QC on various type of pacbio data
Project description




This is the pacbio_qc pipeline from the Sequana project
- Overview:
Quality control and analysis for PacBio long-read sequencing data (BAM files). Generates comprehensive statistics on read quality, length distribution, and GC content, with optional taxonomic classification.
- Input:
BAM files from PacBio sequencers (raw subreads, CCS, or processed reads)
- Output:
Per-sample HTML reports with interactive visualizations, quality metrics, and optional taxonomic classification; comprehensive summary report with all samples
- Status:
production
- Documentation:
This README file, the Wiki from the github repository (link above) and https://sequana.readthedocs.io
- Citation:
Cokelaer et al, (2017), ‘Sequana’: a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI doi:10.21105/joss.00352
Installation
Install via pip:
pip install sequana_pacbio_qc
Optional dependencies:
kraken2: For taxonomic classification (optional, disabled by default)
graphviz: For DAG visualization
apptainer: For containerized execution of tools
Quick Start
# Display help sequana_pacbio_qc --help # Create pipeline in current directory sequana_pacbio_qc --input-directory /path/to/bam/files # With optional Kraken taxonomy sequana_pacbio_qc --input-directory /path/to/bam/files --do-kraken --kraken-databases /path/to/kraken/db # Using apptainer containers sequana_pacbio_qc --input-directory /path/to/bam/files --apptainer-prefix ~/containers
This creates a pacbio_qc directory containing the pipeline and configuration files.
Execution
Execute the pipeline:
cd pacbio_qc bash pacbio_qc.sh
Or with custom Snakemake parameters:
snakemake -s pacbio_qc.rules -c config.yaml --cores 4 --stats stats.txt
Or use the sequanix graphical interface.
Configuration
The pipeline uses config.yaml to control:
Input data: BAM file directory and pattern matching
Kraken: Optional taxonomic database paths (disabled by default)
MultiQC: QC report options
Apptainer: Container image URLs (optional)
Pipeline Overview
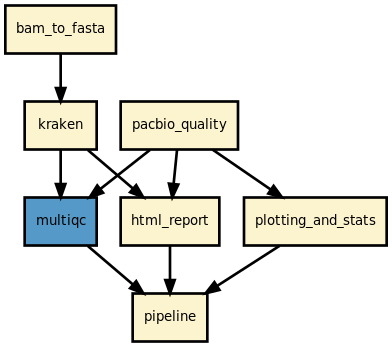
Workflow Details
The pipeline performs the following analyses on PacBio BAM files:
Quality Metrics: Computes read length statistics, GC content distribution, and signal-to-noise ratios
Visualizations: Generates histograms and scatter plots for quality assessment
Per-Sample Reports: Creates individual HTML reports for each sample with:
Read length distribution histograms
GC content analysis
SNR (signal-to-noise ratio) metrics
Quality overview with sample statistics
Taxonomy (Optional): Performs taxonomic classification using Kraken2 when enabled
Summary Report: Generates a comprehensive HTML summary with:
Overview of pipeline and all samples
Summary statistics table with links to per-sample reports
MultiQC aggregated quality metrics
Note: Kraken2 databases are not provided with the pipeline. This step is optional and disabled by default.
Changelog
Version |
Description |
|---|---|
1.0.1 |
HTML reports with pipeline overview; race condition handling for parallel execution with –apptainer-prefix; improved CI/CD workflows |
1.0.0 |
Uses latest wrappers and graphviz apptainers |
0.11.0 |
Release to use latests sequana_pipetools framework |
0.10.0 |
Update to use latest tools from sequana framework |
0.9.0 |
First release of sequana_pacbio_qc using latest sequana rules and modules (0.9.5) |
Contribute & Code of Conduct
To contribute to this project, please take a look at the Contributing Guidelines first. Please note that this project is released with a Code of Conduct. By contributing to this project, you agree to abide by its terms.
Rules and configuration details
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sequana_pacbio_qc-1.1.0.tar.gz.
File metadata
- Download URL: sequana_pacbio_qc-1.1.0.tar.gz
- Upload date:
- Size: 32.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/2.3.4 CPython/3.11.15 Linux/6.14.5-100.fc40.x86_64
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d06442ac9e59b89438e7905153df6ac8f4fdcd333d363e5edd34d3dc0f307713
|
|
| MD5 |
de8d9f22a85ebcde6f1a54555525d35b
|
|
| BLAKE2b-256 |
fbec65d0b9c87173bd9210ac1b70ec696b462770f2242f2adb53b73f0efab9e9
|
File details
Details for the file sequana_pacbio_qc-1.1.0-py3-none-any.whl.
File metadata
- Download URL: sequana_pacbio_qc-1.1.0-py3-none-any.whl
- Upload date:
- Size: 32.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/2.3.4 CPython/3.11.15 Linux/6.14.5-100.fc40.x86_64
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a233498c438de3a72ee846d5b42059d4300bc3ef64b30802576d290a91b6982f
|
|
| MD5 |
bd6c040e2065a92f201b0d13e8c0b920
|
|
| BLAKE2b-256 |
06d50db13993ed41eadfd6497ab16445f7756fd93f64bd1d03712a8daec6f2a7
|











