Positive-unlabeled learning with Python
Project description
Positive-unlabeled learning with Python.
Website: https://pulearn.github.io/pulearn/
Documentation: https://pulearn.github.io/pulearn/doc/pulearn/
from pulearn import ElkanotoPuClassifier
from sklearn.svm import SVC
svc = SVC(C=10, kernel='rbf', gamma=0.4, probability=True)
pu_estimator = ElkanotoPuClassifier(estimator=svc, hold_out_ratio=0.2)
pu_estimator.fit(X, y)1 Documentation
This is the repository for the pulearn package. The readme file is aimed at helping contributors to the project.
To learn more about how to use pulearn, either visit pulearn’s homepage or read the documentation at <https://pulearn.github.io/pulearn/doc/pulearn/>`_.
2 Installation
Install pulearn with:
pip install pulearn3 Extending pulearn
New learner work now goes through a small registry and contributor scaffold:
pulearn.get_algorithm_registry() exposes discoverable metadata for the built-in learners.
doc/new_algorithm_checklist.md defines the required contributor steps.
doc/templates/new_algorithm_doc_stub.md is the docs page starting point.
tests/templates/test_new_algorithm_template.py.tmpl and tests/templates/test_api_contract_template.py.tmpl provide regression and shared API contract scaffolds.
benchmarks/templates/benchmark_entry_template.py.tmpl is the benchmark stub until the benchmark harness lands in the dedicated roadmap milestone.
pulearn.get_scaffold_templates() resolves those scaffold files only from a repository checkout and fails clearly when they are unavailable.
At minimum, every new learner should register metadata, add focused tests, run the shared API contract checks when it inherits from BasePUClassifier, and add docs plus a benchmark placeholder in the same PR.
4 Implemented Classifiers
4.1 Elkanoto
Scikit-Learn wrappers for both the methods mentioned in the paper by Elkan and Noto, “Learning classifiers from only positive and unlabeled data” (published in Proceeding of the 14th ACM SIGKDD international conference on Knowledge discovery and data mining, ACM, 2008).
These wrap the Python code from a fork by AdityaAS (with implementation to both methods) to the original repository by Alexandre Drouin implementing one of the methods.
PU labels are normalized to a canonical internal representation (1 = labeled positive, 0 = unlabeled). Accepted input conventions include {1, -1}, {1, 0}, and {True, False}. Use pulearn.normalize_pu_labels(...) to normalize labels immediately at ingest or estimator/metric boundaries.
4.1.1 Classic Elkanoto
To use the classic (unweighted) method, use the ElkanotoPuClassifier class:
from pulearn import ElkanotoPuClassifier
from sklearn.svm import SVC
svc = SVC(C=10, kernel='rbf', gamma=0.4, probability=True)
pu_estimator = ElkanotoPuClassifier(estimator=svc, hold_out_ratio=0.2)
pu_estimator.fit(X, y)4.1.2 Weighted Elkanoto
To use the weighted method, use the WeightedElkanotoPuClassifier class:
from pulearn import WeightedElkanotoPuClassifier
from sklearn.svm import SVC
svc = SVC(C=10, kernel='rbf', gamma=0.4, probability=True)
pu_estimator = WeightedElkanotoPuClassifier(
estimator=svc, labeled=10, unlabeled=20, hold_out_ratio=0.2)
pu_estimator.fit(X, y)See the original paper for details on how the labeled and unlabeled quantities are used to weigh training examples and affect the learning process: https://cseweb.ucsd.edu/~elkan/posonly.pdf.
4.2 Bagging-based PU-learning
Based on the paper A bagging SVM to learn from positive and unlabeled examples (2013) by Mordelet and Vert. The implementation is by Roy Wright (roywright on GitHub), and can be found in his repository.
Accepted PU label conventions match the package-wide contract: 1/True for labeled positives and 0/-1/False for unlabeled examples.
from pulearn import BaggingPuClassifier
from sklearn.svm import SVC
svc = SVC(C=10, kernel='rbf', gamma=0.4, probability=True)
pu_estimator = BaggingPuClassifier(
estimator=svc, n_estimators=15)
pu_estimator.fit(X, y)4.3 Bayesian PU Classifiers
Bayesian classifiers for PU learning based on the MIT-licensed Bayesian Classifiers for PU Learning project by Chengning Zhang.
