Data Inspection Plugin, designed to streamline file navigation and enhance the efficiency of data inspection.
Project description
Napari Data Inspection
A napari plugin for fast, high-throughput inspection of 2D/3D segmentation datasets. Select one or more images and/or labels layers; files are paired and loaded automatically so you can browse entire datasets without manual effort.
Features
- Multi-folder pairing: any number of image/label folders
- Prefetching & caching for steamless navigation
- Supports common formats (e.g., NIfTI, TIFF, PNG, NRRD, MHA, B2ND) out of the box.
- Extensible loaders (add your own formats if needed).
Installation
# a) Install the plugin
pip install napari-data-inspection
# b) Install the plugin and napari if necessary
pip install napari-data-inspection[all]
Quickstart
napari -w napari-data-inspection
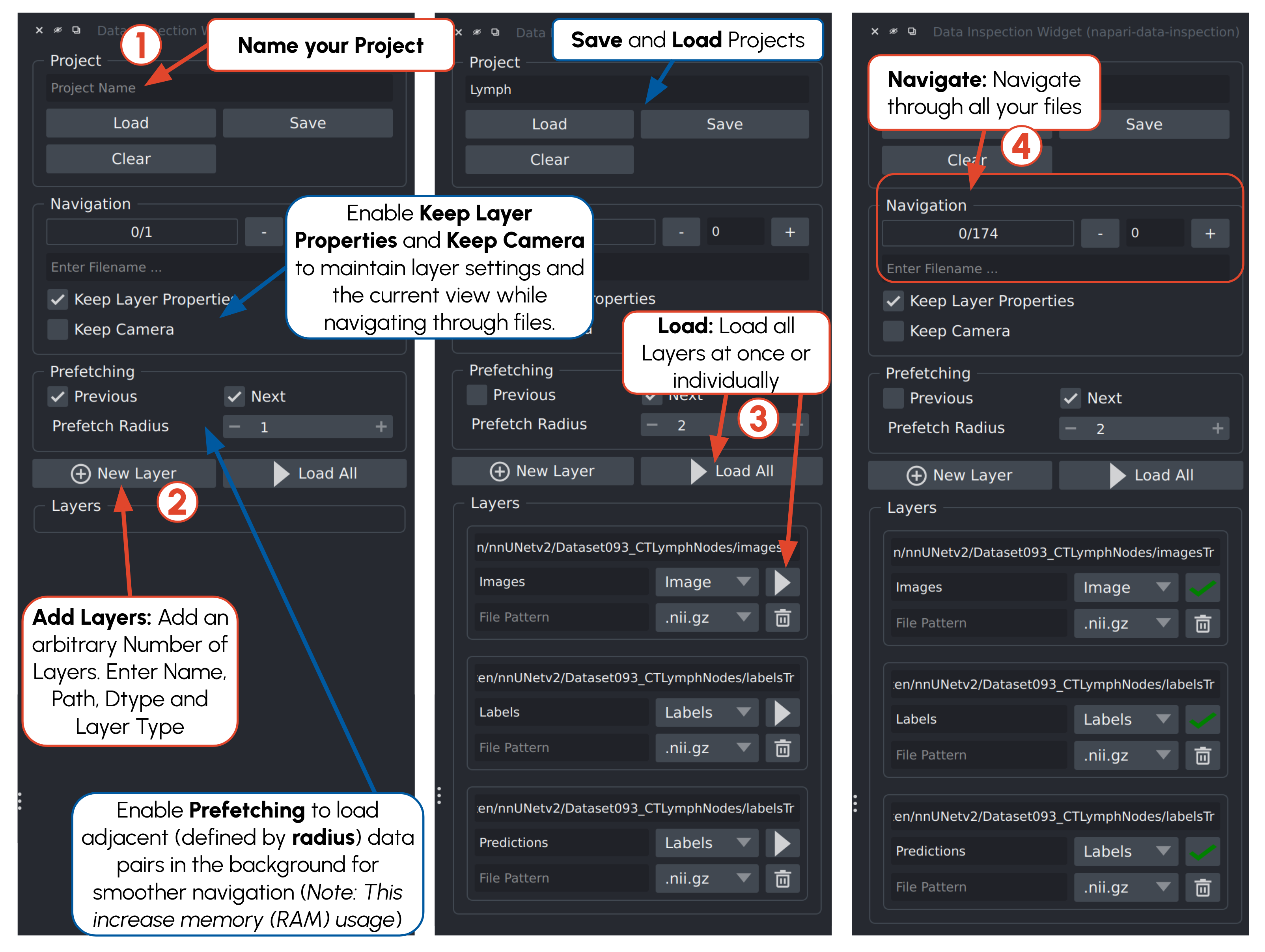
Data organization
- Filter files by patterns (e.g., *_img.nii.gz, *_seg.nii.gz) and/or separate folders per layer.
