A Napari plugin to detect and count nuclei on phase contrast images
Project description
NuclePhaser: Cell Proliferation Measurement & Cell Tracking Assistant Plugin for Timelapse Images
A Napari plugin for automated cell nuclei detection, proliferation and population growth analysis, and single-cell tracking in brightfield and fluorescent nuclei timelapse microscopy images.
WARNING!!! Further description isn't displayed correctly at Napari Hub. For correctly displayed description, visit our GITHUB PAGE.
napari-nuclephaser is an open-source Napari plugin designed for scientists who need to measure cell proliferation rates, analyze population growth, and perform individual cell tracking on timelapse microscopy images. It utilizes Ultralytics YOLO object detection models and obss/sahi sliced inference methods to detect cell nuclei on brightfield and fluorescent images of any size, including large whole slide ones. Learn more with documentation and paper.
NEW! If default NuclePhaser models aren't accurate enough on your specific use case, you can finetune them for free with cloud GPU using Google Colab NuclePhaser finetuning notebook. It is specifically designed for users without coding experience.
Nuclei detection
We trained a series of YOLOv5 and YOLOv11 models to detect nuclei on phase contrast images. It can be used for counting cells or for individual cell tracking (using nuclei detections as tracking markers). Prominent features of this approach are:
- Napari-nuclephaser plugin includes obss/sahi functionality, allowing detection on images of arbitrary sizes.
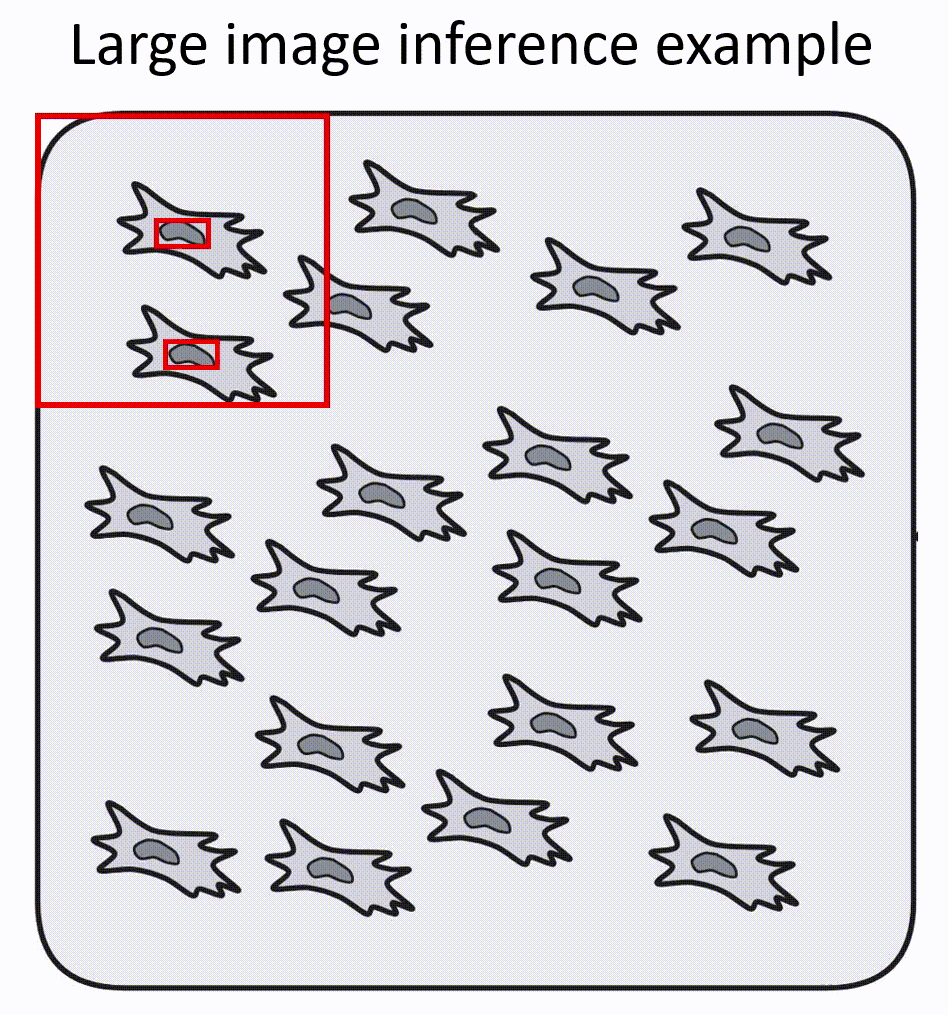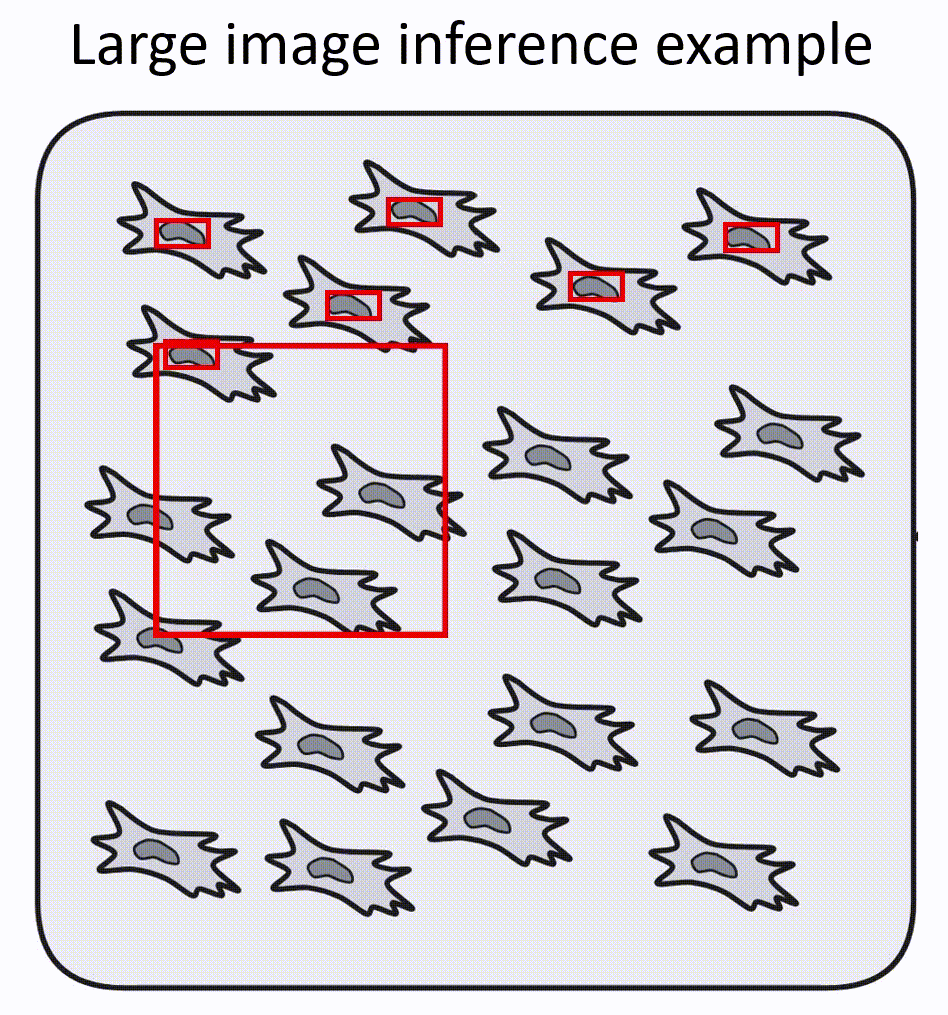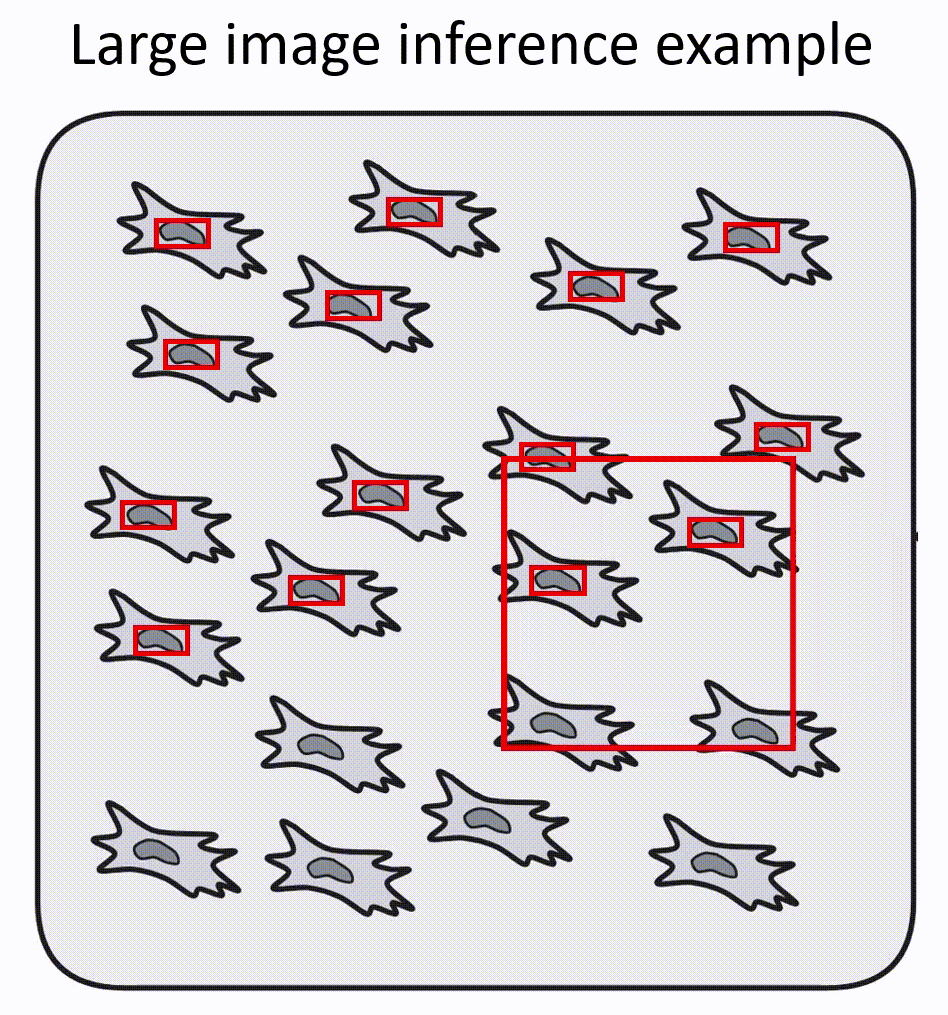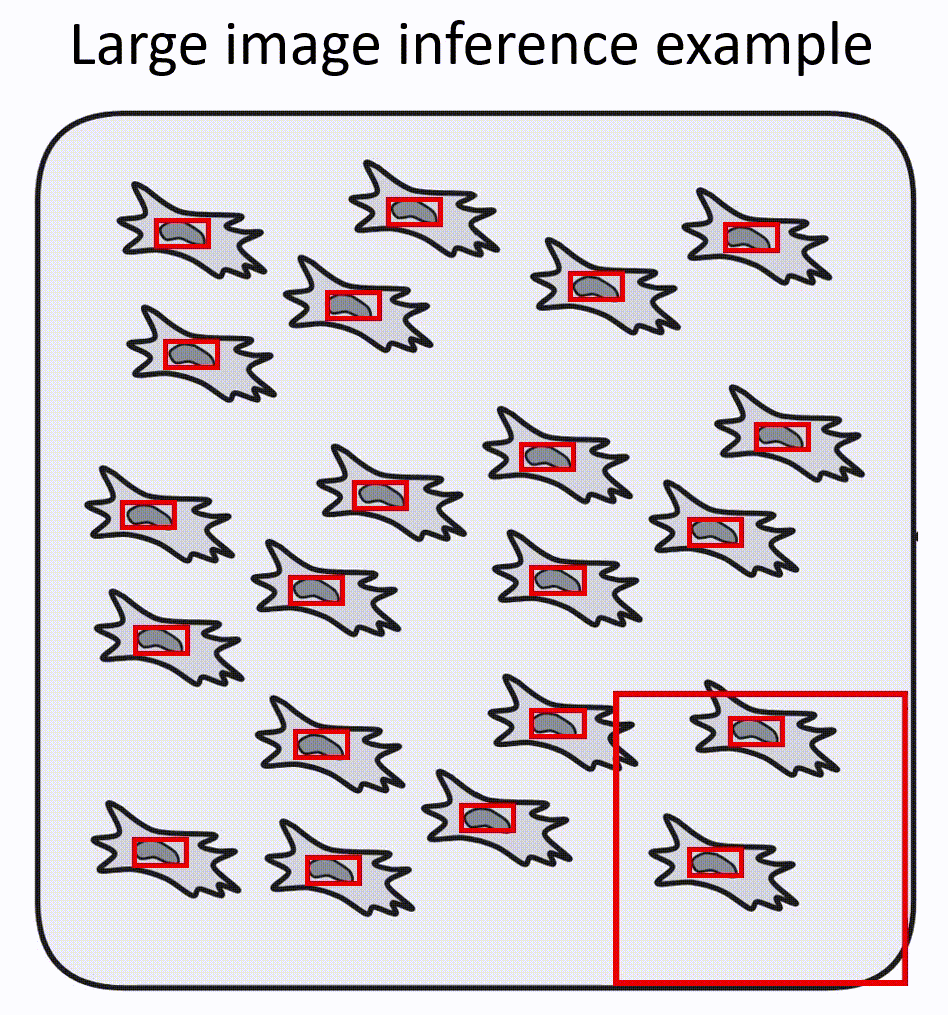
- YOLO models are fast, providing reasonable inference speed even with CPU.
