637], [0.10539, 0.060411, -0.014533], [-0.023637, -0.014533, 11.072504]]
n2 = nmd(X, mu2, Sigma2)
w1 = 0.490892
w2 = 0.509108
return (
w1*n1
+ w2*n2
)
Stage 3: Evaluate the PDF at a point
cmnd([1.0, 0.5, -0.5])Output:
0.011073778538439395
Stage 4: LaTeX output for papers
You can get the fitted PDF as a LaTeX string (suitable for inclusion in papers) with parameter values and the definition of the normal distribution:
latex_str, _ = F.cmnd.get_function(print_code=False, type='latex', decimals=4)
print(latex_str)Output:
where
Here the normal distribution is defined as:
A parameter table in LaTeX is also available via F.cmnd.tabulate(sort_by='weight', type='latex').
Truncated multivariate distributions.
In real problems the domain of the variables is not infinite but bounded into a semi-finite region.
If we start from the unbounded multivariate normal distribution:
Let \(T\subset\{l,\dots,m\}\), where \(l\leq k\) and \(m\leq k\) be the set of indices of the truncated variables, and let \(a_i<b_i\) be the truncation bounds for \(i\in S\). Define the truncation region:
with the remaining coordinates \(i\notin T\) unbounded. The partially-truncated multivariate normal distribution is defined by
where \(\mathbf{1}_{A_T}\) is the indicator function of \(A_T\) and the normalization constant is
Example: univariate truncated mixture
Define a mixture of two Gaussians on the interval \([0, 1]\) with the domain parameter, generate data, and fit with FitCMND(..., domain=[[0, 1]]):
import numpy as np
import multimin as mn
# Truncated mixture of 2 Gaussians on [0, 1]
CMND_1d = mn.ComposedMultiVariateNormal(
mus=[0.2, 0.8],
weights=[0.5, 0.5],
Sigmas=[0.01, 0.03],
domain=[[0, 1]],
)
np.random.seed(1)
data_1d = CMND_1d.rvs(5000)
# Fit with same domain so likelihood and means respect [0, 1]
F_1d = mn.FitCMND(ngauss=2, nvars=1, domain=[[0, 1]])
F_1d.fit_data(data_1d, advance=True)
G = F_1d.plot_fit(hargs=dict(bins=40), sargs=dict(s=0.5, alpha=0.6))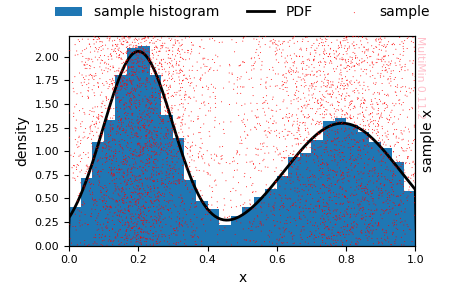
Truncated 1D fit
You can also extract an explicit callable function for the fitted truncated PDF (including the bounds) and evaluate it safely outside the interval.
function, cmnd = F_1d.cmnd.get_function()Output (the printed code, which you can copy):
import numpy as np
from multimin import tnmd
def cmnd(X):
a = 0.0
b = 1.0
mu1_1 = 0.200467
sigma1_1 = 0.009683
n1 = tnmd(X, mu1_1, sigma1_1, a, b)
mu2_1 = 0.801063
sigma2_1 = 0.030392
n2 = tnmd(X, mu2_1, sigma2_1, a, b)
w1 = 0.504151
w2 = 0.495849
return (
w1*n1
+ w2*n2
)
Evaluate the fitted PDF at a point inside the domain and outside the domain:
cmnd(0.5), cmnd(-0.2)Output:
(0.3128645172339761, 0.0)
For papers, you can also generate a LaTeX/Markdown description that includes the truncation information:
function_str, _ = F_1d.cmnd.get_function(print_code=False, type='latex', decimals=4)
print(function_str)Output:
Finite domain. The following variables are truncated (the rest are unbounded):
Variable \(x_{1}\) (index 1): domain \([0.0, 1.0]\).
Truncation region: \(A_T = \{\tilde{U} \in \mathbb{R}^k : a_i \le \tilde{U}_i \le b_i \;\forall i \in T\}\), with \(T\) the set of truncated indices.
where
Truncated normal. The unbounded normal is
The truncation region is \(A_T = \{\tilde{U} \in \mathbb{R}^k : a_i \le \tilde{U}_i \le b_i \;\forall i \in T\}\). The partially truncated normal is
where \(\mathbf{1}_{A_T}\) is the indicator of \(A_T\) and the normalization constant is
See examples/multimin_truncated_tutorial.ipynb for 3D truncated examples and more detail.
Citation
The numerical tools and codes provided in this package have been developed and tested over several years of scientific research.
If you use MultiMin in your research, please cite:
@software{multimin2026,
author = {Zuluaga, Jorge I.},
title = {MultiMin: Multivariate Gaussian fitting},
year = {2026},
url = {https://github.com/seap-udea/multimin}
}What’s New
For a detailed list of changes and new features, see WHATSNEW.md.
Contributing
We welcome contributions! If you’re interested in contributing to MultiMin, please:
Fork the repository
Create a feature branch
Make your changes
Submit a pull request
Please read the CONTRIBUTING.md file for more information.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file multimin-0.9.3.tar.gz.
File metadata
- Download URL: multimin-0.9.3.tar.gz
- Upload date:
- Size: 4.9 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
42f23ce491e64dd1417a3dfd7f74e50da91c263cb8579b0e6ef454ef852ba8b8
|
|
| MD5 |
f074ae9ae32562de6721c03529d62395
|
|
| BLAKE2b-256 |
c5267104c349d4ecbd80d318a56f85e6d4fbafdf8302e71852cce51124bc7310
|
File details
Details for the file multimin-0.9.3-py3-none-any.whl.
File metadata
- Download URL: multimin-0.9.3-py3-none-any.whl
- Upload date:
- Size: 4.9 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6ab65be1545601092208d5bd9a0dc9184338641cf8dd23bae87c19064839c0a2
|
|
| MD5 |
56514cd78c9e9e1b34570435aff5f52d
|
|
| BLAKE2b-256 |
86298f8bb5f1c9084d251a68c4a5da09145fb062d4aecafb8a5350c49bffa09c
|

