- Number of files must match across layers; pairs are made by sorted order.
1. Separate folders per layer
Put each layer’s files in its own directory.
/data/images/
case001.nii.gz
case002.nii.gz
/data/labels/
case001.nii.gz
case002.nii.gz
2. By Patterns
Use patterns (globs) per layer.
/data/
case001_img.nii.gz
case001_seg.nii.gz
case002_img.nii.gz
case002_seg.nii.gz
Patterns
Images: *_img.nii.gz
Labels: *_seg.nii.gz
Supported file types
The data loading is based on the ViData package. The following Extensions and Backends are available.
| Extension(s) | Backend(s) | Notes |
|---|---|---|
.png, .jpg, .jpeg, .bmp |
imageio |
Standard 2D image formats |
.tif, .tiff |
tifffile |
Multipage TIFF; high bit-depths supported |
.nii.gz, .nii, .mha, .nrrd |
sitk |
Medical image formats (3D volumes) |
.nii.gz, .nii |
nibabel |
Alternative medical imaging backend |
.b2nd |
blosc2 |
Compressed N-dimensional arrays |
.b2nd |
blosc2pkl |
Compressed N-dimensional arrays with metadata in a separate .pkl |
.npy |
numpy |
Single NumPy array |
Custom Load Functions
- Register a reader with a decorator.
- Reader must return (numpy_array, metadata_dict).
- Registration happens at import time—make sure this module is imported (e.g., from your package’s init.py).
- See here for an example.
- metadata should contain an "affine" if entry, if any spatial transformation should be applied
# custom_io_template.py — fill in the TODOs and import this module somewhere at startup.
import numpy as np
from typing import Tuple, Dict, List
# TODO: import your backend library (e.g., imageio, tifffile, nibabel, SimpleITK, ...)
# import imageio.v3 as iio
from vidata.registry import register_loader, register_writer
# --------------------------- READER ------------------------------------------
# Replace file extension and backend name to your custom function
@register_loader("image", ".png", ".jpg", ".jpeg", ".bmp", backend="imageio") # To Register Image Loading
@register_loader("mask", ".png", ".bmp", backend="imageio") # To Register Label Loading
def load_custom(file: str) -> tuple[np.ndarray, dict]:
"""
Load a file and return (data, metadata).
metadata can be empty or include keys like: spacing, origin, direction, shear, dtype, etc.
"""
# data = iio.imread(file) # example for imageio
data = ... # TODO: replace
meta = {} # TODO: replace
return data, meta
Torch Dataset Inspection
- Rapid inspection of PyTorch datasets in Napari by browsing images and corresponding labels directly from any torch.utils.data.Dataset
from napari_data_inspection.dataset_inspection import DatasetInspectionWidget, run_dataset_inspection
from torch.utils.data import Dataset
# Example: inspect a PyTorch dataset
dataset = MyCustomDataset(...) # any torch Dataset
run_dataset_inspection(dataset, channel_first=True, rescale=True, no_label=False, bg_class=0)
Acknowledgments


This repository is developed and maintained by the Applied Computer Vision Lab (ACVL) of Helmholtz Imaging and the Division of Medical Image Computing at DKFZ.
This napari plugin was generated with copier using the napari-plugin-template.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_data_inspection-1.0.2.tar.gz.
File metadata
- Download URL: napari_data_inspection-1.0.2.tar.gz
- Upload date:
- Size: 25.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
053d1c26a9d94a4e9467005cab874ea3eaefe07f6ad363a8649d4e2eea430dea
|
|
| MD5 |
60a95eda9a0d8fc0fb57281cd581d562
|
|
| BLAKE2b-256 |
a598cc22b5fc96e7a737d66bbd9fca656f092f773ac9d557961975fe4fa788da
|
File details
Details for the file napari_data_inspection-1.0.2-py3-none-any.whl.
File metadata
- Download URL: napari_data_inspection-1.0.2-py3-none-any.whl
- Upload date:
- Size: 25.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3abb2432d21778e62ac5ea3c3e79bdf4c04d8c066ed0c54bc00291c38010ecf7
|
|
| MD5 |
f1ab83095522bc405d9276a309b47220
|
|
| BLAKE2b-256 |
916cf4aa633abbe91bd60c5d1e0953f6f6b7c918903ff7801c553e12c4d9636c
|











