Generate bacterial nomenclature Latin names using a combinatorial approach, as decribed in Pallen et al. (2021) 'The Next Million Names for Archaea and Bacteria'
Project description

GAN: The Great Automatic Nomenclator
The Next Million Names for Archaea and Bacteria, and the nomenclator Python package
Principle
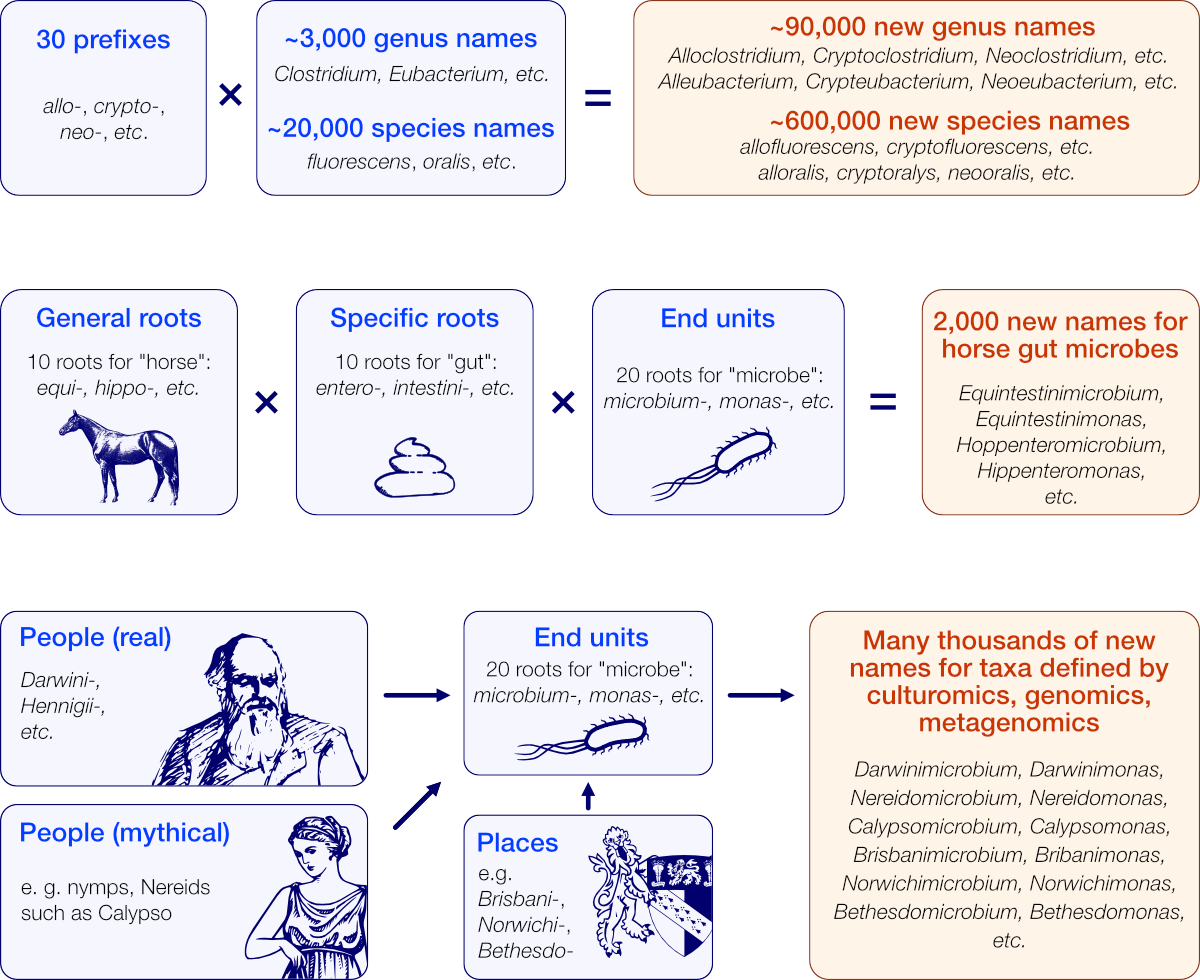
To generate a large number of new names, we apply a combinatorial approach starting with two or three sets of curated roots, that are processed to produce all their possible combinations while keeping trace of their grammatical metadata to draft a valid etymology.
Installation
GAN is available on PyPI as gan-nomenclature and installs with Python 3.8+:
pip install gan-nomenclature
This command installs the library together with its dependencies (pandas, openpyxl, ...).
To work in an isolated environment, you can create one with conda and then install the package from PyPI:
conda create -c conda-forge -n gan python=3.10 pandas pip ipython
conda activate gan
pip install gan-nomenclature
Command-line tools
Installing the package provides a small suite of CLI helpers:
gan-genus: generate JSON/HTML/LaTeX outputs from two or three curated root tables.gan-validate: validate the input Excel files for correct format and content.gan-init: scaffold Excel templates (optionally populated with example rows) for use withgan-genus.gan-aidraft: generate draft etymologies using OpenRouter-hosted LLMs starting from a text file used as context (e.g. a draft of a paper describing the biome where the new taxa were isolated).xls2tsv: convert each worksheet of a workbook into a separate TSV file.tsv2xls: convert TSV files back into Excel format.
Each command offers --help for additional options and usage examples.
