Plot whole genome contact heatmap
Project description
PlotHiC
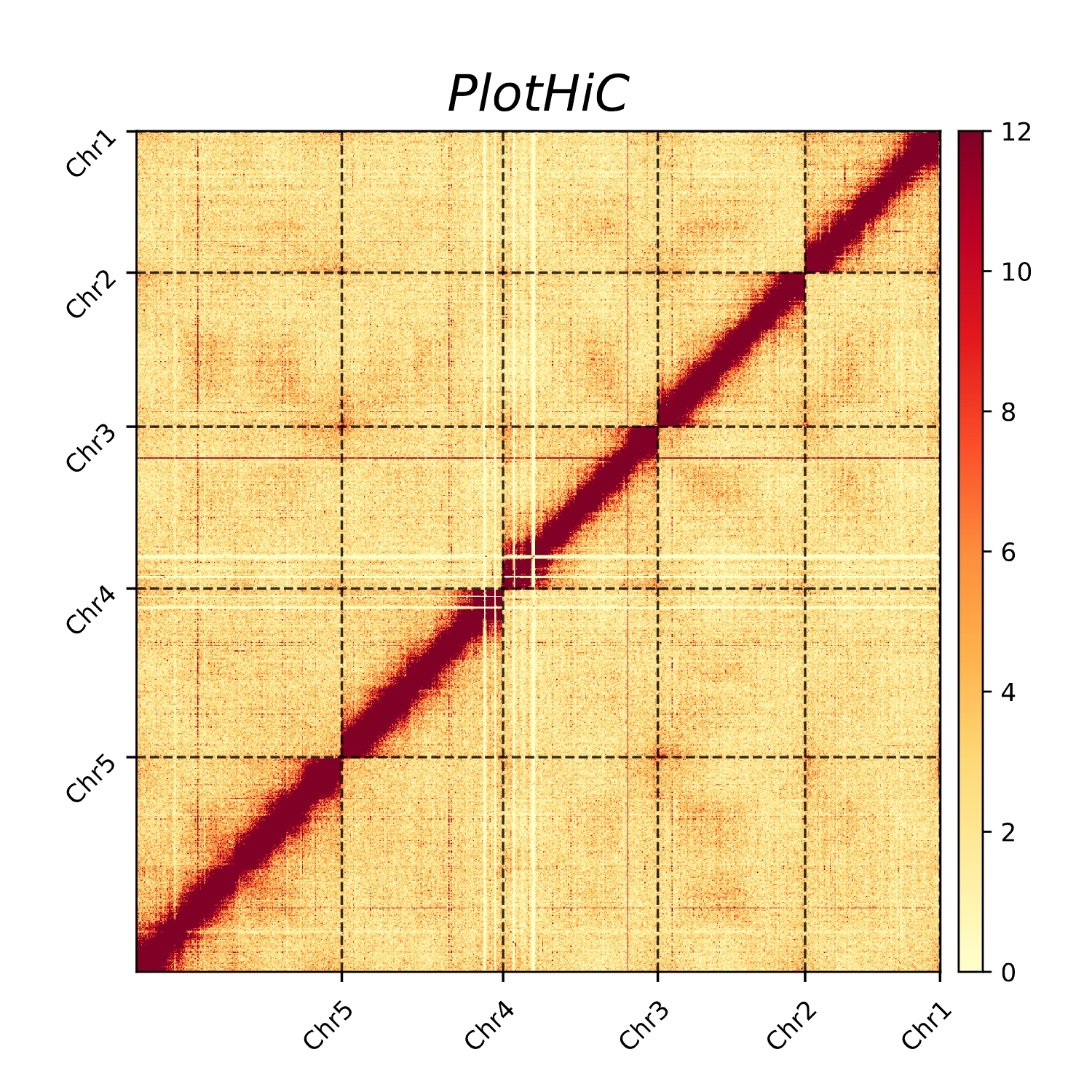
PlotHiC is used to visualize global interaction heatmaps after genome scaffolding.
Note: PlotHiC is currently under development. If you have any questions, please Open Issues or provide us with your comments via the email below.