All four classifiers accept the same package-wide PU label conventions: {1, 0}, {1, -1}, and {True, False}. Continuous features are automatically discretized into equal-width bins.
4.3.1 Positive Naive Bayes (PNB)
The simplest Bayesian PU classifier. Class-conditional distributions \(P(x|y=1)\) and \(P(x|y=0)\) are estimated from the labeled positives and the unlabeled set (treated as approximate negatives) respectively, with Laplace smoothing controlled by alpha.
from pulearn import PositiveNaiveBayesClassifier
clf = PositiveNaiveBayesClassifier(alpha=1.0, n_bins=10)
clf.fit(X_train, y_pu) # y_pu: 1 = labeled positive, 0 = unlabeled
proba = clf.predict_proba(X_test) # shape (n_samples, 2): [P(y=0|x), P(y=1|x)]
labels = clf.predict(X_test)4.3.2 Weighted Naive Bayes (WNB)
Extends PNB by weighting each feature’s log-likelihood contribution by its empirical mutual information with the PU label. Features that are more informative receive higher weights; all weights are non-negative and sum to 1.
from pulearn import WeightedNaiveBayesClassifier
clf = WeightedNaiveBayesClassifier(alpha=1.0, n_bins=10)
clf.fit(X_train, y_pu)
print(clf.feature_weights_) # normalized MI weight per feature (sums to 1)
proba = clf.predict_proba(X_test)4.3.3 Positive Tree-Augmented Naive Bayes (PTAN)
Extends PNB by replacing the naive feature-independence assumption with a tree structure learned via the Chow-Liu algorithm. Pairwise conditional mutual information \(I(X_i; X_j \mid S)\) is computed for all feature pairs, and a maximum spanning tree is built with Prim’s algorithm. Each non-root feature depends on exactly one parent feature in addition to the class label.
from pulearn import PositiveTANClassifier
clf = PositiveTANClassifier(alpha=1.0, n_bins=10)
clf.fit(X_train, y_pu)
print(clf.tan_parents_) # parent index per feature; -1 for the root
proba = clf.predict_proba(X_test)4.3.4 Weighted Tree-Augmented Naive Bayes (WTAN)
Combines PTAN’s tree structure with WNB’s per-feature MI weighting.
from pulearn import WeightedTANClassifier
clf = WeightedTANClassifier(alpha=1.0, n_bins=10)
clf.fit(X_train, y_pu)
print(clf.feature_weights_) # normalized MI weight per feature
print(clf.tan_parents_) # learned tree structure
proba = clf.predict_proba(X_test)A complete end-to-end example on the Wisconsin breast cancer dataset can be found in the examples directory:
python examples/BayesianPULearnersExample.py4.4 Prior Estimation
pulearn.priors now exposes a small, unified class-prior estimation API for SCAR workflows:
LabelFrequencyPriorEstimator returns the observed labeled-positive rate as a naive lower bound for \(\pi\).
HistogramMatchPriorEstimator fits a probabilistic scorer and matches labeled-positive vs. unlabeled score histograms.
ScarEMPriorEstimator runs a soft-label EM refinement loop over latent positives in the unlabeled pool.
All three implement fit(X, y) and estimate(X, y) and return a PriorEstimateResult with the estimated prior, sample counts, and method-specific metadata.
from pulearn import (
HistogramMatchPriorEstimator,
LabelFrequencyPriorEstimator,
ScarEMPriorEstimator,
)
baseline = LabelFrequencyPriorEstimator().estimate(X_train, y_pu)
histogram = HistogramMatchPriorEstimator().estimate(X_train, y_pu)
scar_em = ScarEMPriorEstimator().estimate(X_train, y_pu)
print(baseline.pi, histogram.pi, scar_em.pi)
print(scar_em.metadata["c_estimate"])Bootstrap confidence intervals are available when you need uncertainty estimates or reproducible sensitivity checks:
estimator = ScarEMPriorEstimator().fit(X_train, y_pu)
result = estimator.bootstrap(
X_train,
y_pu,
n_resamples=200,
confidence_level=0.95,
random_state=7,
)
print(result.confidence_interval.lower, result.confidence_interval.upper)Diagnostics helpers can summarize estimator stability across a parameter sweep and optionally drive sensitivity plots:
from pulearn import (
HistogramMatchPriorEstimator,
diagnose_prior_estimator,
)
diagnostics = diagnose_prior_estimator(
HistogramMatchPriorEstimator(),
X_train,
y_pu,
parameter_grid={"n_bins": [8, 12, 20], "smoothing": [0.5, 1.0]},
)
print(diagnostics.unstable, diagnostics.warnings)
print(diagnostics.range_pi, diagnostics.std_pi)
# Optional: requires matplotlib
# from pulearn import plot_prior_sensitivity
# plot_prior_sensitivity(diagnostics)If pi is uncertain, you can sweep corrected metrics across a plausible prior range and inspect best/worst-case behavior:
from pulearn import analyze_prior_sensitivity
sensitivity = analyze_prior_sensitivity(
y_pu,
y_pred=y_pred,
y_score=y_score,
metrics=["pu_precision", "pu_roc_auc"],
pi_min=0.2,
pi_max=0.5,
num=7,
)
print(sensitivity.as_rows())
print(sensitivity.summaries["pu_precision"].best_pi)4.5 Propensity Estimation
pulearn.propensity provides first-class estimators for the SCAR labeling propensity \(c = P(s=1 \mid y=1)\):
MeanPositivePropensityEstimator matches the classic Elkan-Noto mean-on-positives estimate.
TrimmedMeanPropensityEstimator reduces sensitivity to a few badly calibrated labeled positives.
MedianPositivePropensityEstimator and QuantilePositivePropensityEstimator give conservative alternatives when positive scores are noisy or skewed.
CrossValidatedPropensityEstimator uses out-of-fold probabilities from a probabilistic sklearn estimator to reduce optimistic bias.
All score-based estimators implement fit(y_pu, s_proba=...) and estimate(y_pu, s_proba=...). The cross-validated estimator uses the same API but takes X=... plus a base estimator.
from sklearn.linear_model import LogisticRegression
from pulearn import (
CrossValidatedPropensityEstimator,
MeanPositivePropensityEstimator,
MedianPositivePropensityEstimator,
QuantilePositivePropensityEstimator,
TrimmedMeanPropensityEstimator,
)
mean_c = MeanPositivePropensityEstimator().estimate(y_pu, s_proba=y_score)
trimmed_c = TrimmedMeanPropensityEstimator(trim_fraction=0.1).estimate(
y_pu,
s_proba=y_score,
)
median_c = MedianPositivePropensityEstimator().estimate(
y_pu,
s_proba=y_score,
)
quantile_c = QuantilePositivePropensityEstimator(quantile=0.25).estimate(
y_pu,
s_proba=y_score,
)
cv_c = CrossValidatedPropensityEstimator(
estimator=LogisticRegression(max_iter=1000),
cv=5,
random_state=7,
).estimate(y_pu, X=X_train)
print(mean_c.c, trimmed_c.c, median_c.c, quantile_c.c, cv_c.c)
print(cv_c.metadata["fold_estimates"])pulearn.metrics.estimate_label_frequency_c(...) now delegates to the same mean estimator and therefore expects probability-like scores in [0, 1].
Bootstrap confidence intervals are available for propensity estimators when you need uncertainty estimates or a warning that the labeling mechanism looks unstable under resampling:
estimator = TrimmedMeanPropensityEstimator(trim_fraction=0.1).fit(
y_pu,
s_proba=y_score,
)
result = estimator.bootstrap(
y_pu,
s_proba=y_score,
n_resamples=200,
confidence_level=0.95,
random_state=7,
)
print(result.c)
print(result.confidence_interval.lower)
print(result.confidence_interval.upper)
print(result.confidence_interval.warning_flags)Instability warnings flag repeated fit failures, unusually high resample variance, large coefficient of variation, or inconsistent cross-validation fold estimates. Treat those warnings as a signal to inspect calibration, selection bias, or mislabeled positives before using c downstream.
