nnInteractive plugin for Napari
Project description

nnInteractive: Redefining 3D Promptable Segmentation
This repository contains the napari plugin for nnInteractive. Check out the python backend and MITK integration for more.
What is nnInteractive?
Isensee, F.*, Rokuss, M.*, Krämer, L.*, Dinkelacker, S., Ravindran, A., Stritzke, F., Hamm, B., Wald, T., Langenberg, M., Ulrich, C., Deissler, J., Floca, R., & Maier-Hein, K. (2025). nnInteractive: Redefining 3D Promptable Segmentation. https://arxiv.org/abs/2503.08373
*: equal contribution
Abstract:
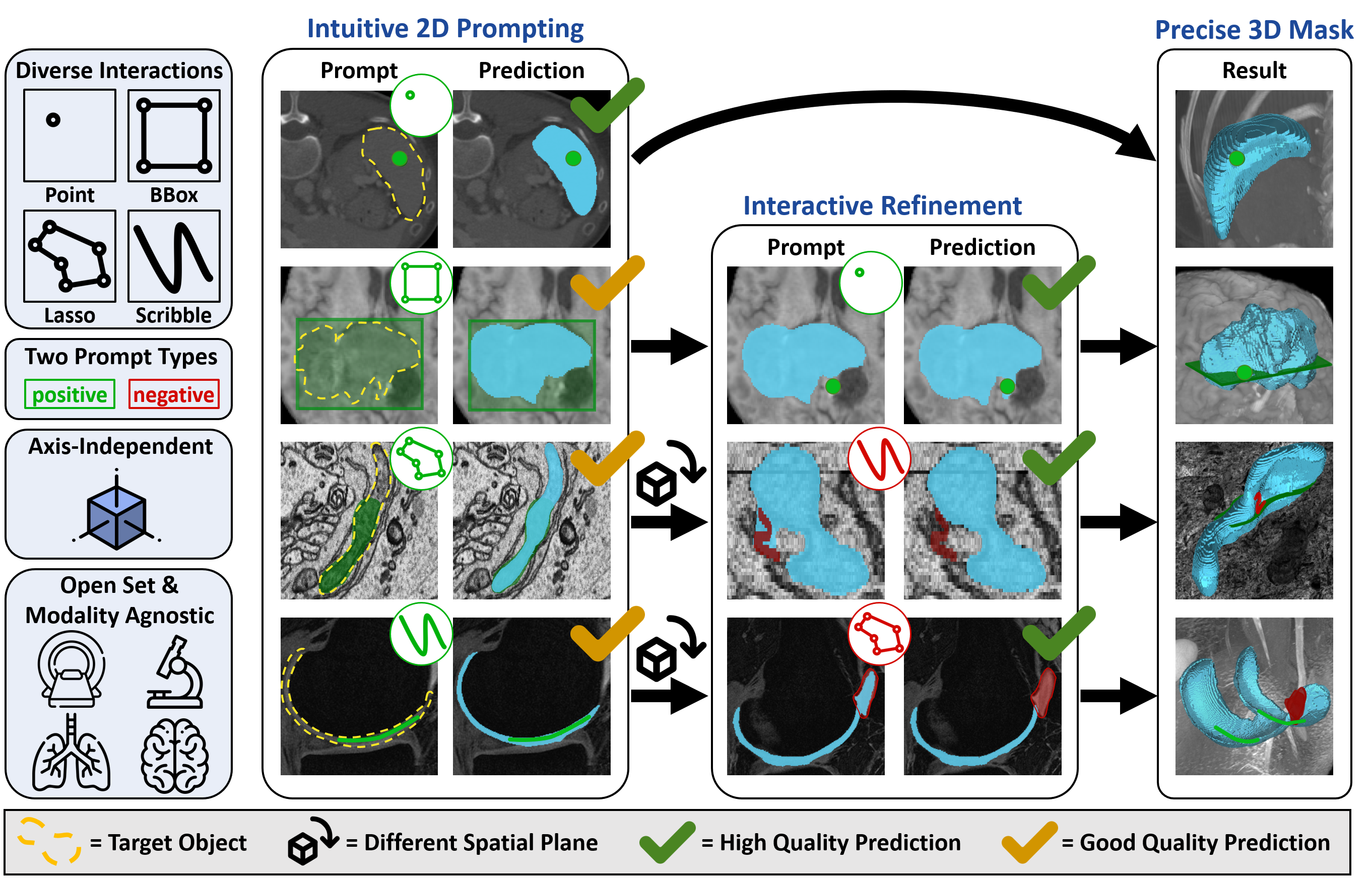
Accurate and efficient 3D segmentation is essential for both clinical and research applications. While foundation models like SAM have revolutionized interactive segmentation, their 2D design and domain shift limitations make them ill-suited for 3D medical images. Current adaptations address some of these challenges but remain limited, either lacking volumetric awareness, offering restricted interactivity, or supporting only a small set of structures and modalities. Usability also remains a challenge, as current tools are rarely integrated into established imaging platforms and often rely on cumbersome web-based interfaces with restricted functionality. We introduce nnInteractive, the first comprehensive 3D interactive open-set segmentation method. It supports diverse prompts—including points, scribbles, boxes, and a novel lasso prompt—while leveraging intuitive 2D interactions to generate full 3D segmentations. Trained on 120+ diverse volumetric 3D datasets (CT, MRI, PET, 3D Microscopy, etc.), nnInteractive sets a new state-of-the-art in accuracy, adaptability, and usability. Crucially, it is the first method integrated into widely used image viewers (e.g., Napari, MITK), ensuring broad accessibility for real-world clinical and research applications. Extensive benchmarking demonstrates that nnInteractive far surpasses existing methods, setting a new standard for AI-driven interactive 3D segmentation.
Demo Videos
Installation
Prerequisites
You need a Linux or Windows computer with a Nvidia GPU. 10GB of VRAM is recommended. Small objects should work with <6GB.
1. Create a virtual environment:
nnInteractive supports Python 3.10+ and works with Conda, pip, or any other virtual environment. Here’s an example using Conda:
conda create -n nnInteractive python=3.12
conda activate nnInteractive
2. Install the correct PyTorch for your system
Go to the PyTorch homepage and pick the right configuration. Note that since recently PyTorch needs to be installed via pip. This is fine to do within your conda environment.
For Ubuntu with a Nvidia GPU, pick 'stable', 'Linux', 'Pip', 'Python', 'CUDA12.6' (if all drivers are up to date, otherwise use and older version):
pip3 install torch torchvision --index-url https://download.pytorch.org/whl/cu126
3. Install this repository + dependencies via
Install napari if necessary
pip install napari[all]
Install the plugin via pip:
pip install napari-nninteractive
Or clone and install this repository:
git clone https://github.com/MIC-DKFZ/napari-nninteractive
cd napari-nninteractive
pip install -e .
Note: Model weights are automatically downloaded on first use. This can take up to a couple of minutes depending on your internet connection
Getting Started
Use one of these three options to start napari and activate the plugin. Afterward, Drag and drop your images into napari.
*Note if getting asked which plugin to use for opening .nii.gz files use napari-nifti.
a) Start napari, then Plugins -> nnInteractive.
napari
b) Run this to start napari with the plugin already started.
napari -w napari-nninteractive
c) Run this to start napari with the plugin and open an image directly
napari demo_data/liver_145_0000.nii.gz -w napari-nninteractive
How to use
Note: To open Nifti (.nii.gz, .nii) files we recommend to select napari-nifti.