Genera generator
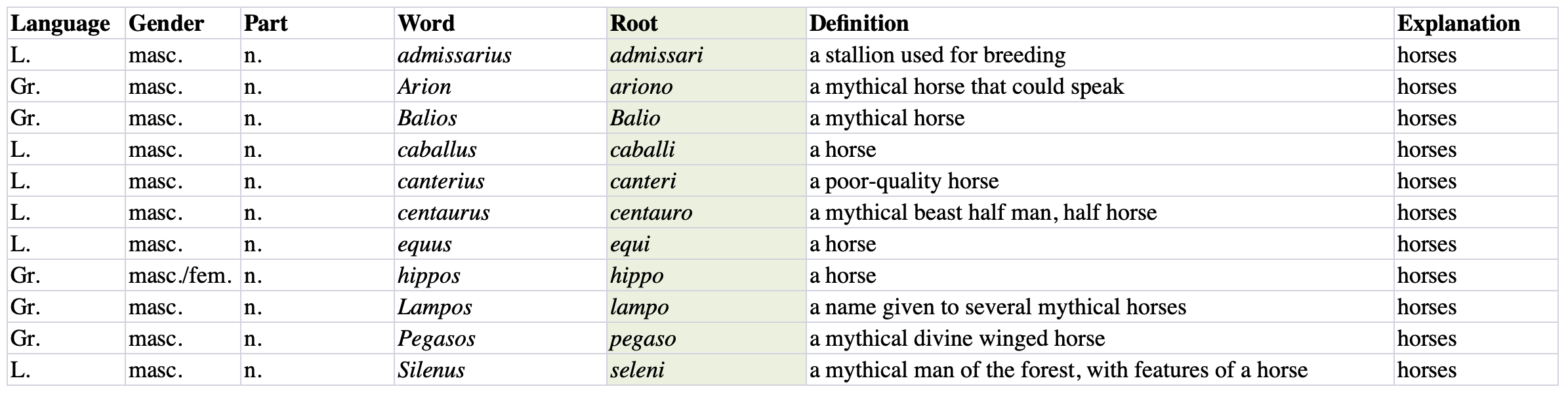
A set of two (or three) Excel tables formatted as shown below is used to generate the list of combinations in JSON, HTML and LaTeX format.
Synopsis:
usage: gan-genus [-h] -1 FIRST -2 SECOND [-3 THIRD] -o OUTDIR [-p PREFIX] [-c CONNECTOR] [-v]
For full usage and installation instructions, please check the documentation.
Example output
Using three small files in the input_test directory (8, 11 and 8 words, respectively), GAN produced 968 (8 x 11 x 8)combinations:
- in PDF format
- in HTML format
Etymology
"The great automatic nomenclaturer" is a reference to a short story ("The Great Automatic Grammatizator") written by the British author Roald Dahl [link].
Citation
Mark J. Pallen et al. The Next Million Names for Archaea and Bacteria, Trends in Microbiology (2020). DOI: 10.1016/j.tim.2020.10.009
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file nomenclator-1.2.0.tar.gz.
File metadata
- Download URL: nomenclator-1.2.0.tar.gz
- Upload date:
- Size: 64.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4fe7677f734114062e21ab2c83e3d88664800109e351a99ad4768c75fa9992a7
|
|
| MD5 |
6792f43a0209ffab4d4d99fa60b63e51
|
|
| BLAKE2b-256 |
9390a05a48ad49285947a6c197e08b3af25a7af2e7980625c15295cf0f237ea8
|
Provenance
The following attestation bundles were made for nomenclator-1.2.0.tar.gz:
Publisher:
pypi-release.yml on telatin/gan
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
nomenclator-1.2.0.tar.gz -
Subject digest:
4fe7677f734114062e21ab2c83e3d88664800109e351a99ad4768c75fa9992a7 - Sigstore transparency entry: 649177655
- Sigstore integration time:
-
Permalink:
telatin/gan@33691b0ee46ca997363d155939c835527e546640 -
Branch / Tag:
refs/tags/v1.2.0 - Owner: https://github.com/telatin
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi-release.yml@33691b0ee46ca997363d155939c835527e546640 -
Trigger Event:
release
-
Statement type:
File details
Details for the file nomenclator-1.2.0-py3-none-any.whl.
File metadata
- Download URL: nomenclator-1.2.0-py3-none-any.whl
- Upload date:
- Size: 48.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8c56086e680f58aa7cd3d8ded2afdc8be274a199aa160143891d823e9ec2c9fc
|
|
| MD5 |
e4ce8bfda4b868193673816bacdd5027
|
|
| BLAKE2b-256 |
10c83e6be5e73a9cfea5aeb8d2a11bc5a959e27519da6f4b2ca5e1c3d0537dfb
|
Provenance
The following attestation bundles were made for nomenclator-1.2.0-py3-none-any.whl:
Publisher:
pypi-release.yml on telatin/gan
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
nomenclator-1.2.0-py3-none-any.whl -
Subject digest:
8c56086e680f58aa7cd3d8ded2afdc8be274a199aa160143891d823e9ec2c9fc - Sigstore transparency entry: 649177697
- Sigstore integration time:
-
Permalink:
telatin/gan@33691b0ee46ca997363d155939c835527e546640 -
Branch / Tag:
refs/tags/v1.2.0 - Owner: https://github.com/telatin
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
pypi-release.yml@33691b0ee46ca997363d155939c835527e546640 -
Trigger Event:
release
-
Statement type: