A tool for quick and easy preprocessing and visualization of fNIRS data
Project description
neuropipeline
neuropipeline is a tool for quick and easy to use preprocessing and visualization of Functional Near-Infrared Spectroscopy (fNIRS) data.
Usage
from neuropipeline.fnirs import fNIRS
import neuropipeline.visualizer as nplv
fnirs = fNIRS("path/to/your_file.snirf")
fnirs.preprocess(optical_density=True,
hemoglobin_concentration=True,
motion_correction=True,
temporal_filtering=True,
detrending=True,
normalization=False
)
nplv.open(fnirs)
snirf.write_snirf("path/to/your_new_file.snirf") # WARNING: Be cautious not to overwrite any data you want to keep.
Analysis Example: Heel Stimulation
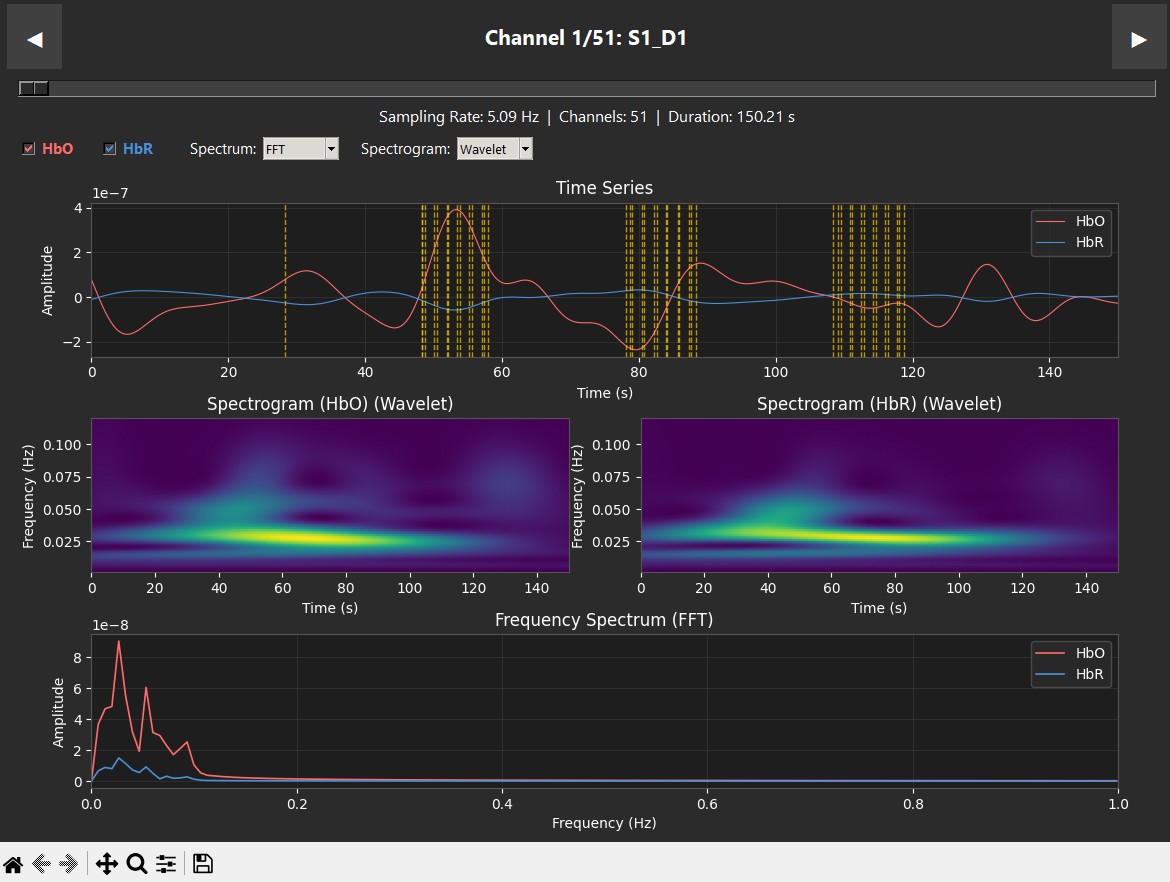
These plots display data from a single subject during a robotic heel-stimulation experiment, showing the Time Series, Spectrogram, and Frequency (PSD/FFT) for two different scenarios. The vertical red dashed lines indicate "markers," which show exactly when a task started or when the robot moved. In this experiment, a robot stimulated the heel 6 times. In the Supination case (left), we see a clear success: oxygenated hemoglobin (HbO) increases right when the stimuli begin. This is supported by the spectrogram, where we see "spikes" of activity at 0.025 Hz (the neurogenic band) that align perfectly with the robot's movements. This confirms the pipeline has successfully captured brain activity in the sensory cortex. In contrast, the Pronation case (right) shows consistently low activity in the spectrogram, and while the time series has some small peaks, they do not show the same clear correlation with the stimulation.
