Self-paced Ensemble for classification on class-imbalanced data.
Project description
Self-paced Ensemble for Highly Imbalanced Massive Data Classification (ICDE 2020)
Links: Paper | Slides | Video | arXiv | PyPI | API Reference | Related Projects | Zhihu/知乎
Self-paced Ensemble (SPE) is an ensemble learning framework for massive highly imbalanced classification. It is an easy-to-use solution to class-imbalanced problems, features outstanding computing efficiency, good performance, and wide compatibility with different learning models. This SPE implementation supports multi-class classification.
| Note: SPE is now a part of imbalanced-ensemble [Doc, PyPI]. Try it for more methods and advanced features! |
Cite Us
If you find this repository helpful in your work or research, we would greatly appreciate citations to the following paper:
@inproceedings{liu2020self-paced-ensemble,
title={Self-paced Ensemble for Highly Imbalanced Massive Data Classification},
author={Liu, Zhining and Cao, Wei and Gao, Zhifeng and Bian, Jiang and Chen, Hechang and Chang, Yi and Liu, Tie-Yan},
booktitle={2020 IEEE 36th International Conference on Data Engineering (ICDE)},
pages={841--852},
year={2020},
organization={IEEE}
}
Installation
It is recommended to use pip for installation.
Please make sure the latest version is installed to avoid potential problems:
$ pip install self-paced-ensemble # normal install
$ pip install --upgrade self-paced-ensemble # update if needed
Or you can install SPE by clone this repository:
$ git clone https://github.com/ZhiningLiu1998/self-paced-ensemble.git
$ cd self-paced-ensemble
$ python setup.py install
Following dependencies are required:
- python (>=3.6)
- numpy (>=1.13.3)
- scipy (>=0.19.1)
- joblib (>=0.11)
- scikit-learn (>=0.24)
- imblearn (>=0.7.0)
- imbalanced-ensemble (>=0.1.3)
Table of Contents
- Cite Us
- Installation
- Table of Contents
- Background
- Documentation
- Examples
- Results
- Miscellaneous
- References
- Related Projects
- Contributors ✨
Background
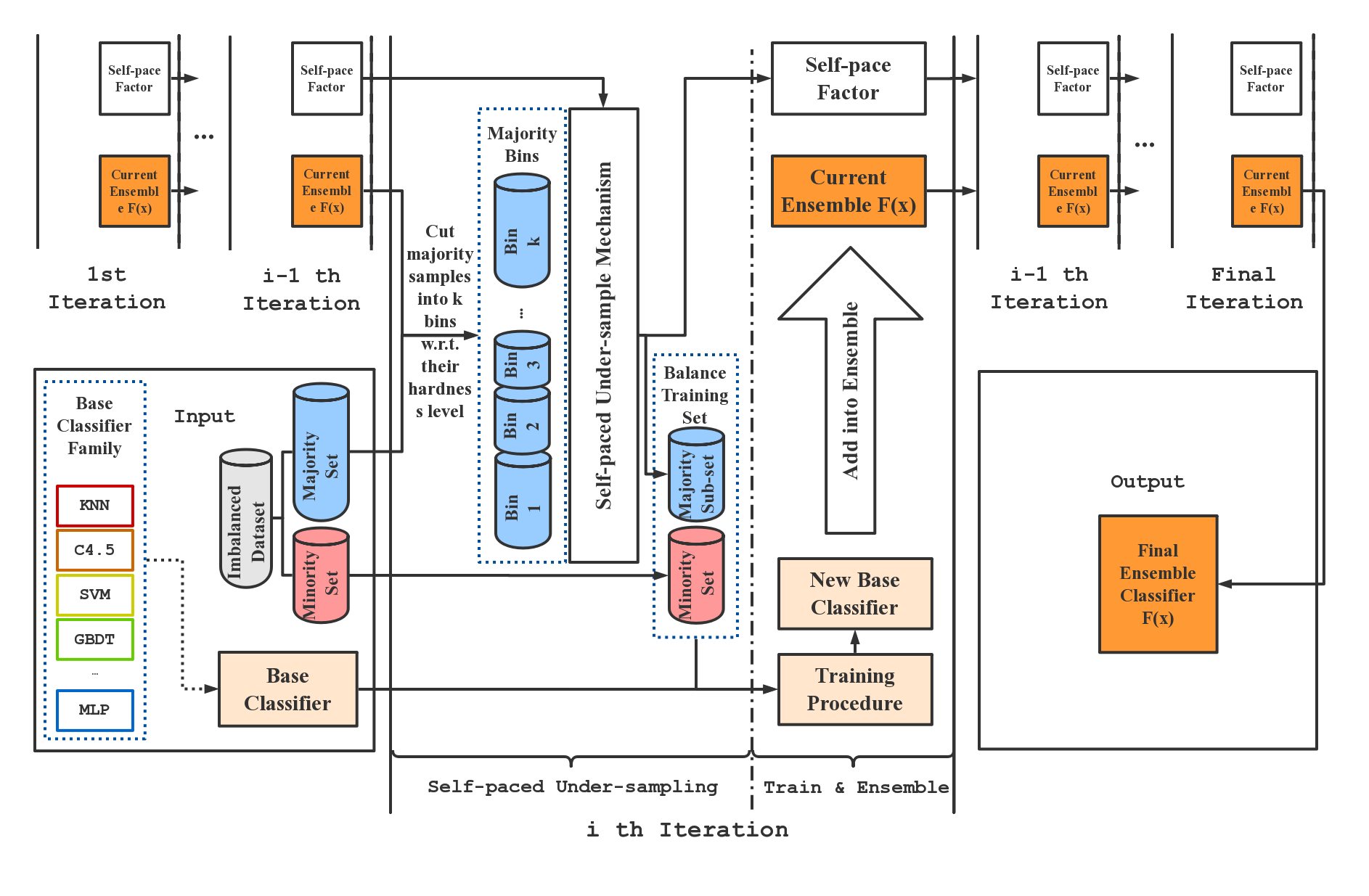
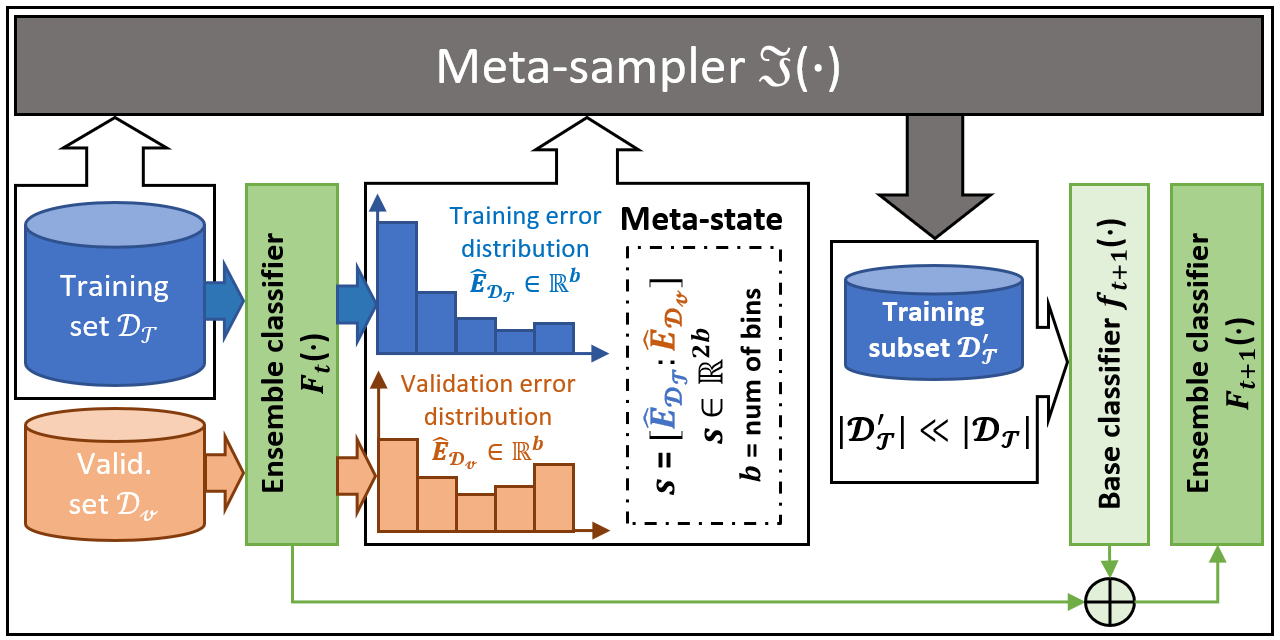
SPE performs strictly balanced under-sampling in each iteration and is therefore very computationally efficient. In addition, SPE does not rely on calculating the distance between samples to perform resampling. It can be easily applied to datasets that lack well-defined distance metrics (e.g. with categorical features / missing values) without any modification. Moreover, as a generic ensemble framework, our methods can be easily adapted to most of the existing learning