- Ability to predict and automatically count nuclei on stacks of images, making it convenient for cell population growth studies and individual cell tracking.
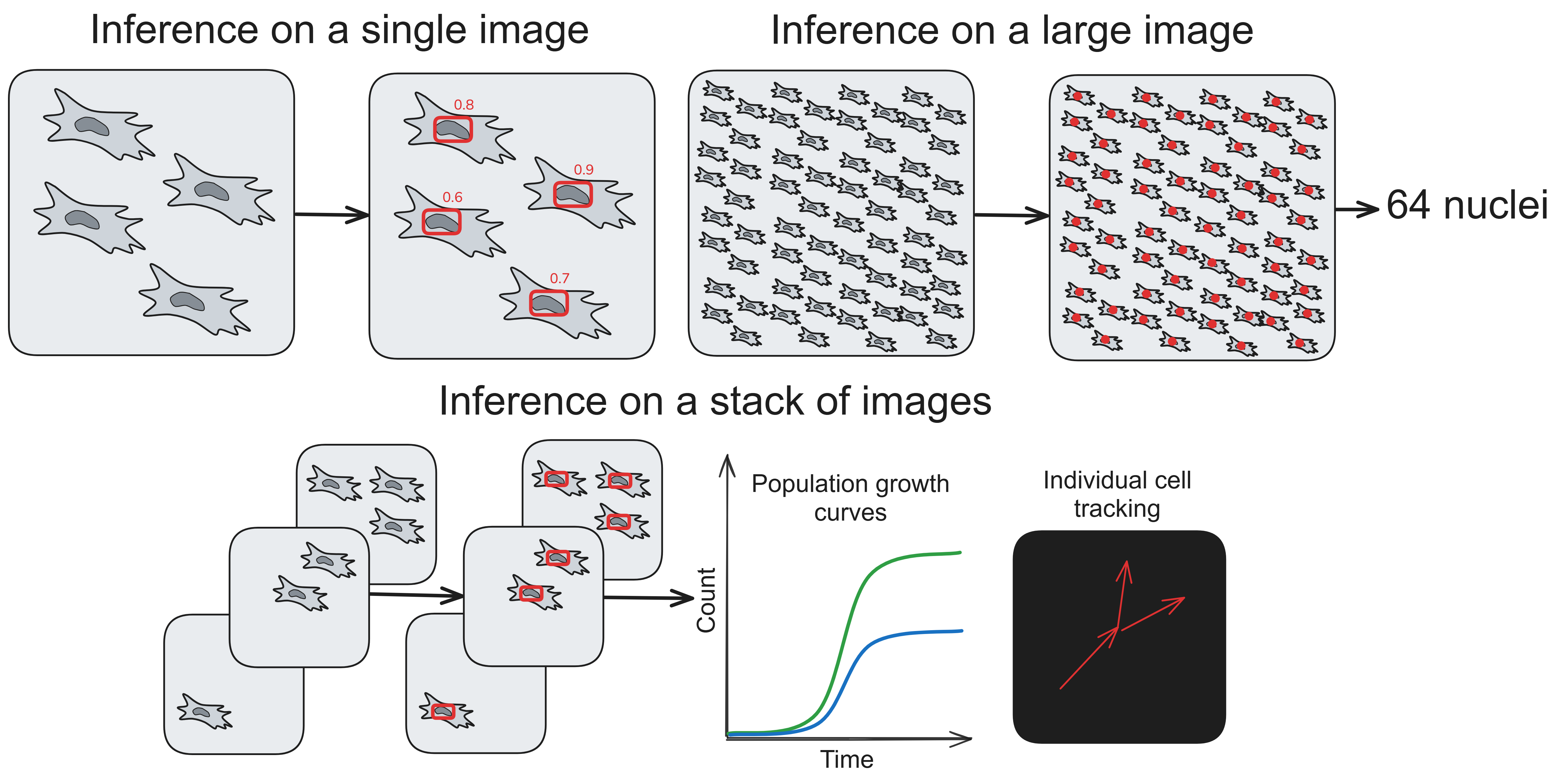
- Calibration algorithm that allows measuring accuracy for each specific use case.
