pca is a python package that performs the principal component analysis and makes insightful plots.
Project description
pca
Star it if you like it!
pca is a python package to perform Principal Component Analysis and to create insightful plots. The core of PCA is build on sklearn functionality to find maximum compatibility when combining with other packages.
But this package can do a lot more. Besides the regular pca, it can also perform SparsePCA, and TruncatedSVD. Depending on your input data, the best approach will be choosen.
Other functionalities are:
- Biplot to plot the loadings
- Determine the explained variance
- Extract the best performing features
- Scatter plot with the loadings
- Outlier detection using Hotelling T2 and/or SPE/Dmodx
This notebook will show some examples.
Contents
Installation
- Install pca from PyPI (recommended). pca is compatible with Python 3.6+ and runs on Linux, MacOS X and Windows.
- It is distributed under the MIT license.
Requirements
- Creation of a new environment is not required but if you wish to do it:
conda create -n env_pca python=3.6
conda activate env_pca
pip install numpy matplotlib sklearn
Installation
pip install pca
- Install the latest version from the GitHub source:
git clone https://github.com/erdogant/pca.git
cd pca
python setup.py install
Import pca package
from pca import pca
Load example data
import numpy as np
from sklearn.datasets import load_iris
# Load dataset
X = pd.DataFrame(data=load_iris().data, columns=load_iris().feature_names, index=load_iris().target)
# Load pca
from pca import pca
# Initialize to reduce the data up to the nubmer of componentes that explains 95% of the variance.
model = pca(n_components=0.95)
# Reduce the data towards 3 PCs
model = pca(n_components=3)
# Fit transform
results = model.fit_transform(X)
X looks like this:
X=array([[5.1, 3.5, 1.4, 0.2],
[4.9, 3. , 1.4, 0.2],
[4.7, 3.2, 1.3, 0.2],
[4.6, 3.1, 1.5, 0.2],
...
[5. , 3.6, 1.4, 0.2],
[5.4, 3.9, 1.7, 0.4],
[4.6, 3.4, 1.4, 0.3],
[5. , 3.4, 1.5, 0.2],
labx=[0, 0, 0, 0,...,2, 2, 2, 2, 2]
label=['label1','label2','label3','label4']
Make scatter plot
fig, ax = model.scatter()

Make biplot
fig, ax = model.biplot(n_feat=4)

Make plot
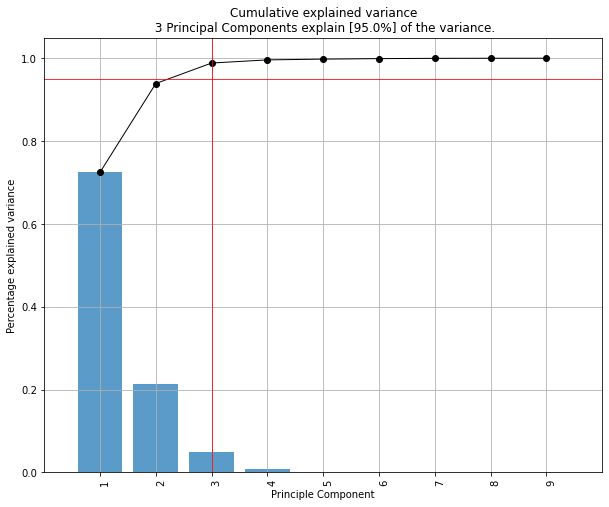
fig, ax = model.plot()

Make 3d plots
fig, ax = model.scatter3d()
fig, ax = model.biplot3d(n_feat=2)

Set alpha transparency
fig, ax = model.scatter(alpha_transparency=1)