You can also compare labeled positives against the highest-scoring unlabeled pool to look for likely SCAR violations:
from pulearn import scar_sanity_check
scar_check = scar_sanity_check(
y_pu,
s_proba=y_score,
X=X_train,
candidate_quantile=0.9,
random_state=7,
)
print(scar_check.group_membership_auc)
print(scar_check.max_abs_smd)
print(scar_check.warnings)Warnings such as group_separable or max_feature_shift mean the high-scoring unlabeled pool looks materially different from labeled positives, which is a practical sign that SCAR may not hold closely enough for naive propensity correction.
4.6 Experimental SAR Hooks
pulearn now exposes a minimal experimental interface for SAR-style selection models. The scope is intentionally narrow: you can plug in a propensity model and derive inverse-propensity weights, but this does not yet implement full SAR learners or SAR-corrected metrics.
from sklearn.linear_model import LogisticRegression
from pulearn import (
ExperimentalSarHook,
compute_inverse_propensity_weights,
predict_sar_propensity,
)
propensity_model = LogisticRegression(max_iter=1000).fit(X_train, s_train)
sar_scores = predict_sar_propensity(propensity_model, X_test)
sar_weights = compute_inverse_propensity_weights(
sar_scores,
clip_min=0.05,
clip_max=1.0,
normalize=True,
)
hook = ExperimentalSarHook(propensity_model)
hook_result = hook.inverse_propensity_weights(X_test, normalize=True)
print(hook_result.weights[:5])
print(hook_result.metadata["propensity_model"])These helpers emit experimental warnings on purpose. Treat them as plumbing for custom SAR research code rather than a stable high-level workflow. Always inspect the clipped counts and weight magnitudes before using them downstream.
5 Evaluation Metrics
Standard binary classification metrics (precision, recall, F1) are systematically biased in PU settings: a trivial classifier that predicts positive for every sample achieves recall = 1.0 with no penalty for false positives in the unlabeled pool. pulearn.metrics provides unbiased estimators that cover the full evaluation lifecycle under the SCAR (Selected Completely At Random) assumption.
5.1 Calibration
Before computing confusion-matrix metrics you must map the model’s observed output \(P(s=1|x)\) to the calibrated posterior \(P(y=1|x)\).
from pulearn.metrics import estimate_label_frequency_c, calibrate_posterior_p_y1
# 1. Estimate the propensity score c = P(s=1 | y=1)
c_hat = estimate_label_frequency_c(y_pu, s_proba)
# 2. Calibrate: P(y=1|x) ≈ P(s=1|x) / c
p_y1 = calibrate_posterior_p_y1(s_proba, c_hat)estimate_label_frequency_c implements the Elkan-Noto estimator \(\hat{c} \approx \mathbb{E}[P(s=1|x) \mid s=1]\). calibrate_posterior_p_y1 clips \(P(s=1|x) / \hat{c}\) to \([0, 1]\).
5.2 Expected-Confusion Metrics
These metrics reconstruct the confusion matrix from calibrated posteriors rather than treating unlabeled data as confirmed negatives.
from pulearn.metrics import (
pu_recall_score,
pu_precision_score,
pu_f1_score,
pu_specificity_score,
)
# Recall on labeled positives (no class prior needed)
rec = pu_recall_score(y_pu, y_pred)
# Unbiased precision and F1 require the class prior π
prec = pu_precision_score(y_pu, y_pred, pi=0.3)
f1 = pu_f1_score(y_pu, y_pred, pi=0.3)
# Expected specificity — returns 0.0 for any all-positive classifier
spec = pu_specificity_score(y_pu, y_score)pu_specificity_score is particularly useful as a sanity-check guard: a degenerate classifier that assigns every sample to the positive class obtains \(\text{spec} = 0\), immediately flagging the model as trivial.
5.3 Ranking Metrics
Two AUC-based metrics correct for the absence of ground-truth negatives.
from pulearn.metrics import pu_roc_auc_score, pu_average_precision_score
# Sakai (2018) adjustment: AUC_pn = (AUC_pu − 0.5π) / (1 − π)
auc = pu_roc_auc_score(y_pu, y_score, pi=0.3)
# Area Under Lift: AUL = 0.5π + (1 − π) · AUC_pu
aul = pu_average_precision_score(y_pu, y_score, pi=0.3)pu_roc_auc_score maps the biased \(AUC_{pu}\) (computed against PU labels) to an unbiased estimator of the true positive-vs-negative AUC. pu_average_precision_score returns the Area Under Lift (AUL), which is more robust to severe class imbalance.