Author: Zijie Jiang
Email: jzjlab@163.com
Content
Citations
If you used PlotHiC in your research, please cite us:
Zijie Jiang, Zhixiang Peng, Zhaoyuan Wei, Jiahe Sun, Yongjiang Luo, Lingzi Bie, Guoqing Zhang, Yi Wang, A deep learning-based method enables the automatic and accurate assembly of chromosome-level genomes, Nucleic Acids Research, 2024;, gkae789, https://doi.org/10.1093/nar/gkae789
Installation
- Dependency :
python = "^3.10"
pip
# pip install
pip install plothic
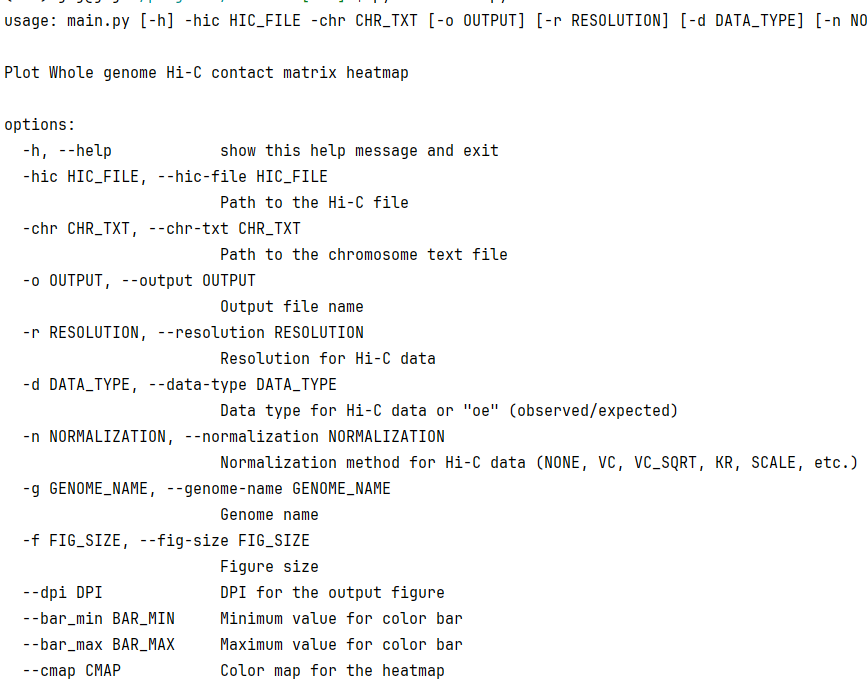
Usage
Input file
genome.hic
This file is taken directly from 3d-dna, you need to select the final hic file (which has already been error adjusted and chromosome boundaries determined).
chr.tx
- This file is used for heatmap labeling. The first column is the name of the chromosome.
- The second column is the length of the chromosome (this length is the length of the hic file in Juicebox and can be manually determined from Juicebox).
- The third column is the order in which the chromosomes are placed, which is used to customize the arrangement of chromosomes (for example, from max to min).
Note: the length is in .hic file, not true base length.
# name length index
Chr1 24800000 5
Chr2 44380000 4
Chr3 63338000 3
Chr4 81187000 2
Chr5 97650000 1
example
- Default order
plothic -hic genome.hic -chr chr.txt -r 100000
# -hic > .hic file
# -chr > chromosome length (in .hic file)
# -r > resolution to visualization
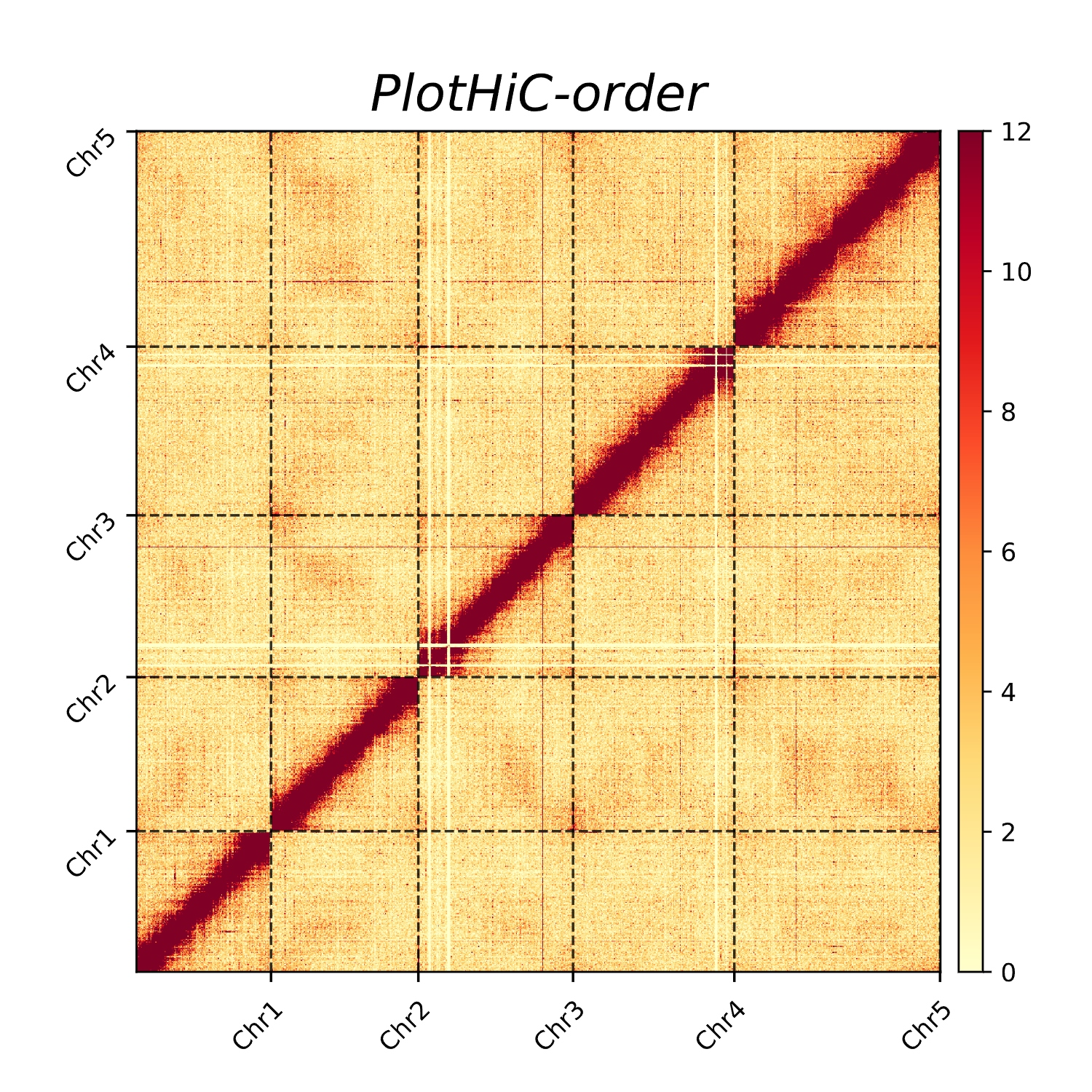
- Custom order
plothic -hic genome.hic -chr chr.txt -r 100000 --order
