Specatalog is a software framework for the systematic management and archival of spectroscopic measurement data, integrating a relational metadata database with structured data storage and analysis workflows.
Project description

Specatalog
Management of spectroscopic measurement data
Overview
Specatalog is a software tool for the systematic management and archival of spectroscopic measurement data in research environments. It combines a structured directory-based data storage system with a relational database (PostgreSQL) for metadata management.
The software is designed to ensure reproducibility, traceability, and long-term accessibility of experimental data. In addition to raw data storage, Specatalog supports the integration of processed data and evaluation results within a unified framework.
Scope and Supported Methods
Specatalog currently supports the following spectroscopic techniques:
- Time-resolved EPR (trEPR)
- Continuous-wave EPR (cwEPR)
- Pulsed EPR (pulseEPR)
- UV/Vis spectroscopy
- Fluorescence spectroscopy
- Transient absorption (TA)
The system is designed to be extensible, allowing the integration of additional spectroscopic methods. Details on extending the data models are provided in the documentation.
Key Features
- Structured archival system for spectroscopic data
- PostgreSQL-based metadata database
- Integration of raw data, processed data, and evaluation results
- Storage of analysis results within HDF5 files
- Loaders for common spectroscopic data formats
- Programmatic access via Python script
- Graphical user interface (GUI) for interactive data management
- Synchronization tools for remote data storage
A central aspect of Specatalog is the tight coupling between metadata and data storage, enabling consistent and reproducible data handling across projects.
Graphical User Interface
The graphical user interface provides tools for interactive data management and inspection.
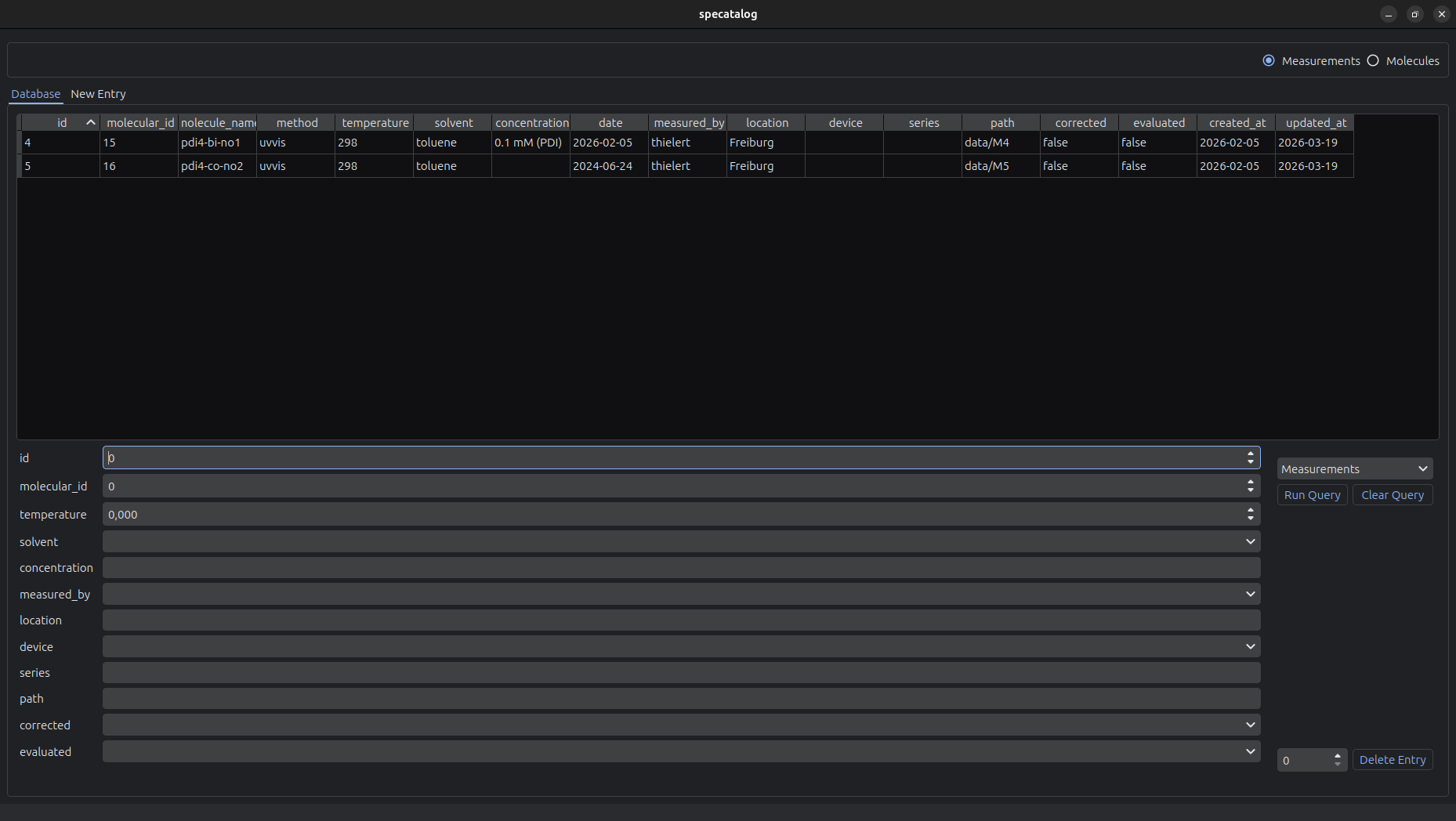
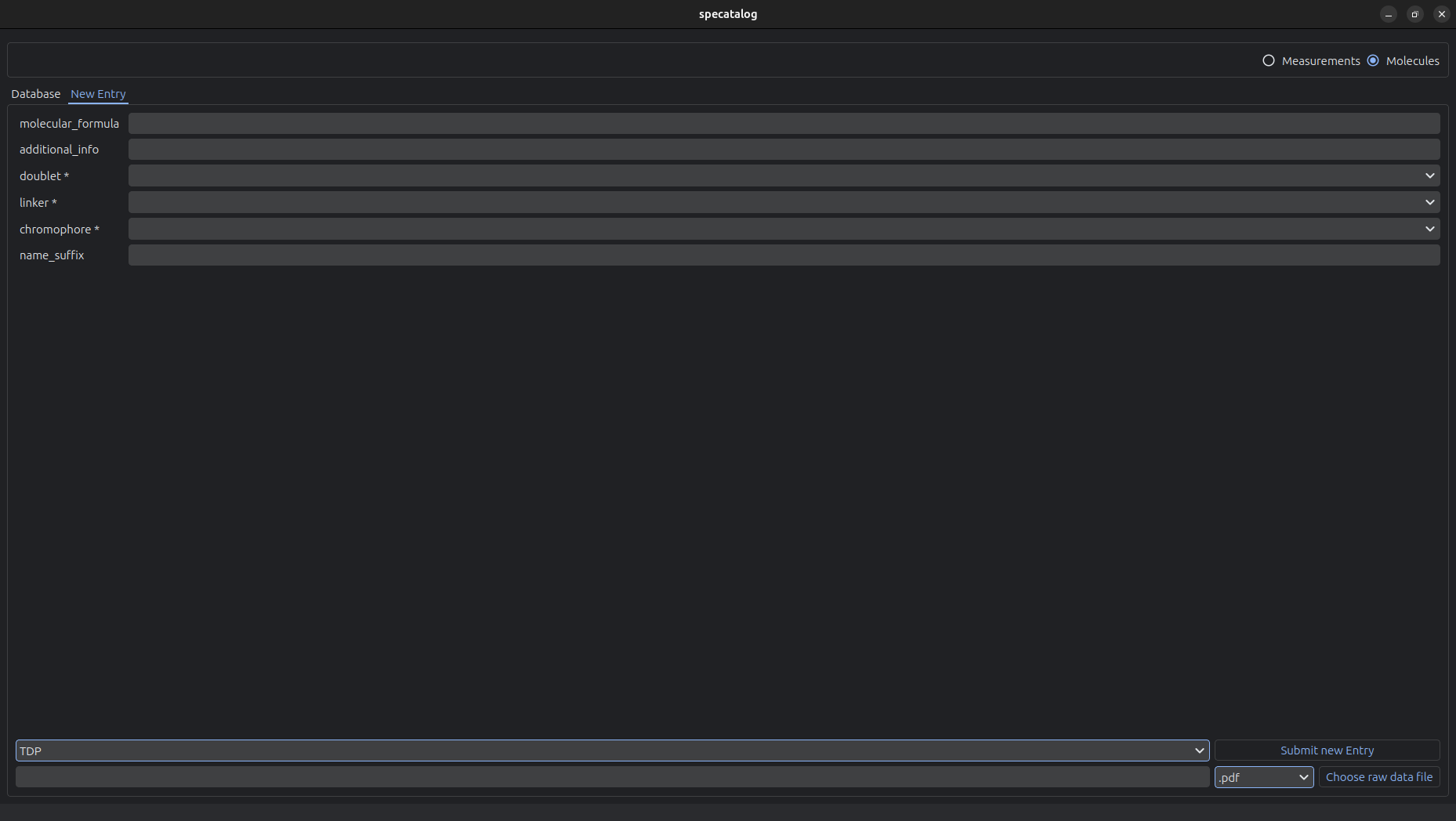
The GUI supports:
- Browsing and searching the database
- Tabular display of entries
- Editing and deletion of existing entries
- Creation of new entries
- Upload of raw measurement data into the archive
Installation
Clone the repository and install the package locally:
git clone https://github.com/TheresiaQuintes/specatalog.git
cd specatalog
pip install .