5.4 Risk Estimators
For flexible models such as deep networks, raw risk estimators are suitable for early stopping and model selection in lieu of black-box accuracy metrics.
from pulearn.metrics import pu_unbiased_risk, pu_non_negative_risk
# uPU: pi * R_p+ + R_u- - pi * R_p- (du Plessis et al., 2015)
risk_upu = pu_unbiased_risk(y_pu, y_score, pi=0.3)
# nnPU: clamps negative component to zero to prevent over-fitting
# (Kiryo et al., 2017)
risk_nnpu = pu_non_negative_risk(y_pu, y_score, pi=0.3)Both functions accept a loss argument (currently "logistic").
5.5 Diagnostics
Two utility functions help detect why a model may be performing poorly.
from pulearn.metrics import pu_distribution_diagnostics, homogeneity_metrics
# KL divergence between labeled and unlabeled score distributions
# Near-zero divergence → model cannot separate positives from unlabeled
diag = pu_distribution_diagnostics(y_pu, y_score)
print(diag["kl_divergence"])
# STD and IQR of predicted-negative scores
# Low STD/IQR → model may be over-relying on trivial features
hom_metrics = homogeneity_metrics(y_pu, y_score)
print(hom_metrics["std"], hom_metrics["iqr"])5.6 Scikit-learn Integration
make_pu_scorer wraps any PU metric as a make_scorer-compatible callable, enabling direct use with GridSearchCV and RandomizedSearchCV.
from sklearn.model_selection import GridSearchCV
from pulearn.metrics import make_pu_scorer
scorer = make_pu_scorer("pu_f1", pi=0.3)
gs = GridSearchCV(estimator, param_grid, scoring=scorer)
gs.fit(X_train, y_pu_train)Supported metric names for make_pu_scorer:
"lee_liu" |
Lee & Liu score (no pi required) |
"pu_recall" |
PU recall (no pi required) |
"pu_precision" |
Unbiased PU precision |
"pu_f1" |
Unbiased PU F1 |
"pu_specificity" |
Expected specificity |
"pu_roc_auc" |
Adjusted ROC-AUC (Sakai 2018) |
"pu_average_precision" |
Area Under Lift (AUL) |
"pu_unbiased_risk" |
uPU risk (lower is better) |
"pu_non_negative_risk" |
nnPU risk (lower is better) |