Calibration algorithm
Result of object detection model inference is highly dependent on confidence threshold parameter.
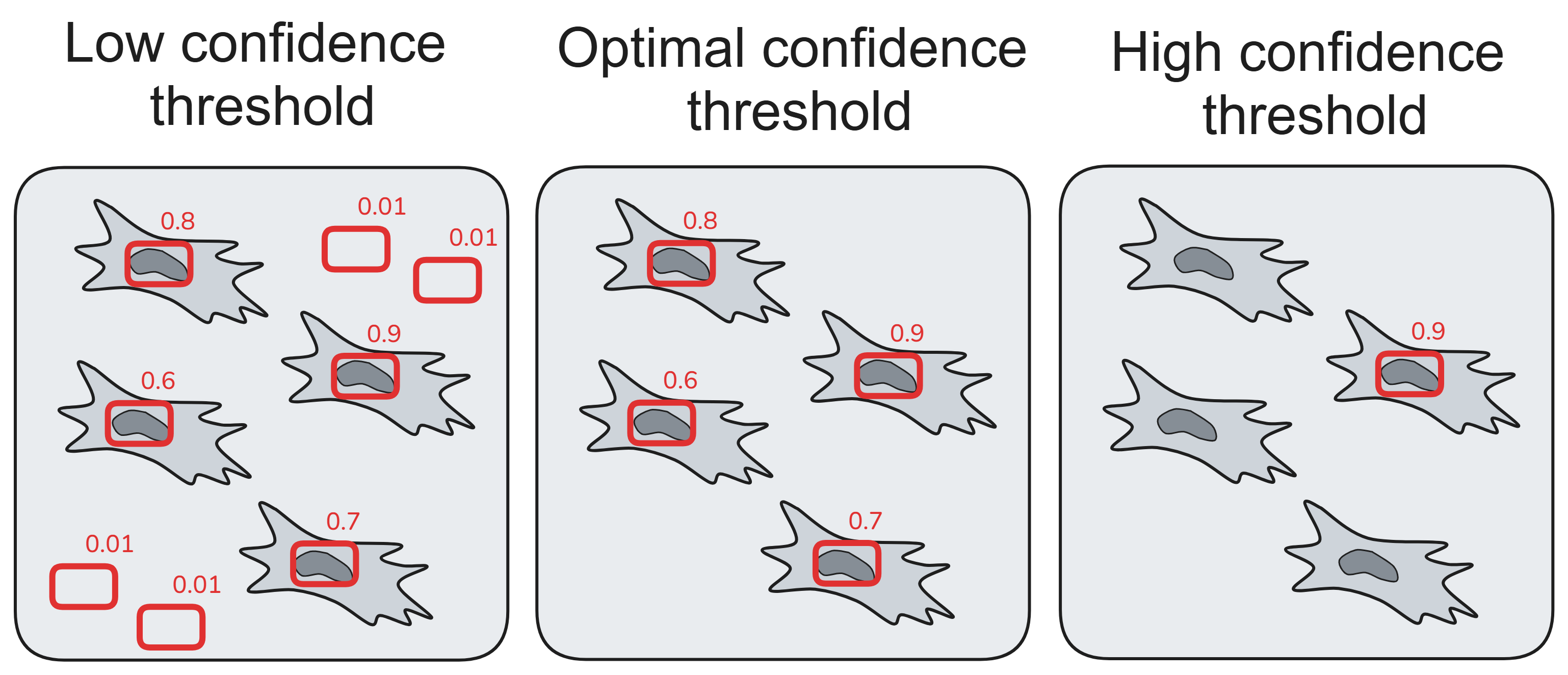
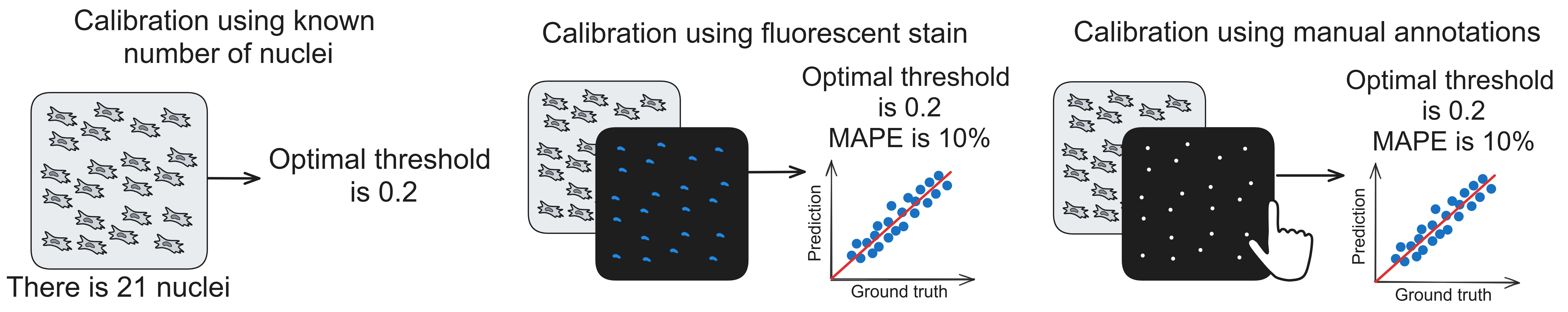
We created several calibration (finding optimal confidence threshold) algorithms that allow adjusting models to specific use cases (cell types, magnifications, illumination settings, cameras etc.):
- Calibration using known number of objects on an image. Doesn't produce accuracy metrics.
- Calibration using fluorescent nuclei stain image (for example, DAPI image). Produces accuracy metrics.
- Calibration using manual annotation of nuclei. Produces accuracy metrics.
Apart from optimal confidence threshold search, these algorithms return accuracy metrics for specific use cases. Given that the calibration image is large, only part of it is used for search of threshold, while the second part is used for evaluation model's accuracy. Accuracy metrics are Mean Absolute Percentage Error (MAPE) and prediction-ground truth scatterplot, which shows how well model performs with different densities of cells.
NEW! If default NuclePhaser models aren't accurate enough on your specific use case, you can finetune them for free with cloud GPU using Google Colab NuclePhaser finetuning notebook. It is specifically designed for users without coding experience.
Learn more about calibration in documentation.
Cell Proliferation Measurement & Population Growth Analysis
With NuclePhaser you can reconstruct population growth curves from timelapse images of growing cell population by counting number of nuclei on each image. Key features of this approach are:
- No special equipment, reagents or dyes required, only regular culture plastic and cell growth medium, microscope with mechanical stage and a PC (even without GPU).
- Accuracy measurement for each specific use case, so you will be sure the tool is working with appropriate precision.
- Measuring the number of cells, not the area occupied by cells, which can be significantly influenced by spreading/narrowing of cells.
- Complete reproducibility of results with metadata.txt files saved for each experiment.

For more detailed information about how NuclePhaser can be used for cell proliferation measurement & population growth analysis, visit our documentation.
Individual cell tracking
NuclePhaser can be used as an assistant for individual cells tracking. This task is extremely difficult, and manual tracking is still the only method with 100% proof against false tracks. With NuclePhaser, you can significantly simplify manual tracking: instead of marking each cell on each image, you can predict nuclei location with NuclePhaser and then correct the result, which is much faster. Learn more in documentation.