PCA normalization.
Normalizing out the 1st and more components from the data. This is usefull if the data is seperated in its first component(s) by unwanted or biased variance. Such as sex or experiment location etc.
print(X.shape)
(150, 4)
# Normalize out 1st component and return data
model = pca()
Xnew = model.norm(X, pcexclude=[1])
print(Xnorm.shape)
(150, 4)
# In this case, PC1 is "removed" and the PC2 has become PC1 etc
ax = pca.biplot(model)
Example to extract the feature importance:
# Import libraries
import numpy as np
import pandas as pd
from pca import pca
# Lets create a dataset with features that have decreasing variance.
# We want to extract feature f1 as most important, followed by f2 etc
f1=np.random.randint(0,100,250)
f2=np.random.randint(0,50,250)
f3=np.random.randint(0,25,250)
f4=np.random.randint(0,10,250)
f5=np.random.randint(0,5,250)
f6=np.random.randint(0,4,250)
f7=np.random.randint(0,3,250)
f8=np.random.randint(0,2,250)
f9=np.random.randint(0,1,250)
# Combine into dataframe
X = np.c_[f1,f2,f3,f4,f5,f6,f7,f8,f9]
X = pd.DataFrame(data=X, columns=['f1','f2','f3','f4','f5','f6','f7','f8','f9'])
# Initialize
model = pca()
# Fit transform
out = model.fit_transform(X)
# Print the top features. The results show that f1 is best, followed by f2 etc
print(out['topfeat'])
# PC feature
# 0 PC1 f1
# 1 PC2 f2
# 2 PC3 f3
# 3 PC4 f4
# 4 PC5 f5
# 5 PC6 f6
# 6 PC7 f7
# 7 PC8 f8
# 8 PC9 f9
Make the plots
model.plot()
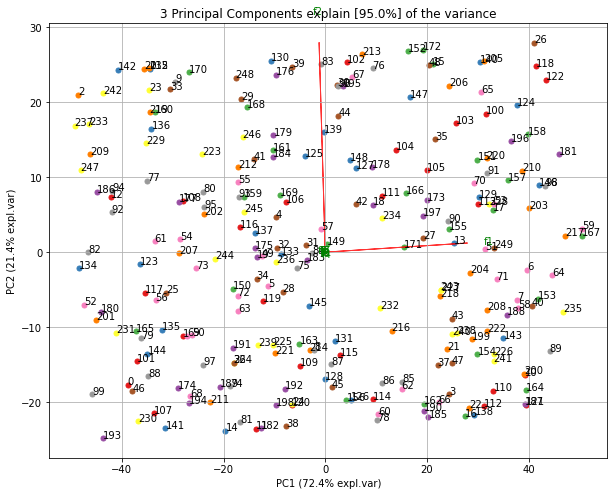
Make the biplot. It can be nicely seen that the first feature with most variance (f1), is almost horizontal in the plot, whereas the second most variance (f2) is almost vertical. This is expected because most of the variance is in f1, followed by f2 etc.
ax = model.biplot(n_feat=10, legend=False)
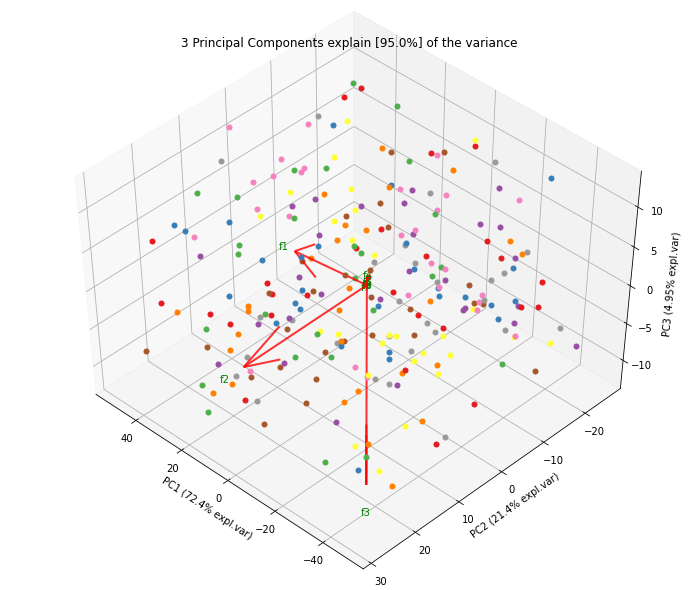
Biplot in 3d. Here we see the nice addition of the expected f3 in the plot in the z-direction.
ax = model.biplot3d(n_feat=10, legend=False)
Example to detect and plot outliers.
To detect any outliers across the multi-dimensional space of PCA, the hotellings T2 test is incorporated. This basically means that we compute the chi-square tests across the top n_components (default is PC1 to PC5). It is expected that the highest variance (and thus the outliers) will be seen in the first few components because of the nature of PCA. Going deeper into PC space may therefore not required but the depth is optional. This approach results in a P-value matrix (samples x PCs) for which the P-values per sample are then combined using fishers method. This approach allows to determine outliers and the ranking of the outliers (strongest tot weak). The alpha parameter determines the detection of outliers (default: 0.05).