Risk metrics are wrapped with greater_is_better=False so that GridSearchCV correctly minimises them.
5.7 Complete Example
An end-to-end demo comparing naive F1 inflation vs. corrected metrics on synthetic SCAR data can be found in the examples directory:
python examples/PUMetricsEvaluationExample.py6 Examples
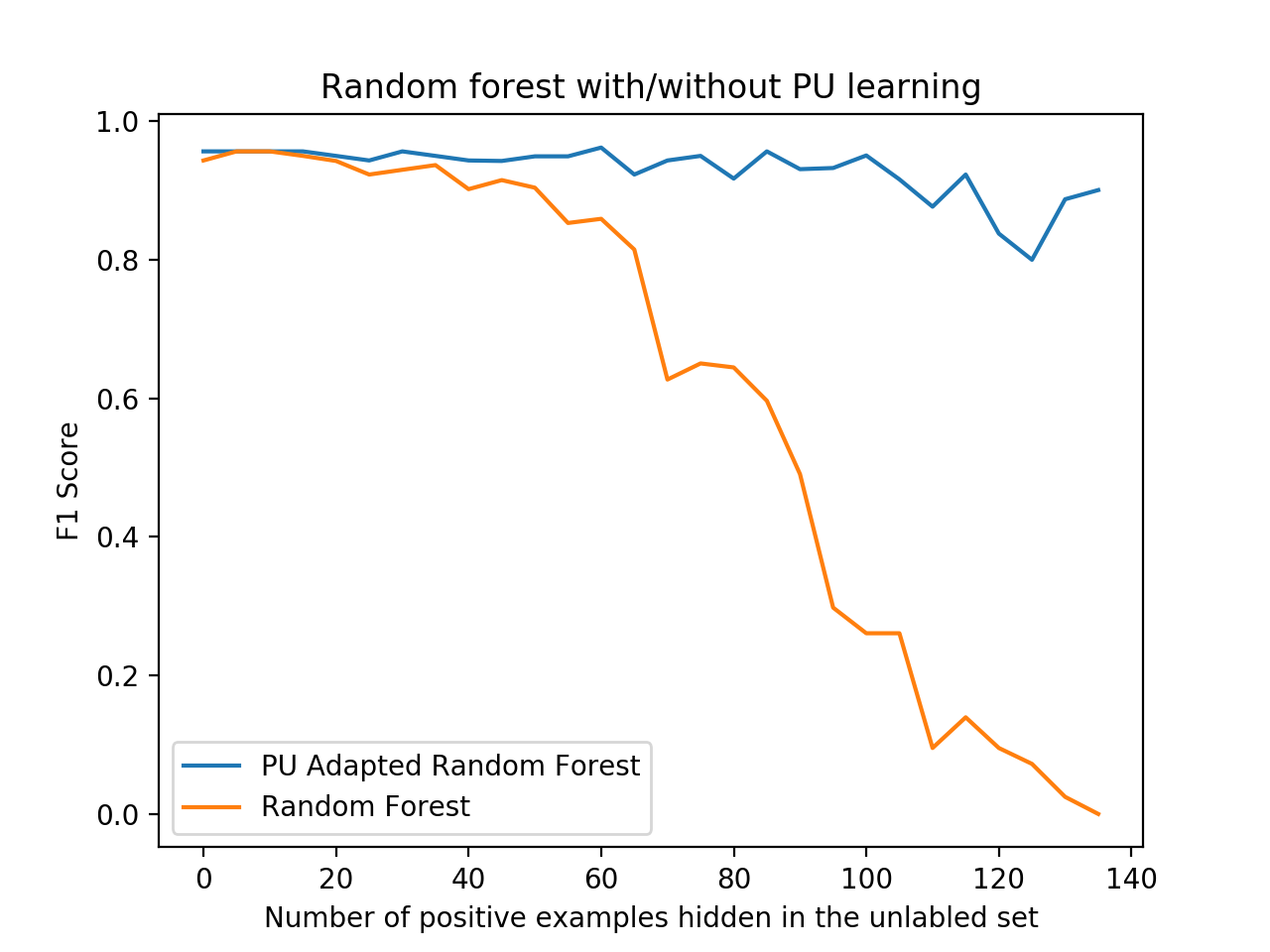
A nice code example of the classic Elkan-Noto classifier used for classification on the Wisconsin breast cancer dataset , comparing it to a regular random forest classifier, can be found in the examples directory.
To run it, clone the repository, and run the following command from the root of the repository, with a python environment where pulearn is installed:
python examples/BreastCancerElkanotoExample.pyYou should see a nice plot like the one below, comparing the F1 score of the PU learner versus a naive learner, demonstrating how PU learning becomes more effective - or worthwhile - the more positive examples are “hidden” from the training set.
7 Contributing
Package author and current maintainer is Shay Palachy (shay.palachy@gmail.com); You are more than welcome to approach him for help. Contributions are very welcomed, especially since this package is very much in its infancy and many other PU Learning methods can be added.
7.1 Installing for development
Clone:
git clone git@github.com:pulearn/pulearn.gitInstall in development mode with test dependencies:
cd pulearn
pip install -e ".[test]"7.2 Running the tests
To run the tests, use:
python -m pytestNotice pytest runs are configured by the pytest.ini file. Read it to understand the exact pytest arguments used.
7.3 Adding tests
At the time of writing, pulearn is maintained with a test coverage of 100%. Although challenging, I hope to maintain this status. If you add code to the package, please make sure you thoroughly test it. Codecov automatically reports changes in coverage on each PR, and so PR reducing test coverage will not be examined before that is fixed.
