A utility for creating sashimi plot from Shiba output
Project description
🐕 shiba2sashimi 🍣 (v0.1.5)
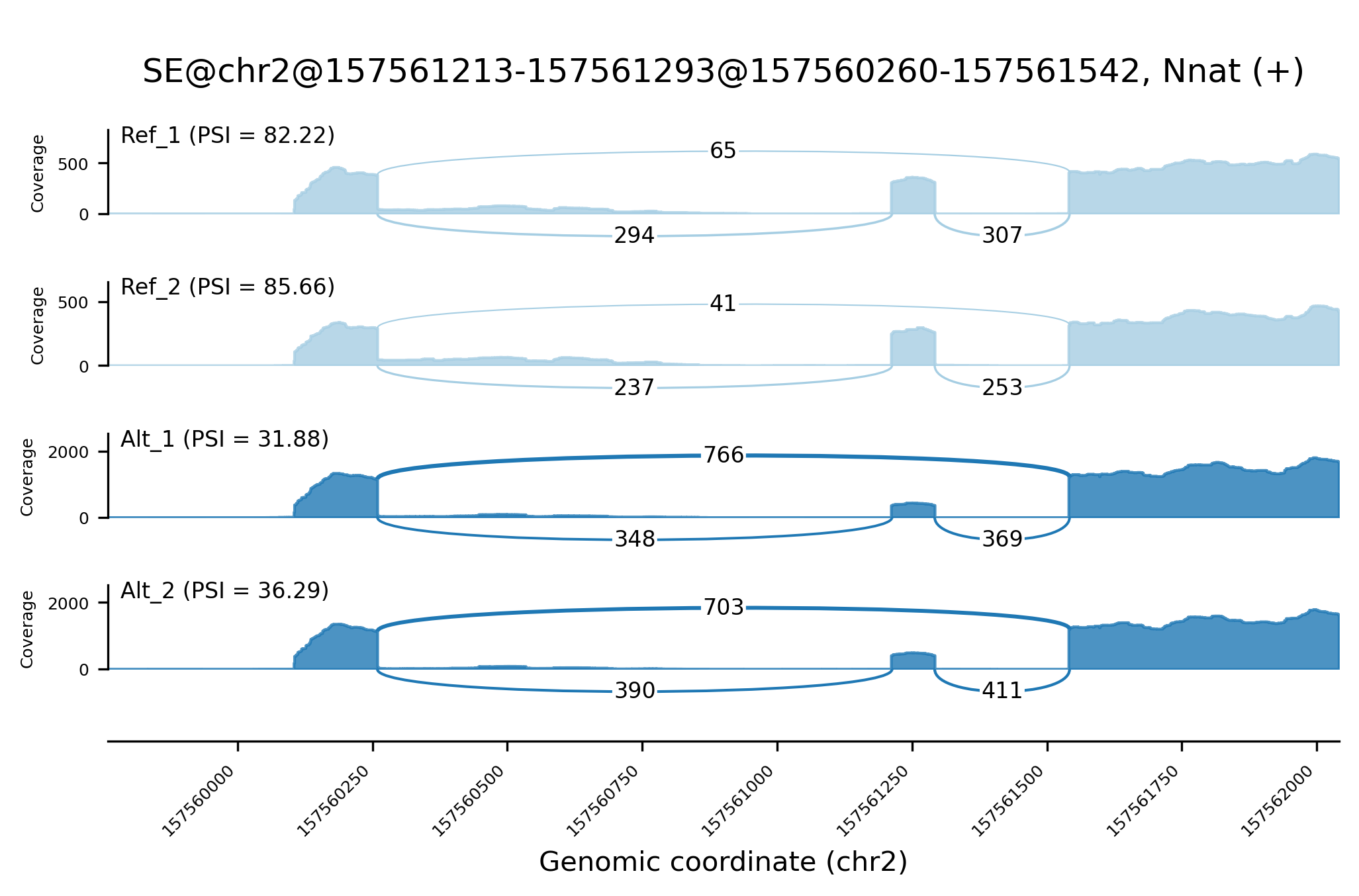
A utility to create Sashimi plots, a publication-quality visualization of RNA-seq data, from Shiba output. Greatly inspired by rmats2sashimiplot and MISO's original implementation.