methods (e.g., C4.5, SVM, GBDT, and Neural Network) to boost their performance on imbalanced data. Compared to existing imbalance learning methods, SPE works particularly well on datasets that are large-scale, noisy, and highly imbalanced (e.g. with imbalance ratio greater than 100:1). Such kind of data widely exists in real-world industrial applications. The figure below gives an overview of the SPE framework.
Documentation
Our SPE implementation can be used much in the same way as the sklearn.ensemble classifiers. Detailed documentation of SelfPacedEnsembleClassifier can be found HERE.
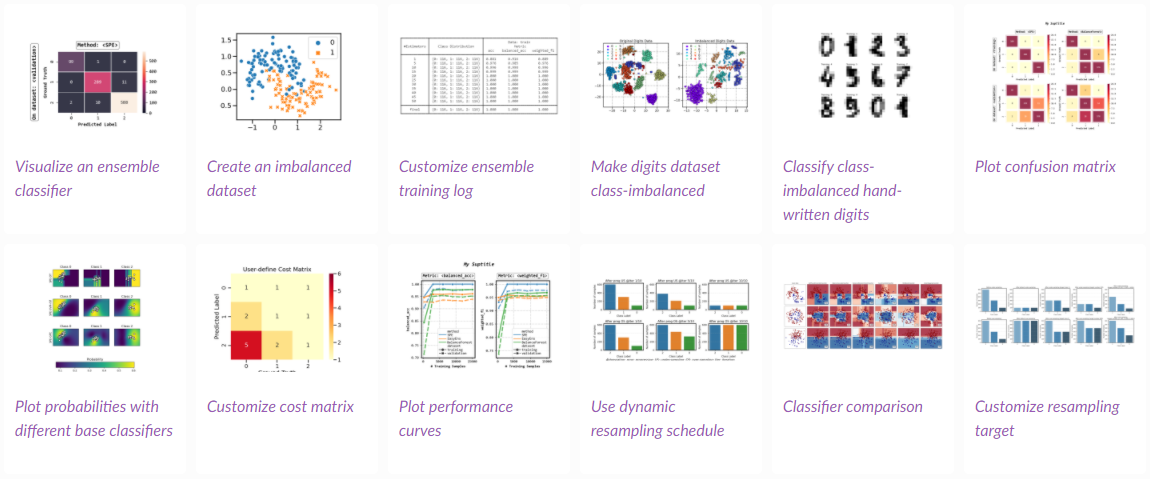
Examples
You can check out examples using SPE for more comprehensive usage examples.
API demo
from self_paced_ensemble import SelfPacedEnsembleClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
# Prepare class-imbalanced train & test data
X, y = make_classification(n_classes=2, random_state=42, weights=[0.1, 0.9])
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.5, random_state=42)
# Train an SPE classifier
clf = SelfPacedEnsembleClassifier(
base_estimator=DecisionTreeClassifier(),
n_estimators=10,
).fit(X_train, y_train)
# Predict with an SPE classifier
clf.predict(X_test)
Advanced usage example
Please see usage_example.ipynb.