from pca import pca
import pandas as pd
import numpy as np
# Create dataset with 100 samples
X = np.array(np.random.normal(0, 1, 500)).reshape(100, 5)
# Create 5 outliers
outliers = np.array(np.random.uniform(5, 10, 25)).reshape(5, 5)
# Combine data
X = np.vstack((X, outliers))
# Initialize model. Alpha is the threshold for the hotellings T2 test to determine outliers in the data.
model = pca(alpha=0.05)
# Fit transform
out = model.fit_transform(X)
# [pca] >The PCA reduction is performed on the [5] columns of the input dataframe.
# [pca] >Column labels are auto-completed.
# [pca] >Row labels are auto-completed.
# [pca] >Fitting using PCA..
# [pca] >Computing loadings and PCs..
# [pca] >Computing explained variance..
# [pca] >Number of components is [4] that covers the [95.00%] explained variance.
# [pca] >Outlier detection using Hotelling T2 test with alpha=[0.05] and n_components=[4]
# [pca] >Outlier detection using SPE/DmodX with n_std=[2]
The information regarding the outliers are stored in the dict 'outliers' (see below). The outliers computed using hotelling T2 test are the columns y_proba, y_score and y_bool. The outliers computed using SPE/DmodX are the columns y_bool_spe, y_score_spe, where y_score_spe is the euclidean distance of the center to the samples. The rows are in line with the input samples.
print(out['outliers'])
# y_proba y_score y_bool y_bool_spe y_score_spe
# 1.0 9.799576e-01 3.060765 False False 0.993407
# 1.0 8.198524e-01 5.945125 False False 2.331705
# 1.0 9.793117e-01 3.086609 False False 0.128518
# 1.0 9.743937e-01 3.268052 False False 0.794845
# 1.0 8.333778e-01 5.780220 False False 1.523642
# .. ... ... ... ... ...
# 1.0 6.793085e-11 69.039523 True True 14.672828
# 1.0 2.610920e-291 1384.158189 True True 16.566568
# 1.0 6.866703e-11 69.015237 True True 14.936442
# 1.0 1.765139e-292 1389.577522 True True 17.183093
# 1.0 1.351102e-291 1385.483398 True True 17.319038
Make the plot
model.biplot(legend=True, SPE=True, hotellingt2=True)
model.biplot3d(legend=True, SPE=True, hotellingt2=True)
# Create only the scatter plots
model.scatter(legend=True, SPE=True, hotellingt2=True)
model.scatter3d(legend=True, SPE=True, hotellingt2=True)


