Markov Clustering in Python
Project description
pyMarkovClustering
Table of contents
Overview
pyMarkovClustering is a python package for Markov Clustering (MCL) and its graph visualization. While there is already a python library markov_clustering that implements the MCL algorithm, it has not been maintained for a long time and lacks some functionality. To address these issues, pyMarkovClustering was developed.
[!NOTE] This library uses scipy sparse matrix for its MCL implementation and can cluster simple graphs with a few thousand nodes without any problems. However, if you need to cluster a complex graph with tens of thousands of nodes or more, I recommend using mcl command-line tool for better runtime performance and memory efficiency.
Installation
Python 3.9 or later is required for installation.
For visualization, networkx and extra packages (e.g. matplotlib, pygraphviz) are also required.
Install PyPI package:
pip install pymarkovclustering
pip install networkx[default,extra] # For visualization
Install conda-forge package:
conda install -c conda-forge pymarkovclustering
conda install -c conda-forge networkx matplotlib-base pygraphviz pydot lxml # For visualization
[!NOTE] pygraphviz installation requires graphviz and C/C++ compiler.
If you encounter installation troubles, see pygraphviz install docs for details.
API Usage
See notebooks and API docs in documents for more details.
Markov Clustering
Simple edges
import pymarkovclustering as pymcl
# List of edges (source, target, weight)
edges = [
("A", "B", 10),
("A", "C", 10),
("B", "C", 2),
("D", "E", 5),
("F", "G", 2),
("H", "I", 0.0),
]
# load edges as matrix, MCL, extract clusters
matrix, labels = pymcl.edges_to_sparse_matrix(edges)
mcl_matrix = pymcl.mcl(matrix, quiet=False)
clusters = pymcl.extract_clusters(mcl_matrix, labels)
for i, cluster in enumerate(clusters, 1):
print(f"Cluster{i:03d}: {cluster}")
Output:
Cluster001: ['A', 'B', 'C']
Cluster002: ['D', 'E']
Cluster003: ['F', 'G']
Cluster004: ['H']
Cluster005: ['I']
Random generated edges
import pymarkovclustering as pymcl
# Generate random edges for MCL test
edges = pymcl.random_edges(30, min_cluster_size=2, max_cluster_size=6)
print(f"Edges: {edges}\n")
# easymcl automates load edges as matrix, MCL, extract clusters
clusters = pymcl.easymcl(edges, inflation=2.0)
for i, cluster in enumerate(clusters, 1):
print(f"Cluster{i:03d}: {cluster}")
Output:
Edges: [('5_2', '5_5', 0.625), ('2_1', '2_5', 0.602), ('6_4', '6_5', 0.301), ('5_3', '5_6', 0.73), ('5_2', '5_6', 0.612), ('5_3', '5_5', 0.333), ('2_3', '2_5', 0.33), ('5_1', '5_3', 0.918), ('1_2', '1_4', 0.218), ('7_1', '7_2', 0.291), ('4_2', '4_3', 0.553), ('3_1', '3_2', 0.354), ('5_3', '5_4', 0.828), ('2_2', '2_4', 0.099), ('6_2', '6_5', 0.875), ('2_1', '2_3', 0.533), ('2_1', '2_4', 0.705), ('5_4', '5_5', 0.704), ('1_1', '1_4', 0.968), ('2_2', '2_5', 0.074), ('5_1', '5_5', 0.093), ('1_2', '1_3', 0.892), ('6_2', '6_3', 0.091), ('1_3', '1_5', 0.095), ('6_2', '6_4', 