Save & Load model
We recommend to use joblib or pickle for saving and loading SPE models, e.g.,
from joblib import dump, load
# save the model
dump(clf, filename='clf.joblib')
# load the model
clf = load('clf.joblib')
You can also use the alternative APIs provided in SPE:
from self_paced_ensemble.utils import save_model, load_model
# save the model
clf.save('clf.joblib') # option 1
save_model(clf, 'clf.joblib') # option 2
# load the model
clf = load_model('clf.joblib')
Compare SPE with other methods
Please see comparison_example.ipynb.
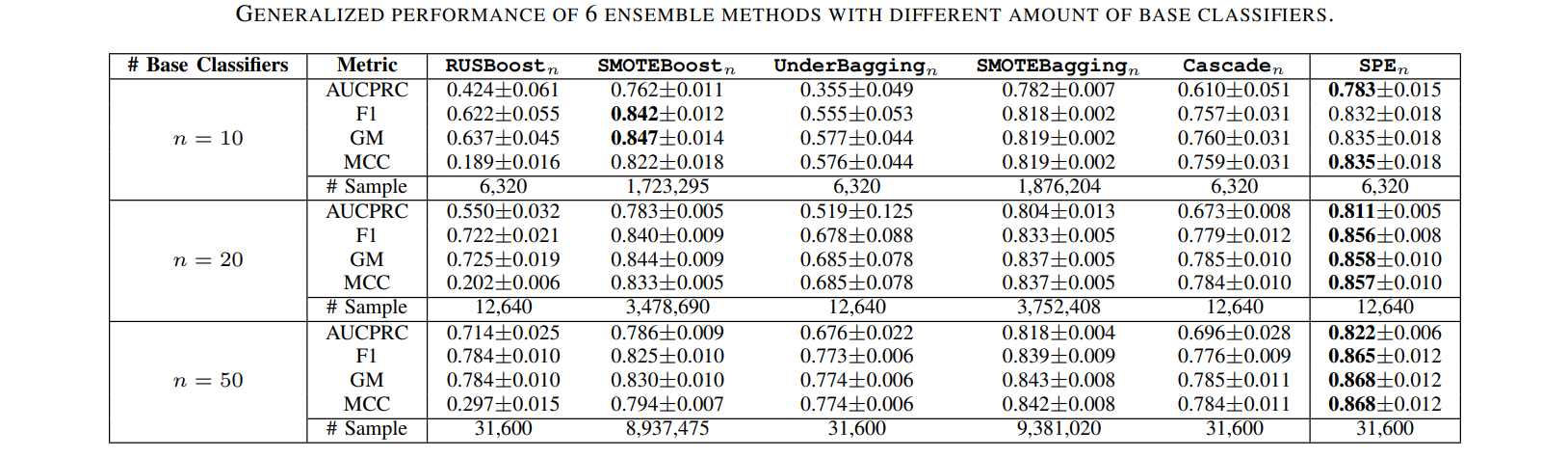
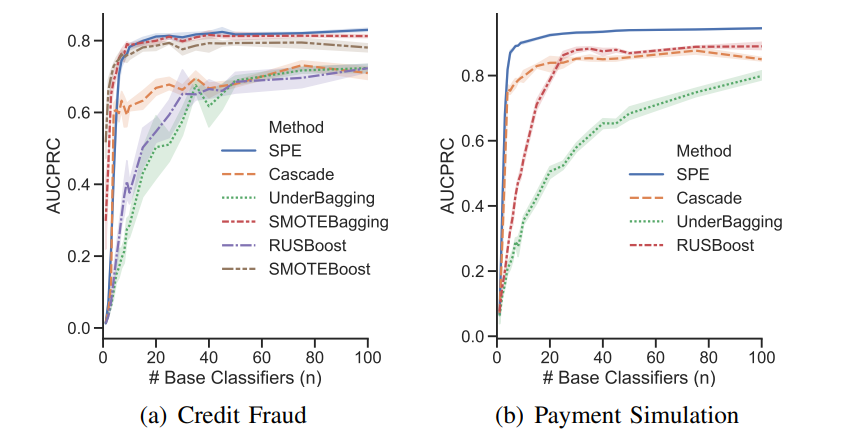
Results
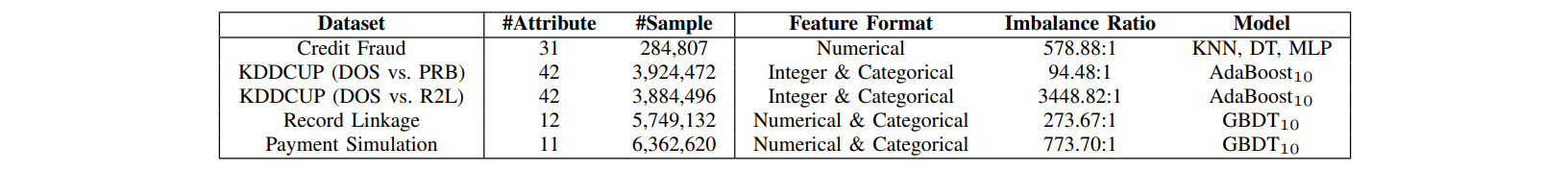
Dataset links: Credit Fraud, KDDCUP, Record Linkage, Payment Simulation.
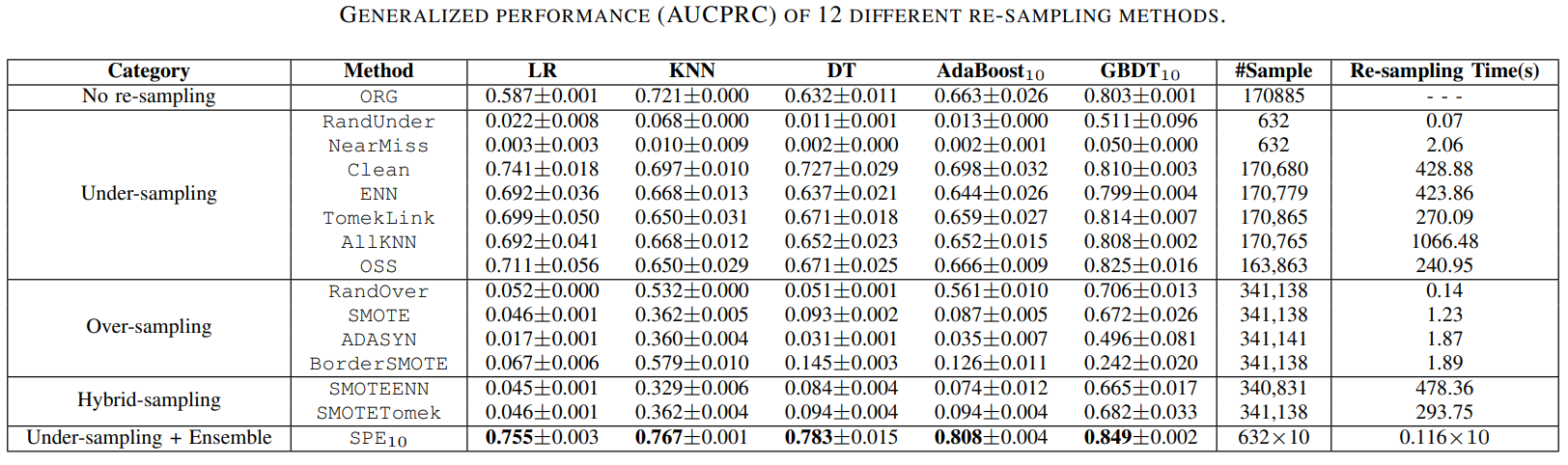
Comparisons of SPE with traditional resampling/ensemble methods in terms of performance & computational efficiency.
