A utility for creating sashimi plot from Shiba output
Project description
🐕 shiba2sashimi 🍣 (v0.1.2)
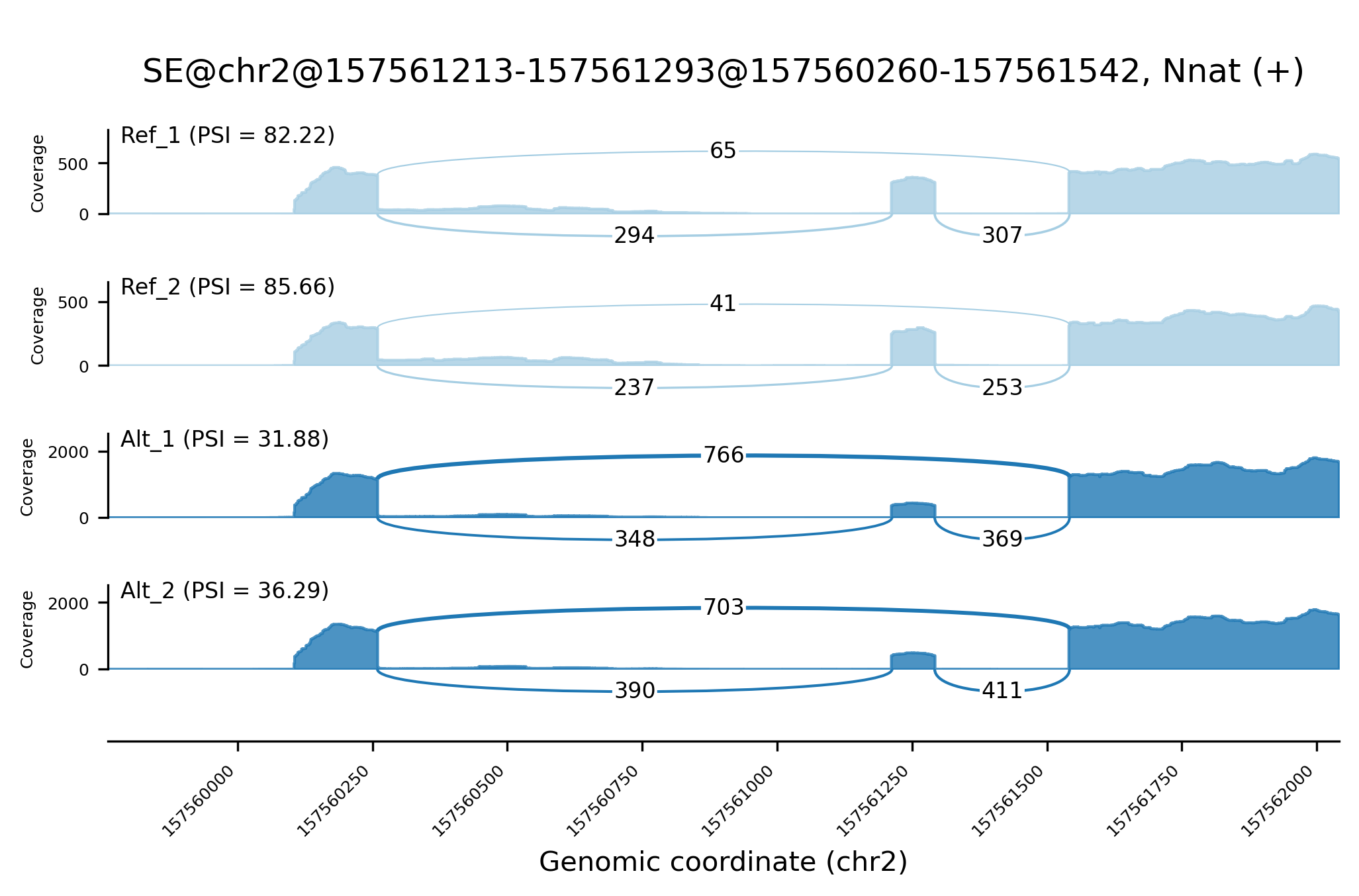
A utility to create Sashimi plots, a publication-quality visualization of RNA-seq data, from Shiba output. Greatly inspired by rmats2sashimiplot and MISO's original implementation.