Quick start
shiba2sashimi -e /path/to/Shiba/experiment_table.tsv \
-s /path/to/Shiba/workdir/ -o img/sashimi_example.png \
--id "SE@chr2@157561213-157561293@157560260-157561542"
How to install
pip
pip install shiba2sashimi
or
git clone https://github.com/Sika-Zheng-Lab/shiba2sashimi.git
cd shiba2sashimi
pip install .
conda
conda create -n shiba2sashimi -c bioconda -c conda-forge shiba2sashimi
conda activate shiba2sashimi
Docker
docker pull naotokubota/shiba2sashimi
Dependencies
- python (>=3.9)
- numpy (>=1.18.0,<2.0.0)
- matplotlib (>=3.1.0)
- pysam (>=0.22.0)
Usage
usage: shiba2sashimi [-h] -e EXPERIMENT -s SHIBA -o OUTPUT [--id ID] [-c COORDINATE] [--samples SAMPLES] [--groups GROUPS] [--colors COLORS] [--extend_up EXTEND_UP] [--extend_down EXTEND_DOWN]
[--smoothing_window_size SMOOTHING_WINDOW_SIZE] [--font_family FONT_FAMILY] [--dpi DPI] [-v]
shiba2sashimi v0.1.5 - Create Sashimi plot from Shiba output
optional arguments:
-h, --help show this help message and exit
-e EXPERIMENT, --experiment EXPERIMENT
Experiment table used for Shiba
-s SHIBA, --shiba SHIBA
Shiba working directory
-o OUTPUT, --output OUTPUT
Output file
--id ID Positional ID (pos_id) of the event to plot
-c COORDINATE, --coordinate COORDINATE
Coordinates of the region to plot
--samples SAMPLES Samples to plot. e.g. sample1,sample2,sample3 Default: all samples in the experiment table
--groups GROUPS Groups to plot. e.g. group1,group2,group3 Default: all groups in the experiment table. Overrides --samples
--colors COLORS Colors for each group. e.g. red,orange,blue
--extend_up EXTEND_UP
Extend the plot upstream. Only used when not providing coordinates. Default: 500
--extend_down EXTEND_DOWN
Extend the plot downstream. Only used when not providing coordinates. Default: 500
--smoothing_window_size SMOOTHING_WINDOW_SIZE
Window size for median filter to smooth coverage plot. Greater value gives smoother plot. Default: 21
--font_family FONT_FAMILY
Font family for labels
--dpi DPI DPI of the output figure. Default: 300
-v, --verbose Increase verbosity
Contributing
Thank you for wanting to improve shiba2sashimi! If you have any bugs or questions, feel free to open an issue or pull request.
Citation
If you use shiba2sashimi in your research, please cite the original Shiba paper:
Kubota N, Chen L, Zheng S. Shiba: a versatile computational method for systematic identification of differential RNA splicing across platforms. Nucleic Acids Research 53(4), 2025, gkaf098.
Authors
- Naoto Kubota (0000-0003-0612-2300)
- Liang Chen (0000-0001-6164-4553)
- Sika Zheng (0000-0002-0573-4981)
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file shiba2sashimi-0.1.5.tar.gz.
File metadata
- Download URL: shiba2sashimi-0.1.5.tar.gz
- Upload date:
- Size: 13.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.22
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5ae7255d56a3aa440a6bd26593a35baacf04a36469bf84309ed750597efa5672
|
|
| MD5 |
7897744c4fdafbe6c337a7985bca1f3d
|
|
| BLAKE2b-256 |
7946bb406b35043b897e48664d2e1ac328d52b5dd5e1f78b6546022e9402b649
|
File details
Details for the file shiba2sashimi-0.1.5-py3-none-any.whl.
File metadata
- Download URL: shiba2sashimi-0.1.5-py3-none-any.whl
- Upload date:
- Size: 14.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.22
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a7e2ec5b1a4d89fd4270d8a883667f6f8b2a058c5062cf7d859b9ba7b44a27c8
|
|
| MD5 |
160b03e6a0d3196a66972e18baa58128
|
|
| BLAKE2b-256 |
1997986c765072423c88984df82ab44e2d48a5e50cf1c63f9ac184feddd17307
|