 |
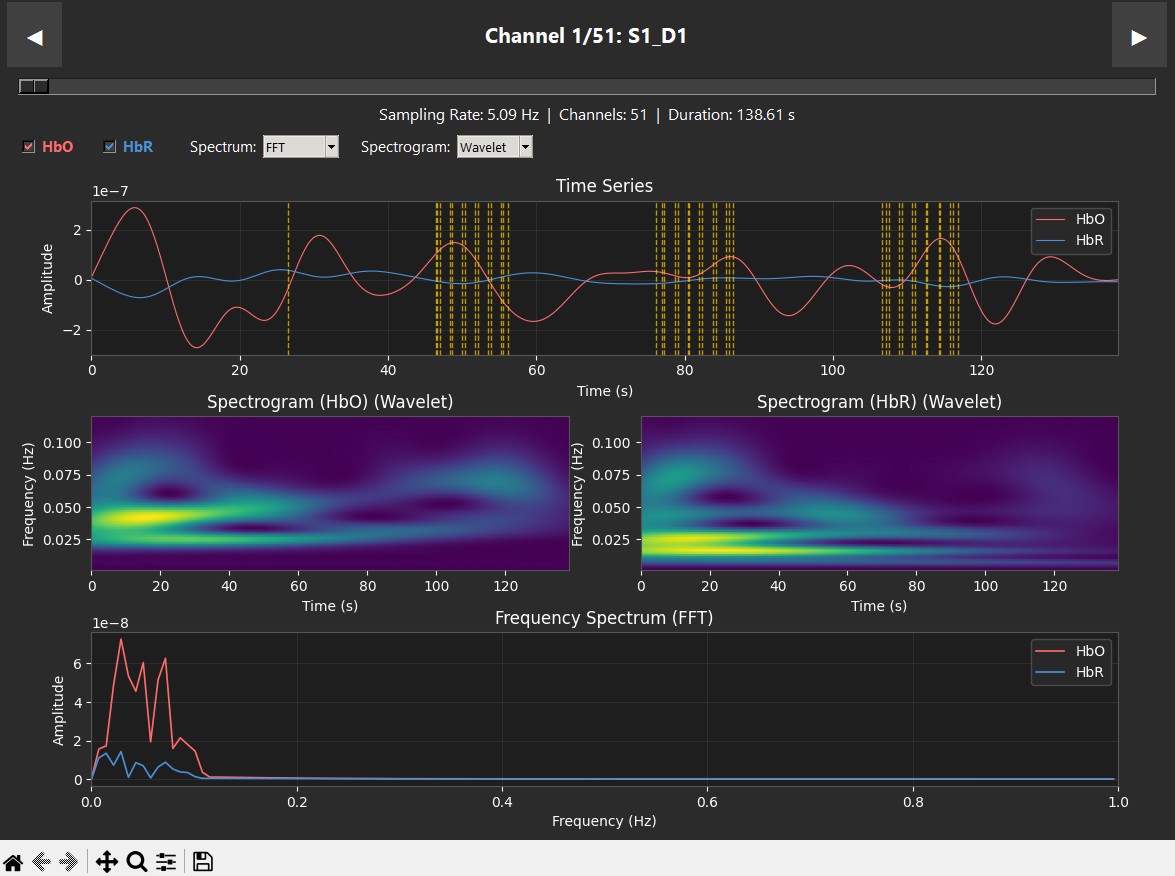 |
| Supination | Pronation |
Installation
python -m pip install neuropipeline
Advanced Usage
from neuropipeline import fNIRS, fNIRSPreprocessor
from neuropipeline.fnirs import visualizer as nplvf
from neuropipeline.eeg import visualizer as nplve
fnirs = fNIRS("path/to/your_file.snirf")
# Advanced Preprocessing Configuration
pp = fNIRSPreprocessor(fnirs) # Create preprocesssor
pp.set_optical_density(True) # Configure
pp.set_hemoglobin_concentration(True)
pp.set_motion_correction(True)
pp.set_temporal_filtering(True, lowcut=0.01, highcut=0.2, order=15)
pp.set_detrending(True)
pp.set_normalization(False)
pp.print() # Inspect the settings
fnirs.preprocess(pp) # Pass the preprocesser only
fnirs.write_snirf("path/to/your_new_file.snirf") # WARNING : Dont overwrite data you want to keep
nplvf.set_spectrogram_limits(0.0, 0.2) # The spectrogram will show frequencies from 0 to 0.2 Hz
nplvf.set_marker_dictionary({2:"Rest", # Display as text rather than indices
3:"Stimuli A",
4:"Stimuli B"})
nplvf.set_spectrum_mode("FFT") # What type of spectrum to show: "FFT" or "PSD"
# NOTE : The wavelet method is computationally intensive
# Try "STFT" first, then "Wavelet" if needed
nplvf.set_spectrogram_method("Wavelet")
nplvf.open(fnirs)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file neuropipeline-0.2.0.tar.gz.
File metadata
- Download URL: neuropipeline-0.2.0.tar.gz
- Upload date:
- Size: 27.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2aecc11e684d2a9ecebfd5d15ac58579120fa5841fda3149f1f35b1ff1b00362
|
|
| MD5 |
f07415c6d1d8a39d2d1b0ecda5204596
|
|
| BLAKE2b-256 |
cb18d87056888e536877e0383ce284f5fb3adc38db6a1230c23a91d4f559418e
|
File details
Details for the file neuropipeline-0.2.0-py3-none-any.whl.
File metadata
- Download URL: neuropipeline-0.2.0-py3-none-any.whl
- Upload date:
- Size: 28.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9efca4508a2873804ecc7f7ee598bec95070be42d7321cd09a211d4aa4df6e18
|
|
| MD5 |
c817c4e27775337fe68681b86a0bf5e3
|
|
| BLAKE2b-256 |
ece68dd39d9b2b51599624c5594d7779b1b0daac7a5b453519d1c5be6fb46dfd
|