0.993), ('5_2', '5_4', 0.785), ('1_1', '1_3', 0.83), ('4_3', '4_4', 0.521), ('6_1', '6_2', 0.222), ('4_1', '4_3', 0.64), ('2_3', '2_4', 0.85), ('4_1', '4_2', 0.316), ('6_1', '6_5', 0.543), ('6_3', '6_5', 0.489), ('5_1', '5_6', 0.84), ('4_1', '4_4', 0.204), ('1_3', '1_4', 0.14), ('1_2', '1_5', 0.139), ('7_1', '7_3', 0.125), ('6_1', '6_3', 0.803), ('5_4', '5_6', 0.063), ('2_2', '2_3', 0.147), ('2_1', '2_2', 0.987), ('4_2', '4_4', 0.443), ('5_2', '5_3', 0.71), ('7_2', '7_3', 0.333), ('6_3', '6_4', 0.998), ('1_4', '1_5', 0.799), ('1_1', '1_5', 0.358), ('5_1', '5_4', 0.916), ('5_1', '5_2', 0.062), ('2_4', '2_5', 0.56), ('1_1', '1_2', 0.918), ('5_5', '5_6', 0.917), ('6_1', '6_4', 0.142)]
Cluster001: ['5_2', '5_5', '5_3', '5_6', '5_1', '5_4']
Cluster002: ['2_1', '2_5', '2_3', '2_2', '2_4']
Cluster003: ['6_4', '6_5', '6_2', '6_3', '6_1']
Cluster004: ['1_2', '1_4', '1_1', '1_3', '1_5']
Cluster005: ['4_2', '4_3', '4_4', '4_1']
Cluster006: ['7_1', '7_2', '7_3']
Cluster007: ['3_1', '3_2']
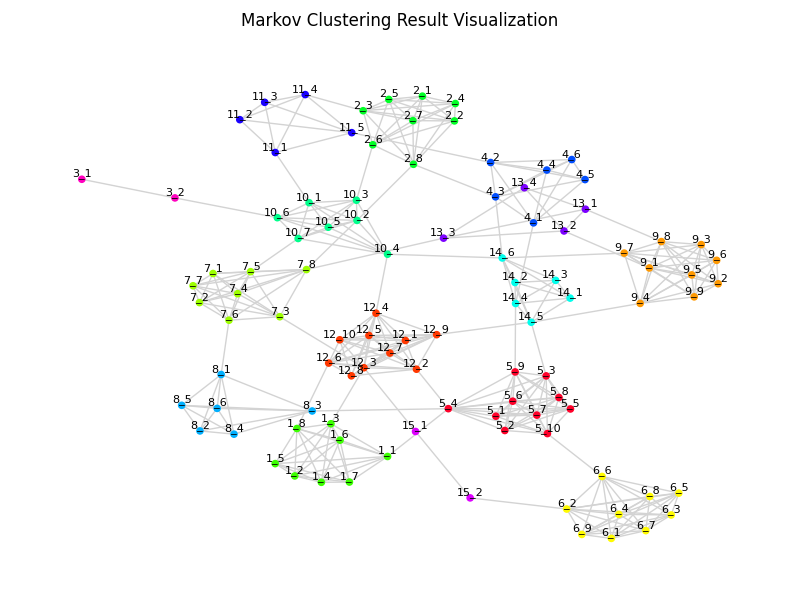
Visualization
import pymarkovclustering as pymcl
# Generate random edges for MCL test
edges = pymcl.random_edges(100, random_add_rate=0.1, min_cluster_size=2)
# easymclviz automates load edges as matrix, MCL, extract clusters, visualization
fig = pymcl.easymclviz(edges, inflation=2.0, show_label=True)
fig.suptitle("Markov Clustering Result Visualization")
fig.savefig("clusters.png", dpi=100)
CLI Usage
pyMarkovClustering provides simple CLI for running MCL and extract clusters from edges file.
Option
$ pymcl --help
usage: pymcl [options] edges.tsv -o clusters.tsv
Markov Clustering in Python
positional arguments:
edges Input edges(source, target, weight) tab-delimited file
optional arguments:
-o , --outfile Output tab-delimited clusters file (default: stdout)
-I , --inflation Inflation factor (default: 2.0)
--max_iter Max number of iteration (default: 100)
-q, --quiet No print log on screen (default: OFF)
-v, --version Print version information
-h, --help Show this help message and exit
Example Command
pymcl edges.tsv -I 2.0 -o clusters.tsv
e.g. edges.tsv >>> clusters.tsv
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pymarkovclustering-0.1.0.tar.gz.
File metadata
- Download URL: pymarkovclustering-0.1.0.tar.gz
- Upload date:
- Size: 277.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
51893d8614532c52888edbdc57340abfc1b58d06ad64516465268e807b6adbf1
|
|
| MD5 |
e32845e06d1ebcf0063ac9b7a11f303a
|
|
| BLAKE2b-256 |
f49b6a23615f3133b7a102af4668c204145a3773831b04d129aa5da2df47eed0
|
File details
Details for the file pymarkovclustering-0.1.0-py3-none-any.whl.
File metadata
- Download URL: pymarkovclustering-0.1.0-py3-none-any.whl
- Upload date:
- Size: 15.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d892150e41b6c2ffe90f242abe51b44a63db1fcb45bf9f434b5264b64c6994da
|
|
| MD5 |
82036eacfb53b5aa769359d52c453935
|
|
| BLAKE2b-256 |
323874710090052aec78309e5733028ab2e8edf205773f8c30d55e6f524ae01d
|