Quick start
shiba2sashimi -e /path/to/Shiba/experiment_table.tsv \
-s /path/to/Shiba/output/ -o img/sashimi_example.png \
--id "SE@chr2@157561213-157561293@157560260-157561542"
How to install
pip
pip install shiba2sashimi
or
git clone https://github.com/Sika-Zheng-Lab/shiba2sashimi.git
cd shiba2sashimi
pip install .
conda
conda create -n shiba2sashimi python=3.9
conda activate shiba2sashimi
conda install -c bioconda shiba2sashimi
Docker
docker pull naotokubota/shiba2sashimi
Dependencies
- python (>=3.9)
- numpy (>=1.18.0,<2.0.0)
- matplotlib (>=3.1.0)
- pysam (>=0.22.0)
Usage
usage: shiba2sashimi [-h] -e EXPERIMENT -s SHIBA -o OUTPUT [--id ID] [-c COORDINATE] [--samples SAMPLES] [--groups GROUPS] [--colors COLORS] [--extend_up EXTEND_UP] [--extend_down EXTEND_DOWN]
[--smoothing_window_size SMOOTHING_WINDOW_SIZE] [--font_family FONT_FAMILY] [--dpi DPI] [-v]
shiba2sashimi v0.1.2 - Create Sashimi plot from Shiba output
optional arguments:
-h, --help show this help message and exit
-e EXPERIMENT, --experiment EXPERIMENT
Experiment table used for Shiba
-s SHIBA, --shiba SHIBA
Shiba output directory
-o OUTPUT, --output OUTPUT
Output file
--id ID Positional ID (pos_id) of the event to plot
-c COORDINATE, --coordinate COORDINATE
Coordinates of the region to plot
--samples SAMPLES Samples to plot. e.g. sample1,sample2,sample3 Default: all samples in the experiment table
--groups GROUPS Groups to plot. e.g. group1,group2,group3 Default: all groups in the experiment table. Overrides --samples
--colors COLORS Colors for each group. e.g. red,orange,blue
--extend_up EXTEND_UP
Extend the plot upstream. Only used when not providing coordinates. Default: 500
--extend_down EXTEND_DOWN
Extend the plot downstream. Only used when not providing coordinates. Default: 500
--smoothing_window_size SMOOTHING_WINDOW_SIZE
Window size for median filter to smooth coverage plot. Greater value gives smoother plot. Default: 21
--font_family FONT_FAMILY
Font family for labels
--dpi DPI DPI of the output figure. Default: 300
-v, --verbose Increase verbosity
Authors
- Naoto Kubota (0000-0003-0612-2300)
- Liang Chen (0000-0001-6164-4553)
- Sika Zheng (0000-0002-0573-4981)
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
shiba2sashimi-0.1.2.tar.gz
(12.2 kB
view details)
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file shiba2sashimi-0.1.2.tar.gz.
File metadata
- Download URL: shiba2sashimi-0.1.2.tar.gz
- Upload date:
- Size: 12.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
44175c6ef2d3799a33d8ef9d625637346e9bf9563b1e5691bd221a66e088cafe
|
|
| MD5 |
1393ce0ba9f5f35fbe34fdc5d91d5e86
|
|
| BLAKE2b-256 |
0ee506d8aa090903e3b82dbdb55a2151487bb7725b76c54bc54f28be70edcda8
|
File details
Details for the file shiba2sashimi-0.1.2-py3-none-any.whl.
File metadata
- Download URL: shiba2sashimi-0.1.2-py3-none-any.whl
- Upload date:
- Size: 13.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3af635bb976bdd5a806ad02498b4690933884c7d01cff95ccb9f1ed001fad022
|
|
| MD5 |
a8c5ed79f8ec0b68a20abed02fa2ca89
|
|
| BLAKE2b-256 |
c499b9d63f7792a9a508a34ac04f8c4e794461a63e0c14c38f6cd57d022d13bc
|