Pipeline that runs bcl2fastq and creates additional plots within a Snakemake workflow
Project description
This is is the demultiplex pipeline from the Sequana projet
- Overview:
Runs bcl2fastq on raw BCL data and creates plots to ease the QC validation
- Input:
A valid Illumina base calling directory
- Output:
a set of PNG files and the expected FastQ files
- Status:
production
- Citation:
Cokelaer et al, (2017), ‘Sequana’: a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI https://doi:10.21105/joss.00352
Installation
You must install Sequana first:
pip install sequana
Then, just install this package:
pip install sequana_demultiplex
Usage
sequana_pipelines_demultiplex --help sequana_pipelines_demultiplex --working-directory DATAPATH --bcl-directory bcldata
This creates a directory fastq. You just need to execute the pipeline:
cd demutliplex sh demutliplex.sh # for a local run
This launch a snakemake pipeline. If you are familiar with snakemake, you can retrieve the demutliplex.rules and config.yaml files and then execute the pipeline yourself with specific parameters:
snakemake -s demutliplex.rules --cores 4 --stats stats.txt
Or use sequanix interface.
Requirements
This pipelines requires the following executable(s):
bcl2fastq 2.20.0
Details
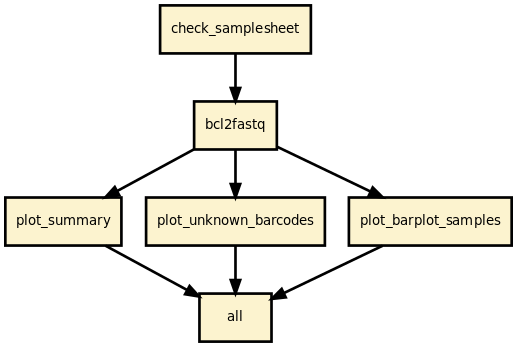
This pipeline runs bcl2fastq 2.20 and creates a set of diagnostics plots to help deciphering common issues such as missing index and sample sheet errors.
Rules and configuration details
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
Changelog
Version |
Description |
|---|---|
0.9.5 |
|
0.9.4 |
|
0.9.3 |
Fix regression bug |
0.9.2 |
remove warning due to relative paths. |
0.9.1 |
Make the merging options compulsory. Users must tell whether they want to merge the lanes or not. This avoid to do the merging or not whereas the inverse was expected. |
0.8.6 |
Uses 64G/biomics queue and 16 cores on a SLURM scheduler |
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file sequana_demultiplex-0.9.5.tar.gz.
File metadata
- Download URL: sequana_demultiplex-0.9.5.tar.gz
- Upload date:
- Size: 29.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.15.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/40.4.3 requests-toolbelt/0.9.1 tqdm/4.39.0 CPython/3.5.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e22ec7b883da5eaaebbaca8f177dfe0e08aa45869caa2e9b658aaf670dcafe64
|
|
| MD5 |
c7f1f4ea9fefeb86820c0a144616cc7a
|
|
| BLAKE2b-256 |
bde272c45152c18d0b6a111b48a05586e461bc32159d3b797626566ee8e707d9
|