Models
Currently only YOLOv5n, YOLOv5s, YOLOv11n and YOLOv11s models, as well as fluorescent nuclei detector YOLOv5n are downloaded automatically with pip install napari-nuclephaser. To use larger models, download them with these links:
Fluorescent nuclei detectors
| Model | Link |
|---|---|
| Fluorescence_v5n | Donwload |
| Fluorescence_v5s | Donwload |
| Fluorescence_v5m | Donwload |
| Fluorescence_v5l | Donwload |
| Fluorescence_v5x | Donwload |
| Fluorescence_v11n | Donwload |
| Fluorescence_v11s | Donwload |
| Fluorescence_v11m | Donwload |
| Fluorescence_v11l | Donwload |
| Fluorescence_v11x | Donwload |
Brighfield nuclei detectors
| Model | Link |
|---|---|
| Brightfield_v5n | Donwload |
| Brightfield_v5s | Donwload |
| Brightfield_v5m | Donwload |
| Brightfield_v5l | Donwload |
| Brightfield_v5x | Donwload |
| Brightfield_v11n | Donwload |
| Brightfield_v11s | Donwload |
| Brightfield_v11m | Donwload |
| Brightfield_v11l | Donwload |
| Brightfield_v11x | Donwload |
[!NOTE] Feel free to use the models published here without the plugin!
Plugin functionality
napari-nuclephaser plugin offers following widgets:
- Widget for inference on single image. Result can be in the form of points or boxes with or without confidence scores. Automatically returns number of cells in the name of result layer.
- Widget for inference on stack of images. Optionally can create .csv or .xlsx file at given location with counting results.
- Widget for calibration using known number of cells.
- Widget for calibration using fluorescent nuclei image (fluorescent nuclei detection model is used as a perfect predictor).
- Widget for calibration using manual annotations.
- Widget for transforming Napari Points layer into Labels layer, which allows turning detection in tracking algorithms-digestible form (in particular, btrack).
- Widget for counting number of points in Points layer.
Learn more about widgets and their functionality at documentation.
Citation
If you use NuclePhaser in your work, please cite our preprint:
@article {Voloshin2025.05.13.653705,
author = {Voloshin, Nikita and Putlyaev, Egor and Chechekhina, Elizaveta and Usachev, Vladimir and Karagyaur, Maxim and Bozov, Kirill and Grigorieva, Olga and Tyurin-Kuzmin, Pyotr and Kulebyakin, Konstantin},
title = {NuclePhaser: a YOLO-based framework for cell nuclei detection and counting in phase contrast images of arbitrary size with support of fast calibration and testing on specific use cases},
year = {2025},
doi = {10.1101/2025.05.13.653705},
URL = {https://www.biorxiv.org/content/early/2025/05/16/2025.05.13.653705},
eprint = {https://www.biorxiv.org/content/early/2025/05/16/2025.05.13.653705.full.pdf},
journal = {bioRxiv}
}
Installation
For detailed installation instructions, visit our documentation.
Contributing
Contributions are very welcome. Tests can be run with tox, please ensure the coverage at least stays the same before you submit a pull request.
License
Distributed under the terms of the MIT license, "napari-nuclephaser" is free and open source software
Issues
If you encounter any problems, please file an issue along with a detailed description.
This napari plugin was generated with copier using the napari-plugin-template.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_nuclephaser-0.2.4.tar.gz.
File metadata
- Download URL: napari_nuclephaser-0.2.4.tar.gz
- Upload date:
- Size: 42.5 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5d11aeabe432c751a803c9eb82de6f1754c21c80c92e0ced4bca8330fd12c71a
|
|
| MD5 |
dce799c02527fe506b83bff5cad35674
|
|
| BLAKE2b-256 |
fb781055564086b311543972882bfc118540309f72603738d98e82e3d032ce35
|
File details
Details for the file napari_nuclephaser-0.2.4-py3-none-any.whl.
File metadata
- Download URL: napari_nuclephaser-0.2.4-py3-none-any.whl
- Upload date:
- Size: 42.5 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
31e43ebd6731dc089f82b9bf49b37717e397afd3a7d1e52e8d8baa453ffe7ccc
|
|
| MD5 |
e2ac7f8c3839ef10f33642b1ebf9ee25
|
|
| BLAKE2b-256 |
0bb08b41de8878f170211458237ec9003c32e57772ba171ac8e394f330d8c6f9
|


