Tests reside under the tests directory in the root of the repository. Each model has a separate test folder, with each class - usually a pipeline stage - having a dedicated file (always starting with the string “test”) containing several tests (each a global function starting with the string “test”). Please adhere to this structure, and try to separate tests cases to different test functions; this allows us to quickly focus on problem areas and use cases. Thank you! :)
7.4 Code style
pulearn code is written to adhere to the coding style dictated by flake8. Practically, this means that one of the jobs that runs on the project’s Travis for each commit and pull request checks for a successful run of the flake8 CLI command in the repository’s root. Which means pull requests will be flagged red by the Travis bot if non-flake8-compliant code was added.
To solve this, please run flake8 on your code (whether through your text editor/IDE or using the command line) and fix all resulting errors. Thank you! :)
7.5 Adding documentation
This project is documented using the numpy docstring conventions, which were chosen as perhaps the most widelspread conventions both supported by common tools such as Sphinx and resulting in human-readable docstrings (in my personal opinion, of course). When documenting code you add to this project, please follow these conventions.
Additionally, if you update this README.rst file, use python setup.py checkdocs to validate it compiles.
8 License
This package is released as open-source software under the BSD 3-clause license. See LICENSE_NOTICE.md for the different copyright holders of different parts of the code.
9 Credits
Implementations code by:
Elkan & Noto - Alexandre Drouin and AditraAS.
Bagging PU Classifier - Roy Wright.
Bayesian PU Classifiers (PNB, WNB, PTAN, WTAN) - ported from Bayesian Classifiers for PU Learning by Chengning Zhang (MIT License).
Packaging, testing and documentation by Shay Palachy.
Fixes and feature contributions by:
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pulearn-0.2.0.tar.gz.
File metadata
- Download URL: pulearn-0.2.0.tar.gz
- Upload date:
- Size: 77.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
73be132a3129458b4f517aeab3f4b72dd7bd7c60f1e08b97669f0c039d4ec843
|
|
| MD5 |
81c3ef517fde2924c04a35695447aca8
|
|
| BLAKE2b-256 |
06c922c0b98439ee488eb818bd68c11f444199353afe923235fd7171bac364a3
|
Provenance
The following attestation bundles were made for pulearn-0.2.0.tar.gz:
Publisher:
release.yml on pulearn/pulearn
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
pulearn-0.2.0.tar.gz -
Subject digest:
73be132a3129458b4f517aeab3f4b72dd7bd7c60f1e08b97669f0c039d4ec843 - Sigstore transparency entry: 1104914148
- Sigstore integration time:
-
Permalink:
pulearn/pulearn@d4683e941b047f536462ac8a3f8aacfbfc500e30 -
Branch / Tag:
refs/heads/master - Owner: https://github.com/pulearn
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@d4683e941b047f536462ac8a3f8aacfbfc500e30 -
Trigger Event:
push
-
Statement type:
File details
Details for the file pulearn-0.2.0-py3-none-any.whl.
File metadata
- Download URL: pulearn-0.2.0-py3-none-any.whl
- Upload date:
- Size: 77.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d04af28e198e3d99362a4559fe0bc1b100707c621c915b007d6bcd8c29249635
|
|
| MD5 |
025691109b5ac4493e67ee66754bba56
|
|
| BLAKE2b-256 |
e8879b76408de3cdc7e774dc7e512497b5e84f63e996938940f533c6a40aa872
|
Provenance
The following attestation bundles were made for pulearn-0.2.0-py3-none-any.whl:
Publisher:
release.yml on pulearn/pulearn
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
pulearn-0.2.0-py3-none-any.whl -
Subject digest:
d04af28e198e3d99362a4559fe0bc1b100707c621c915b007d6bcd8c29249635 - Sigstore transparency entry: 1104914222
- Sigstore integration time:
-
Permalink:
pulearn/pulearn@d4683e941b047f536462ac8a3f8aacfbfc500e30 -
Branch / Tag:
refs/heads/master - Owner: https://github.com/pulearn
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@d4683e941b047f536462ac8a3f8aacfbfc500e30 -
Trigger Event:
push
-
Statement type:

















