Utility functions to ease interactive work with pydna.
Project description
pydna-utils is a package containing utilities for pydna facilitating interactive use.
- open a Dsecrecord in the ApE plasmid editor or snapgene
- a global PCR primer list
- a global restriction enzyme list
- cached access to genbank
Install:
pip install pydna-utils
pydna_utils creates a user settings file (pydna_config.toml) where user
information and links to useful data can be placed.
On my linux mint laptop, this is located at /home/bjorn/.config/pydna/pydna_config.toml.
The platformdirs package is used to decide where this file should be located.
The settings file is a TOML file and has this content by default:
pydna_ape_cmd = "/usr/bin/tclsh /home/bjorn/.ApE/ApE.tcl"
pydna_snapgene_cmd = "/opt/gslbiotech/snapgene/snapgene.sh"
pydna_enzymes = "/home/bjorn/.ApE/Enzymes/LGM_group.txt"
pydna_primers = "/home/bjorn/myvault/PRIMERS.md"
pydna_email = "someone@example.com"
pydna_ncbi_cache_dir = "/home/bjorn/.cache/pydna_utils"
pydna_ncbi_expiration = "604800"
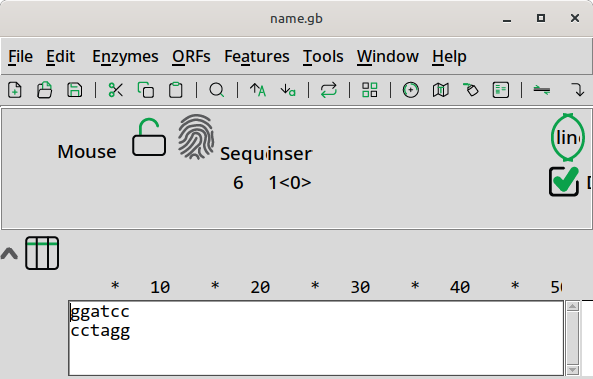
pydna_utils.editor.ape starts the ApE plasmid editor.
>>> from pydna_utils.editor import ape
>>> from pydna.dseqrecord import Dseqrecord
>>> sequence = Dseqrecord("GGATCC")
>>> sequence.seq
Dseq(-6)
GGATCC
CCTAGG
>>> ape(sequence)
pydna_utils.myprimers.PrimerList enables a global primer list for pydna
The primer list is typically a text file containing primer sequences in a format that pydna understands, such as FASTA.
This feature is most useful if the laboratory keeps a plain text file with primer sequences for all lab members. As visible below, our list had 1821 primers when this example was created.
You may not want to ship this list with your pydna code. The list may be long nd contain mostly irrelevant primers. It may also have information that best be kept inside the lab. For this reason, PrimerList remembers which primers have been accessed in a particular session and can also create pydna code ready to be pasted into a pydna script or notebook that contain the relevant part of the list. See example below:
>>> from pydna_utils.myprimers import PrimerList
>>> pl = PrimerList()
>>> len(pl)
1821
>>> pl[1]
1_5CYC1clone 35-mer:5'-GATCGGCCGGATCCA..CCG-3'
>>> pl[2]
2_3CYC1clon 35-mer:5'-CGATGTCGACTTAGA..AAG-3'
>>> print(pl[1].format("fasta"))
>>> print(pl[2].format("fasta"))
>>> pl.accessed
[1_5CYC1clone 35-mer:5'-GATCGGCCGGATCCA..CCG-3',
2_3CYC1clon 35-mer:5'-CGATGTCGACTTAGA..AAG-3']
>>> pl.code(pl.accessed)
from pydna.parsers import parse_primers
p = {}
p[1], p[2] = parse_primers('''
>1_5CYC1clone 35-mer
GATCGGCCGGATCCAAATGACTGAATTCAAGGCCG
>2_3CYC1clon 35-mer
CGATGTCGACTTAGATCTCACAGGCTTTTTTCAAG
''')
pydna_utils.myenzymes.myenzymes enables a global restriction enzyme batch
pydna_enzymes should contain a path to a text file containing restriction enzyme names. No particular formatting is required, but the names have to be separated by white space and exactly as they appear on REBASE.
>>> from pydna_utils.myenzymes import myenzymes
>>> myenzymes
RestrictionBatch(['AatII', 'Acc65I', 'AflII', 'AjiI', 'BamHI', 'BglI'])
pydna_utils.genbank.genbank is a cached version of pydna.genbank.genbank
>>> from pydna_utils.genbank import genbank
>>> gb_sequence = genbank("A23695.1")
>>> gb_sequence
Gbnk(-4 A23695.1)
>>> gb_sequence.seq
Dseq(-4)
AAAA
TTTT
The settings pydna_email, pydna_ncbi_cache_dir and pydna_ncbi_expiration
are important for how the cache works.
The user only needs to set the email as this is a NCBI requirement.
>>> import pydna_utils
>>> pydna_utils.get_settings()
+-----------------------+-----------------------------------------+
| Setting | Value |
+-----------------------+-----------------------------------------+
| pydna_ape_cmd | /usr/bin/tclsh /home/bjorn/.ApE/ApE.tcl |
| pydna_snapgene_cmd | /opt/gslbiotech/snapgene/snapgene.sh |
| pydna_enzymes | /home/bjorn/.ApE/Enzymes/LGM_group.txt |
| pydna_primers | /home/bjorn/myvault/PRIMERS.md |
| pydna_email | b*******b@gmail.com |
| pydna_ncbi_cache_dir | /home/bjorn/.cache/pydna_utils |
| pydna_ncbi_expiration | 604800 |
+-----------------------+-----------------------------------------+
>>> from pydna_utils import open_config_file
>>> open_config_file() # opens the config file for editing in system text editor.
>>> from pydna_utils import open_cache_folder
>>> open_cache_folder() # opens the cache folder in system file explorer.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pydna_utils-0.0.4.tar.gz.
File metadata
- Download URL: pydna_utils-0.0.4.tar.gz
- Upload date:
- Size: 11.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/2.2.1 CPython/3.12.7 Linux/6.14.0-36-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e6521b16d85cdc86926036e09cf117d22b429b285e8f27ef5b88475e84a35615
|
|
| MD5 |
df9009096ae8beedc1fec0545edb38d6
|
|
| BLAKE2b-256 |
b8957cb809fc11a9ef9f13dafcf01f9c33201ddd94e406859aeb06f617f8bec5
|
File details
Details for the file pydna_utils-0.0.4-py3-none-any.whl.
File metadata
- Download URL: pydna_utils-0.0.4-py3-none-any.whl
- Upload date:
- Size: 12.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/2.2.1 CPython/3.12.7 Linux/6.14.0-36-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9a0e67c4b89b83d06850874d6905ea21f424e5423b795b9a7623c130cb3eddb4
|
|
| MD5 |
7d89f2b17e41e31085caf22cbdbb52f8
|
|
| BLAKE2b-256 |
5b8100c50945cb5f179d016ed30137995d8dd960ec65a093452d85f2929f4cce
|