Computational microscopy toolkit for label-free imaging
Project description
recOrder
This package offers a comprehensive pipeline, command line interface, and napari plugin for quantitative label-free microscopy.
In this repo you will find python tools and UI that allow the user to calibrate microscope hardware, acquire multi-modal data, reconstruct density and anisotropy, and visualize the data.
The acquisition, calibration, background correction, reconstruction, and applications of QLIPP are described in the following E-Life Paper:
Syuan-Ming Guo, Li-Hao Yeh, Jenny Folkesson, Ivan E Ivanov, Anitha P Krishnan, Matthew G Keefe, Ezzat Hashemi, David Shin, Bryant B Chhun, Nathan H Cho, Manuel D Leonetti, May H Han, Tomasz J Nowakowski, Shalin B Mehta, "Revealing architectural order with quantitative label-free imaging and deep learning," eLife 2020;9:e55502 DOI: 10.7554/eLife.55502 (2020).
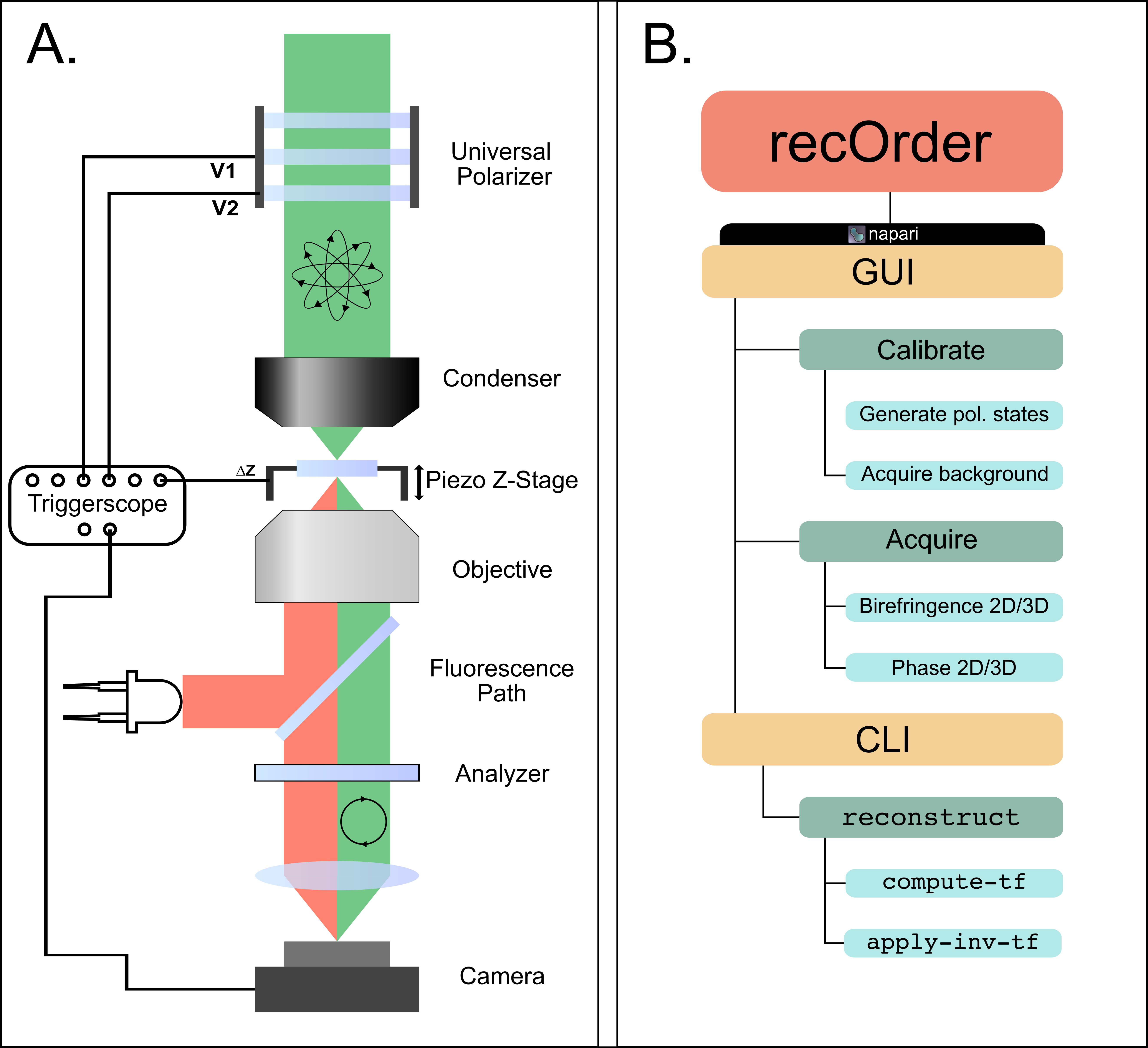
recOrder is to be used alongside the QLIPP module, whose design has been optimized to fit on a conventional widefield microscope (Panel A below). The QLIPP module allows for the collection of label-free information consisting of the intrinsic anisotropy of the sample and its relative phase (density). All of these measurements are collected through compensated, polarization diverse illumination and quantitatively recovered through recOrder's computational reconstruction pipeline. The overall structure of recOrder can be visualized below in Panel B, highlighting the two different usage modes and their features: graphical user interface (GUI) through napari and command line interfact (CLI).
