A plugin to quickly generate dense ground truth with sparse labels
Project description
napari-bootstrapper
A napari plugin to quickly generate dense 3D segmentations from sparse 2D labels.
Sparse 2D annotations made in ~10 minutes on a single section can produce dense 3D segmentations that are good starting points for refinement. Based on the 2D→3D method described in Sparse Annotation is Sufficient for Bootstrapping Dense Segmentation.
For larger volumes, dedicated 3D models, and block-wise processing, see the Bootstrapper CLI.
| Dataset | Data Type | Video |
|---|---|---|
| CREMI C | 3D volumetric stack | 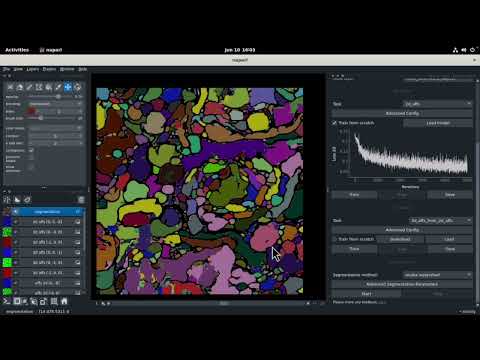 |
| Fluo-C2DL-Huh7 | 2D + time series | 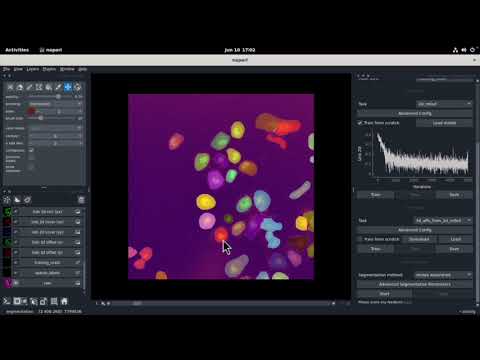 |
Getting Started
Install
conda create -n napari-bootstrapper -c conda-forge python==3.11 napari pyqt
conda activate napari-bootstrapper
pip install napari-bootstrapper
Or install the latest development version:
pip install git+https://github.com/ucsdmanorlab/napari-bootstrapper.git
Launch
conda activate napari-bootstrapper
napari
Open the Bootstrapper widget from Plugins → napari-bootstrapper.
How It Works
The plugin has four sections that follow a simple workflow:
1. Data
Load a 3D image (or 4D with a channels dimension) and create sparse 2D labels on one or a few slices. Click "Make mask" to generate a binary training mask.
We recommend using a foundation model to make sparse 2D labels, like micro-sam.
2. Train a 2D Model
Train a 2D model on your sparse labels. Three task types are available:
2d_affs— affinities2d_lsd— local shape descriptors2d_mtlsd— multi-task (both)
Or load a pretrained checkpoint.
3. Load a 3D Model
The 3D model lifts 2D predictions into 3D affinities. Pretrained weights are recommended — just click "Download".
4. Segment
Click "Start" to run the full 2D→3D inference pipeline. The output is an instance segmentation produced via mutex watershed or connected components.
Proofreading
We provide a separate widget for refining segmentations quickly. Select labels by placing points on them or entering label IDs manually. Operations can be applied per-slice (2D) or on the full volume (3D).
- Morphology — Dilate, erode, open, close, fill holes. Stenciled (3×3×3) or spherical (variable radius). Uses fastmorph.
- Merge / Split — Merge labels, split with watershed markers, or delete. Uses fastremap.
- Filter — Remove by size (min/max voxels), keep K largest, remove outliers by sigma, filter by Z-slice count, relabel connected components. Uses cc3d.
Citation
If you find this useful, please cite our preprint:
@article {Thiyagarajan2024.06.14.599135,
author = {Thiyagarajan, Vijay Venu and Sheridan, Arlo and Harris, Kristen M. and Manor, Uri},
title = {Sparse Annotation is Sufficient for Bootstrapping Dense Segmentation},
year = {2024},
doi = {10.1101/2024.06.14.599135},
URL = {https://www.biorxiv.org/content/10.1101/2024.06.14.599135v2},
}
Issues
If you encounter any problems, please file an issue along with a detailed description.
Acknowledgements
- micro-sam — Making foundation models like SAM accessible to the community.
- empanada-napari — Proofreading widget inspiration.
- napari-cellulus — Development scaffolding.
Funding
Chan-Zuckerberg Imaging Scientist Award DOI https://doi.org/10.37921/694870itnyzk from the Chan Zuckerberg Initiative DAF, an advised fund of Silicon Valley Community Foundation (funder DOI 10.13039/100014989).
NSF NeuroNex Technology Hub Award (1707356), NSF NeuroNex2 Award (2014862)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_bootstrapper-0.3.0.tar.gz.
File metadata
- Download URL: napari_bootstrapper-0.3.0.tar.gz
- Upload date:
- Size: 11.5 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a86eeb4dcb49427923975acd93d617d1dcd32a5ff4e7025fd3305d5585c6a156
|
|
| MD5 |
8b7e992a2aeed1361e23c28fff5c2e31
|
|
| BLAKE2b-256 |
591ffdb9f7eb720780e602dd4b4d68458081544fbe6b1fdcefff035d0fbbb495
|
File details
Details for the file napari_bootstrapper-0.3.0-py3-none-any.whl.
File metadata
- Download URL: napari_bootstrapper-0.3.0-py3-none-any.whl
- Upload date:
- Size: 11.6 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4272723d85962a9af66b0516b301069366c9dc02c114cf05fbad8e9e7ad94a28
|
|
| MD5 |
31ad1df326e0e6f9b1a869b8973be884
|
|
| BLAKE2b-256 |
422b1e4ffa7448ccc467cef2ecf9a90edae39dc46806b51db803179cad7cb060
|


