Miscellaneous
This repository contains:
- Implementation of Self-paced Ensemble
- Implementation of 5 ensemble-based imbalance learning baselines
SMOTEBoost[1]SMOTEBagging[2]RUSBoost[3]UnderBagging[4]BalanceCascade[5]
- Implementation of resampling based imbalance learning baselines [6]
- Additional experimental results
NOTE: The implementations of other ensemble and resampling methods are based on imbalanced-ensemble and imbalanced-learn.
References
| # | Reference |
|---|---|
| [1] | N. V. Chawla, A. Lazarevic, L. O. Hall, and K. W. Bowyer, Smoteboost: Improving prediction of the minority class in boosting. in European conference on principles of data mining and knowledge discovery. Springer, 2003, pp. 107–119 |
| [2] | S. Wang and X. Yao, Diversity analysis on imbalanced data sets by using ensemble models. in 2009 IEEE Symposium on Computational Intelligence and Data Mining. IEEE, 2009, pp. 324–331. |
| [3] | C. Seiffert, T. M. Khoshgoftaar, J. Van Hulse, and A. Napolitano, “Rusboost: A hybrid approach to alleviating class imbalance,” IEEE Transactions on Systems, Man, and Cybernetics-Part A: Systems and Humans, vol. 40, no. 1, pp. 185–197, 2010. |
| [4] | R. Barandela, R. M. Valdovinos, and J. S. Sanchez, “New applications´ of ensembles of classifiers,” Pattern Analysis & Applications, vol. 6, no. 3, pp. 245–256, 2003. |
| [5] | X.-Y. Liu, J. Wu, and Z.-H. Zhou, “Exploratory undersampling for class-imbalance learning,” IEEE Transactions on Systems, Man, and Cybernetics, Part B (Cybernetics), vol. 39, no. 2, pp. 539–550, 2009. |
| [6] | Guillaume Lemaître, Fernando Nogueira, and Christos K. Aridas. Imbalanced-learn: A python toolbox to tackle the curse of imbalanced datasets in machine learning. Journal of Machine Learning Research, 18(17):1–5, 2017. |
Related Projects
Check out Zhining's other open-source projects!
 Imbalanced-Ensemble [PythonLib] 
|
 Imbalanced Learning [Awesome] 
|
 Machine Learning [Awesome] 
|
 Meta-Sampler [NeurIPS] 
|
Contributors ✨
Thanks goes to these wonderful people (emoji key):
 Zhining Liu 💻 📖 💡 |
 Yuming Fu 💻 🐛 |
 Thúlio Costa 💻 🐛 |
 Neko Null 🚧 |
 lirenjieArthur 🐛 |
 AC手动机 🐛 |
 Carlo Moro 🤔 |
This project follows the all-contributors specification. Contributions of any kind welcome!
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file self-paced-ensemble-0.1.7.tar.gz.
File metadata
- Download URL: self-paced-ensemble-0.1.7.tar.gz
- Upload date:
- Size: 46.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f2ce7095975af7f2c39bd33159e06447e929c5cb0d93f80f4f051f592ab8be72
|
|
| MD5 |
9fad22e31adfc368c008c26da529b413
|
|
| BLAKE2b-256 |
3b8d41d8e2c6091d0107a2da763c317677c1f786b7693c689338ffe1e9155f78
|
File details
Details for the file self_paced_ensemble-0.1.7-py2.py3-none-any.whl.
File metadata
- Download URL: self_paced_ensemble-0.1.7-py2.py3-none-any.whl
- Upload date:
- Size: 48.1 kB
- Tags: Python 2, Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
958a75d6220ef4c3185c28c7f2dc8b66a5ace88f703067ced14a1c53b55b9674
|
|
| MD5 |
31f64e4b15309190535d7c3355b832c2
|
|
| BLAKE2b-256 |
102c0c6196b95bbfe10ff38dbbb48b96f9813553c7e30a8dd01f75a179edcab2
|