Dataset
Slides and a dataset shared during a workshop on QLIPP and recOrder can be found on Zenodo.
Installation
Easy installation:
(Optional but recommended) install anaconda and create a virtual environment
conda create -n recorder python
conda activate recorder
Install napari:
pip install "napari[all]"
Open napari and use the Plugin > Install/Uninstall Plugins... menu to install recOrder-napari.
Developer installation:
Install git and conda (either anaconda or miniconda).
Create a conda environment dedicated to recOrder:
conda create -n recorder python
conda activate recorder
Clone this repository:
git clone https://github.com/mehta-lab/recOrder.git
Install recOrder and its dependencies:
cd recOrder
pip install -e ".[dev]"
Optional napari plugin: recOrder includes a napari plugin that can be used to acquire data via MicroManager. To install napari use
pip install "napari[all]"
To run the recOrder plugin use
napari -w recOrder-napari
To acquire data via MicroManager, follow the instructions on the wiki.
Optional GPU: recOrder supports NVIDIA GPU computation with the cupy package. Follow these instructions to install cupy and check its installation with import cupy. To enable gpu processing, set use_gpu: True in the config files.
Command-line usage
Type recOrder.help for instructions on the two command-line modes: recOrder.reconstruct and recOrder.convert.
recOrder.reconstruct
recOrder.reconstruct uses configuration files to select reconstruction parameters. Start with an example configuration file /examples/example_configs/config_example_qlipp.yml and modify the parameters to match your dataset.
Run the reconstruction with
recOrder.reconstruct --config <path/to/config.yml>
The following command-line arguments override parameters specified in the configuration file:
--method (str) method of reconstruction: QLIPP,IPS,UPTI'
--mode (str) mode of reconstruction: 2D, 3D'
--data_dir (str) path to raw data folder'
--save_dir (str) path to folder where reconstructed data will be saved'
--name (str) name under which to save the reconstructed data'
--config (str) path to configuration file (see /examples/example_configs')
--overwrite (bool) True/False whether or not to overwrite data that exists under save_dir/name'
For example, this command uses the QLIPP reconstruction method even if the configuration file specifies a different reconstruction method
recOrder.reconstruct --config /path/to/config.yml --method QLIPP
recOrder.convert
recOrder.convert converts MicroManager tif files to ome-zarr files. For example
recOrder.convert --input <path/to/mm/tifs> --output <path/to/output.zarr> --data_type ometiff
License
Chan Zuckerberg Biohub Software License
This software license is the 2-clause BSD license plus clause a third clause that prohibits redistribution and use for commercial purposes without further permission.
Copyright © 2019. Chan Zuckerberg Biohub. All rights reserved.
Redistribution and use in source and binary forms, with or without modification, are permitted provided that the following conditions are met:
-
Redistributions of source code must retain the above copyright notice, this list of conditions and the following disclaimer.
-
Redistributions in binary form must reproduce the above copyright notice, this list of conditions and the following disclaimer in the documentation and/or other materials provided with the distribution.
-
Redistributions and use for commercial purposes are not permitted without the Chan Zuckerberg Biohub's written permission. For purposes of this license, commercial purposes are the incorporation of the Chan Zuckerberg Biohub's software into anything for which you will charge fees or other compensation or use of the software to perform a commercial service for a third party. Contact ip@czbiohub.org for commercial licensing opportunities.
THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS" AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file recOrder-napari-0.1.2.tar.gz.
File metadata
- Download URL: recOrder-napari-0.1.2.tar.gz
- Upload date:
- Size: 11.6 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
64728d224b36a5453f7ca995e5b4f4ef6ade34906502c84b37637e5b8c41e2da
|
|
| MD5 |
dcb5120696a4701c73259b06a61dcac7
|
|
| BLAKE2b-256 |
bf2e2783dd0c2db2ef350a18ed88038c655da2bdea0a4113a2fc47322eb62cf4
|
File details
Details for the file recOrder_napari-0.1.2-py3-none-any.whl.
File metadata
- Download URL: recOrder_napari-0.1.2-py3-none-any.whl
- Upload date:
- Size: 11.6 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cccd498d9a0c79bdf96849df75c778b358b9d3471a8cf53fc6418034d66dc8eb
|
|
| MD5 |
7f95486e2e47af243f1997752ec70563
|
|
| BLAKE2b-256 |
4a949d07b3589710974716f1182d3c04f60823b1f022be9e11e01580fff58340
|