Use point and shape layer to edit swc format in napari
Project description
napari-swc-editor
Use point and shape layer to edit swc format in napari.
This napari plugin was generated with copier using the napari-plugin-template.
Features
https://github.com/user-attachments/assets/cba1820f-d0b5-436c-a981-62bae0e1a6ba
IO
READER
- Your .swc should follow the following specs: http://www.neuronland.org/NLMorphologyConverter/MorphologyFormats/SWC/Spec.html
- the reader will create 2 napari layer:
point_layerandshape_layer. Onlypoint_layeris interactive,shape_layeris used to render path between swc points. - The raw swc can be accessed in the point layer metadata. Such as
point_layer.metadata["raw_swc"] - A
pd.DataFrameobject is also saved in the metadata:point_layer.metadata["swc_data"]
WRITER
- With the
point_layerselected, you can use napari interface to save with.swcextension name. - You can also do it in command line:
napari.save_layers('test.swc', [point_layer])
Napari Interface
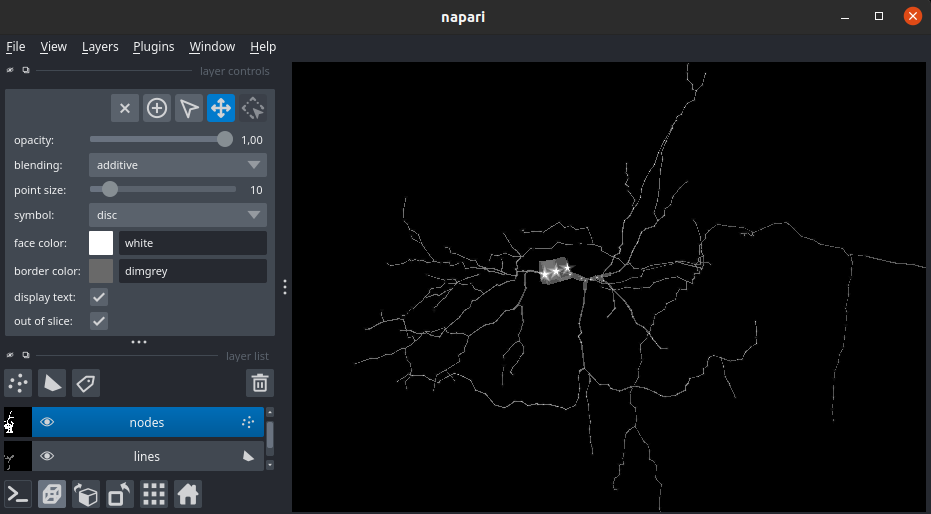
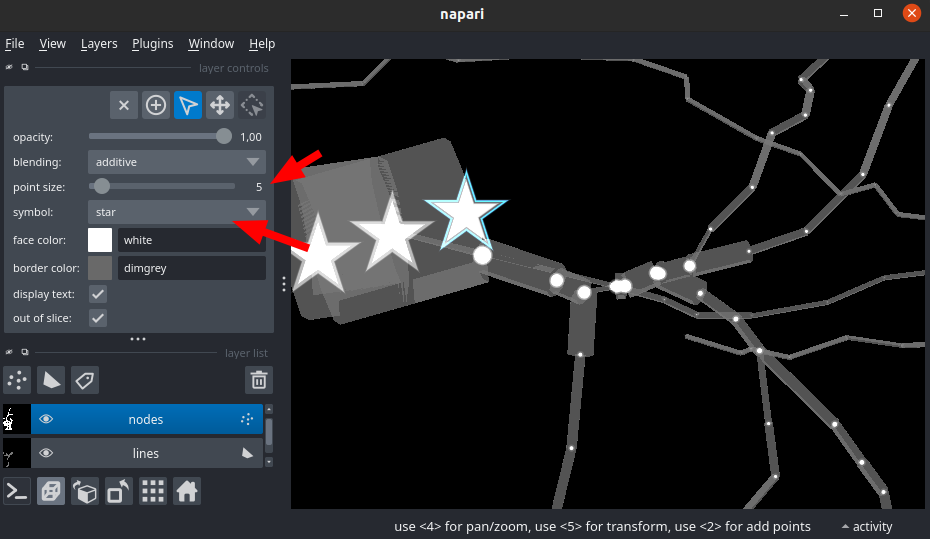
Structure ID and point symbol
In swc, structure id allow to label the type of neuron structure the point belongs to. In this plugin by default, the points will follow this symbol mapping:
SWC_SYMBOL = {
0: "clobber", # undefined
1: "star", # soma
2: "disc", # axon
3: "triangle_down", # basal dendrite
4: "triangle_up", # apical dendrite
}
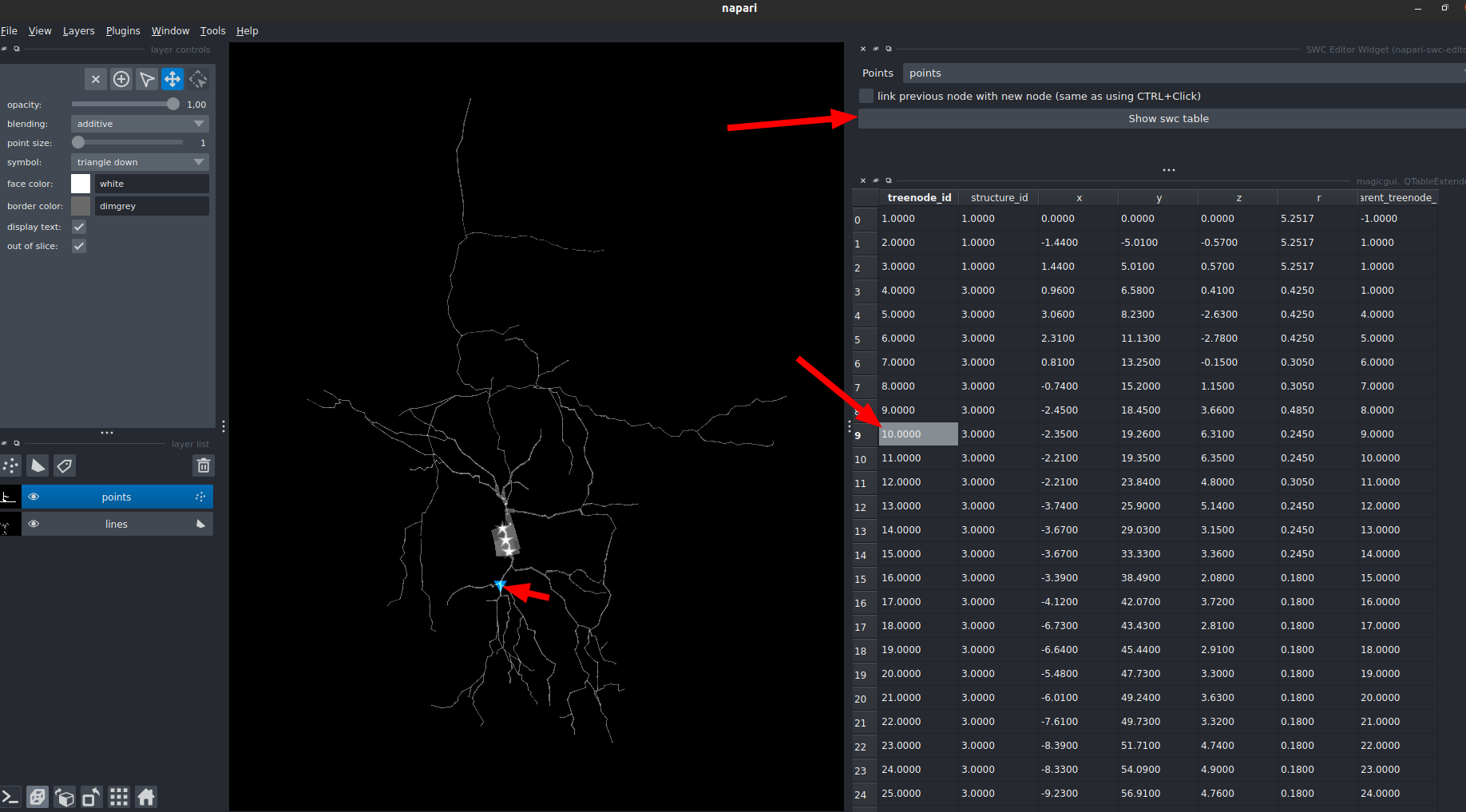
You can also visualize the swc data in a table using the widget under Plugin > SWC Editor Widget
When using the "Show swc table" you will have an interactive table widget:
- left-click on table: highlight + center on the corresponding point
- double-left-click on table: highlight + center on the correspongind point + zoom
- selection on the point layer: highlight the corresponding row on the table
SWC Edition
ALL INTERACTIONS ARE ONLY BOUND TO THE point_layer
THERE IS NO CTRL-Z (please save your progress)
- Add point: You can edit the "r" and the "structure_id" using the
point_sizeandsymbol - Remove point: (Select the point and press