All dependencies are specified in the pyproject.toml file.
Requirements
- Python ≥ 3.11
- PostgreSQL database
- Operating systems: Linux, macOS, Windows
Command Line Interface
Specatalog provides several command-line tools:
-
specatalog-welcomeVerifies installation and prints the current working directory -
specatalog-configurationInitial configuration of the system -
specatalog-guiLaunches the graphical user interface -
specatalog-sync-download,specatalog-sync-uploadSynchronization with remote directories -
specatalog-update-dbUpdates database models
Data Model and Storage Concept
Specatalog separates metadata and data storage while maintaining a strict linkage between both.
Directory Structure
base_dir/
├── data/
│ ├── M1/
│ ├── M2/
│ ├── ...
│ └── M{ms_id}/
│ ├── additional_info/
│ ├── figures/
│ ├── literature/
│ ├── raw/
│ ├── scripts/
│ └── measurement_M{ms_id}.h5
├── molecules/
│ ├── MOL1/
│ ├── MOL2/
│ ├── ...
│ └── MOL{mol_id}/
│ ├── file_with_structure.cdxml
│ └── file_with_structure.pdf
└── allowed_values.py
HDF5 Data Structure
Each measurement is associated with a structured HDF5 file:
measurement_{ms_id}.h5
├── raw_data
│ ├── data
│ ├── data_imag
│ ├── data_real
│ └── xaxis
├── corrected_data
│ └── <user-defined datasets>
└── evaluations
└── <user-defined datasets>
This structure allows storing both raw data and derived results in a consistent and extendable format.
Example Usage
Creating a New Measurement Entry
from specatalog.models import creation_pydantic_measurements as ms
from datetime import date
from specatalog.helpers.full_entry import create_full_measurement
new_measurement = ms.TREPRModel(
molecular_id=1,
temperature=80,
solvent="toluene",
date=date(2025, 12, 24),
measured_by="your_name",
device="ELEXSYS",
frequenc_band="Q",
attenuation="20dB",
exciation_wl=530
)
fm = create_full_measurement(new_measurement, BASE_PATH, raw_data, "bruker_bes3t")
print(fm.success)
print(fm.measurement_id)
Querying the Database
from specatalog.crud_db import read as r
filter_model = r.TREPRFilter(
molecular_id=1,
temperature__le=80,
measured_by="your name"
)
results = r.run_query(filter_model, ordering_model)
print(results)
print(results[0].id)
print(results[0].molecule.name)
Working with HDF5 Data
from specatalog.data_management.hdf5_reader import load_from_id
import matplotlib.pyplot as plt
import numpy as np
dat, file = load_from_id(1, mode="a")
x = dat.raw_data.xaxis
intensity = dat.raw_data.data
plt.plot(x, np.real(intensity))
offset = 2.3
corrected = x - offset
fit = -2 * (x - 12000)**2 + 2500
dat.corrected_data.set_attr("x-offset", 2.3)
dat.corrected_data.set_dataset("xaxis", corrected)
dat.evaluations.set_dataset("fit1", fit)
Documentation
Comprehensive documentation, including installation details and instructions for extending the data models, is available at:
https://theresiaquintes.github.io/specatalog/
Intended Use
Specatalog is intended for the archival and management of spectroscopic data in academic and industrial research environments. It is particularly suited for laboratories that require structured data storage, standardized metadata handling, and reproducible analysis workflows.
License
This project is licensed under the GNU General Public License v3.0 (GPL-3.0).
Contributing
Contributions, issue reports, and feature requests are welcome.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file specatalog-1.0.0.tar.gz.
File metadata
- Download URL: specatalog-1.0.0.tar.gz
- Upload date:
- Size: 86.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8a47ea4eb7d5a9c40a2959b95cbe488fbb3252bdd370304a94f4630f30774caa
|
|
| MD5 |
138a002fb8e7c0d58e912e52046f8563
|
|
| BLAKE2b-256 |
4cb35a2a0bdf1456b6f0431e749b132f048d94cadf4945b914c2c8c1c8b7abba
|
File details
Details for the file specatalog-1.0.0-py3-none-any.whl.
File metadata
- Download URL: specatalog-1.0.0-py3-none-any.whl
- Upload date:
- Size: 83.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
199ad5561da389b3c0b3f3a0bc418aae4e86429a8f1b7aa1884935529812e84f
|
|
| MD5 |
9ec9775cf479a90240574708d712a990
|
|
| BLAKE2b-256 |
575fc7b3a39e17cc86636bd33a5a123fcdea419065dd3b11b3ac129da4e52172
|