# -hic > .hic file
# -chr > chromosome length (in .hic file)
# -r > resolution to visualization
# --order >
other parameter
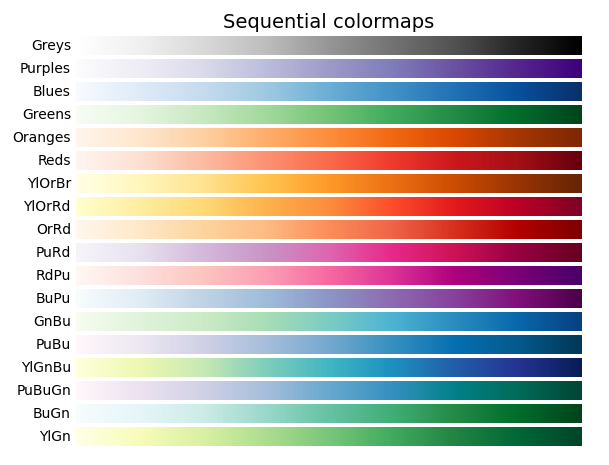
Color map
PlotHiC uses YlOrRd by default, you can choose more colors from Matplotlib.
Dev
Currently only use hic and chr txt are supported for visualization, and assembly files will be supported in the future. However, from the perspective of usage, using chr txt files seems to be more convenient. If you have better suggestions or other requirements, please Open Issues.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file plothic-0.3.1.tar.gz.
File metadata
- Download URL: plothic-0.3.1.tar.gz
- Upload date:
- Size: 20.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.4 CPython/3.10.12 Linux/5.10.16.3-microsoft-standard-WSL2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1a09d21ef224896c2813a3471af3efd1142b282c1137d714427edfc3abd1837e
|
|
| MD5 |
9b9369a1d081d53a1f1eca39900b9cbe
|
|
| BLAKE2b-256 |
c165192e73583c024855c4c1732aa3ebc6ade1280dee024496d0baeefeea0268
|
File details
Details for the file plothic-0.3.1-py3-none-any.whl.
File metadata
- Download URL: plothic-0.3.1-py3-none-any.whl
- Upload date:
- Size: 22.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.4 CPython/3.10.12 Linux/5.10.16.3-microsoft-standard-WSL2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
503917ff2e9677cfbff997ddb87dab48ceba64f91a10171cd74e81818c232a48
|
|
| MD5 |
a0c4eb7b0bdfe5583f7024ef0830115e
|
|
| BLAKE2b-256 |
db46251b5f306fe92195b15bf9709f8312393709e974195c7c327e7e167d0bc6
|