Citation
When using nnInteractive, please cite the following paper:
Isensee, F.*, Rokuss, M.*, Krämer, L.*, Dinkelacker, S., Ravindran, A., Stritzke, F., Hamm, B., Wald, T., Langenberg, M., Ulrich, C., Deissler, J., Floca, R., & Maier-Hein, K. (2025). nnInteractive: Redefining 3D Promptable Segmentation. https://arxiv.org/abs/2503.08373
*: equal contribution
License
Note that while this repository is available under Apache-2.0 license (see LICENSE), the model checkpoint is Creative Commons Attribution Non Commercial Share Alike 4.0!
Acknowledgments


This repository is developed and maintained by the Applied Computer Vision Lab (ACVL) of Helmholtz Imaging and the Division of Medical Image Computing at DKFZ.
This napari plugin was generated with copier using the napari-plugin-template.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_nninteractive-1.0.5.tar.gz.
File metadata
- Download URL: napari_nninteractive-1.0.5.tar.gz
- Upload date:
- Size: 33.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7481c1675f7cbf256e0aed06d2cd25ba26c3a66d4a390ef9c4b1027e47a793a9
|
|
| MD5 |
0bda5782b1c0b44dfde6d5d9b370d705
|
|
| BLAKE2b-256 |
f374208e24afb4ed5140a84a463228e1080eeef8a4ba52585c4c5c5101b1eb6e
|
File details
Details for the file napari_nninteractive-1.0.5-py3-none-any.whl.
File metadata
- Download URL: napari_nninteractive-1.0.5-py3-none-any.whl
- Upload date:
- Size: 36.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9afdab4540fff80488fbc0628e45da1ff95ccfbe29ab0ed9a0746220dace873e
|
|
| MD5 |
ae828d5fdfcf15fcec71a2f6b6259937
|
|
| BLAKE2b-256 |
0a6ec983758576d6b981a25a3e6de46204a15712809743382c2b1a53fc541c25
|