1orsupprordelete) All the link pointing to this point will be removed - Add edge: Select 2 or more point(s) and press on your keyboard
l(aka: link). - Remove edge: Select 1 or more point(s) and press on your keyboard
u(aka: unlink).
If you want to link point as you are adding them you have two solutions:
- press "CTRL" while you add points, this will create a link with the previously selected point
- use the
Plugin > SWC Editor WidgetCheckbox ("link previous node with new node (same as using CTRL+Click)"): when selected, all new points will be selected with the previously selected point
https://github.com/user-attachments/assets/273f1221-2882-4a7c-ab7f-6d3ecb7f3fa6
Installation
You can install napari-swc-editor via pip:
pip install napari-swc-editor
To install latest development version :
pip install git+https://github.com/LaboratoryOpticsBiosciences/napari-swc-editor.git
Contributing
Contributions are very welcome. Tests can be run with tox, please ensure the coverage at least stays the same before you submit a pull request.
License
Distributed under the terms of the BSD-3 license, "napari-swc-editor" is free and open source software
Issues
If you encounter any problems, please file an issue along with a detailed description.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_swc_editor-0.0.5.tar.gz.
File metadata
- Download URL: napari_swc_editor-0.0.5.tar.gz
- Upload date:
- Size: 492.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a8ff09121f5e5e69245f734caa5c4d1ade9e2d3a70fdaa6b439aeb7430255eb5
|
|
| MD5 |
ed48637e061b8b7df9683bdab8df4b8f
|
|
| BLAKE2b-256 |
2013cd2eb51ef27dacaa730ad30eceb4440266656e5df430a2499e5c5a795dca
|
File details
Details for the file napari_swc_editor-0.0.5-py3-none-any.whl.
File metadata
- Download URL: napari_swc_editor-0.0.5-py3-none-any.whl
- Upload date:
- Size: 19.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
707701833678d8a3d397a42dfff9250ffbc13c20811616b98a07480ee3bac966
|
|
| MD5 |
9c08c3211d56f59c1d84a76d01588b53
|
|
| BLAKE2b-256 |
fa2c889ef646fcf182a31f59542110706d2a689cd5f3c5aa6f6fe00a070ac8cd
|