The outliers can can easily be selected:
# Select the outliers
Xoutliers = X[out['outliers']['y_bool'],:]
# Select the other set
Xnormal = X[~out['outliers']['y_bool'],:]
If desired, the outliers can also be detected directly using the hotelling T2 and/or SPE/DmodX functionality.
import pca
outliers_hot = pca.hotellingsT2(out['PC'].values, alpha=0.05)
outliers_spe = pca.spe_dmodx(out['PC'].values, n_std=2)
Example to only plot the directions (arrows).
from pca import pca
# Initialize
model = pca()
# Example with DataFrame
X = np.array(np.random.normal(0, 1, 500)).reshape(100, 5)
X = pd.DataFrame(data=X, columns=np.arange(0, X.shape[1]).astype(str))
# Fit transform
out = model.fit_transform(X)
# Make plot with parameters: set cmap to None and label and legend to False. Only directions will be plotted.
model.biplot(cmap=None, label=False, legend=False)

Set visible status of figures.
from pca import pca
# Initialize
model = pca()
# Example with DataFrame
X = np.array(np.random.normal(0, 1, 500)).reshape(100, 5)
X = pd.DataFrame(data=X, columns=np.arange(0, X.shape[1]).astype(str))
# Fit transform
out = model.fit_transform(X)
# Make plot with parameters.
fig, ax = model.biplot(visible=False)
# Set the figure again to True and show the figure.
fig.set_visible(True)
fig
Example to Transform unseen datapoints into fitted space.
import matplotlib.pyplot as plt
from sklearn.datasets import load_iris
import pandas as pd
from pca import pca
# Initialize
model = pca(n_components=2, normalize=True)
# Dataset
X = pd.DataFrame(data=load_iris().data, columns=load_iris().feature_names, index=load_iris().target)
# Get some random samples across the classes
idx=[0,1,2,3,4,50,51,52,53,54,55,100,101,102,103,104,105]
X_unseen = X.iloc[idx, :]
# Label original dataset to make sure the check which samples are overlapping
X.index.values[idx]=3
# Fit transform
model.fit_transform(X)
# Transform new "unseen" data. Note that these datapoints are not really unseen as they are readily fitted above.
# But for the sake of example, you can see that these samples will be transformed exactly on top of the orignial ones.
PCnew = model.transform(X_unseen)
# Plot PC space
model.scatter()
# Plot the new "unseen" samples on top of the existing space
plt.scatter(PCnew.iloc[:, 0], PCnew.iloc[:, 1], marker='x')
Citation
Please cite distfit in your publications if this is useful for your research. Here is an example BibTeX entry:
@misc{erdogant2019pca,
title={pca},
author={Erdogan Taskesen},
year={2019},
howpublished={\url{https://github.com/erdogant/pca}},
}
Maintainer
Erdogan Taskesen, github: [erdogant](https://github.com/erdogant)
Contributions are welcome.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pca-1.5.5.tar.gz.
File metadata
- Download URL: pca-1.5.5.tar.gz
- Upload date:
- Size: 28.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.7.0 requests/2.25.1 setuptools/52.0.0.post20210125 requests-toolbelt/0.9.1 tqdm/4.61.2 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7772cc67f6d8b74fb2272869ea7ca52d8371a53e2b51635abcd60949883df111
|
|
| MD5 |
a527aa2f5123407c4ad995aad1c2b09f
|
|
| BLAKE2b-256 |
ea7bdb447427a3c5354a7c07fb0f467784ed46b91e8a7f371607546f89792696
|
File details
Details for the file pca-1.5.5-py3-none-any.whl.
File metadata
- Download URL: pca-1.5.5-py3-none-any.whl
- Upload date:
- Size: 24.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.7.0 requests/2.25.1 setuptools/52.0.0.post20210125 requests-toolbelt/0.9.1 tqdm/4.61.2 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a6ac4aa8ad2ca1d2651842ac08f3b5ec88e694aad82d3e5ea86dbd790dd4ecfe
|
|
| MD5 |
bcd79fb5cf3be8e4d0bc7ebae72e2f68
|
|
| BLAKE2b-256 |
eecf585e701329dbde6a5b6deb826e09edcb46b2602f01290eb441b11bfce0fb
|