Spatial Single Cell Analysis in Python
Project description
Squidpy - Spatial Single Cell Analysis in Python
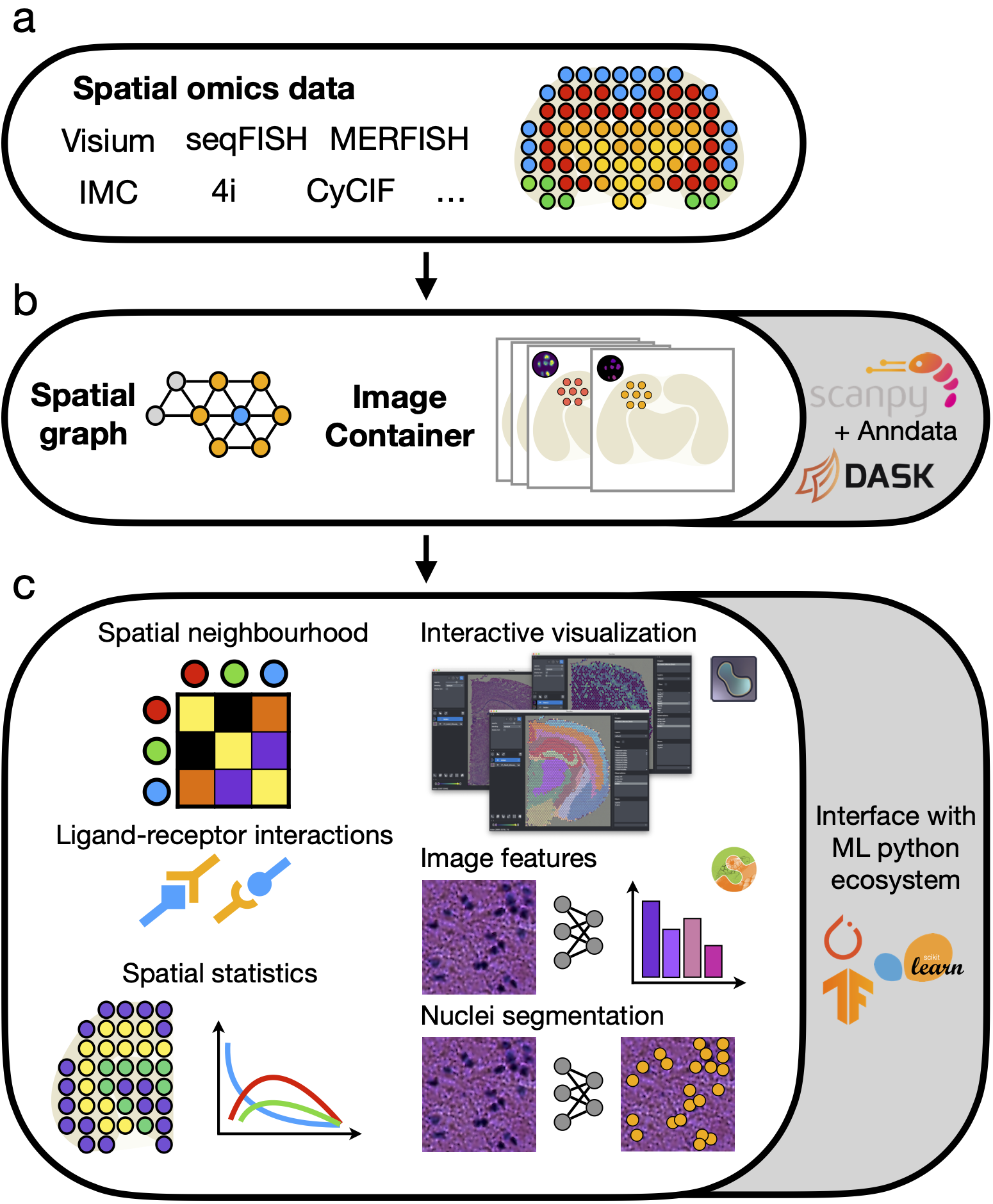
Squidpy is the scverse toolkit for scalable analysis and visualization of spatial molecular data. It builds on scanpy and anndata, providing streamlined APIs for feature extraction, spatial statistics, and interactive exploration of tissue sections together with microscopy images.
Documentation
Head over to the documentation for installation instructions, tutorials, how-to guides, and reference material.
Installation
We recommend running Squidpy on a recent Linux or macOS system with Python ≥3.11, but it also works on Windows via WSL.
Install from PyPI with:
pip install squidpy
or from conda-forge:
conda install -c conda-forge squidpy
Interactive visualization
To get optional dependencies required for the napari-based interactive plotting APIs, install the interactive extra:
pip install 'squidpy[interactive]'
Key capabilities
- Build and analyze spatial neighbor graphs directly from Visium, Slide-seq, Xenium, and other spatial omics assays.
- Compute spatial statistics for cell types and genes, including neighborhood enrichment, co-occurrence, and Moran's I.
- Efficiently store, featurize, and visualize high-resolution tissue microscopy images via scikit-image.
- Explore annotated datasets interactively with napari and scverse visualization tooling.
Contributing
Contributions are welcome! Please read the contributing guide for instructions on setting up your environment, running tests, and submitting pull requests.
Citation
If you use Squidpy in your research, cite the original publication:
@article{palla:22,
author = {Palla, Giovanni and Spitzer, Hannah and Klein, Michal and Fischer, David and Schaar, Anna Christina
and Kuemmerle, Louis Benedikt and Rybakov, Sergei and Ibarra, Ignacio L. and Holmberg, Olle
and Virshup, Isaac and Lotfollahi, Mohammad and Richter, Sabrina and Theis, Fabian J.},
title = {Squidpy: a scalable framework for spatial omics analysis},
journal = {Nature Methods},
year = {2022},
month = {Feb},
volume = {19},
number = {2},
pages = {171--178},
issn = {1548-7105},
doi = {10.1038/s41592-021-01358-2},
}
Squidpy is part of the scverse® project (website, governance) and is fiscally sponsored by NumFOCUS. Please consider making a tax-deductible donation to help the project pay for developer time, professional services, travel, workshops, and a variety of other needs.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file squidpy-1.8.1.tar.gz.
File metadata
- Download URL: squidpy-1.8.1.tar.gz
- Upload date:
- Size: 6.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d692581ffe1d897605d0825eb40b786482bdf16a4c5f7d5e71ebe3fb46dfc7fd
|
|
| MD5 |
68c47d090006617f4ef2b78a29b5c1e1
|
|
| BLAKE2b-256 |
dc76e2b54dc1859c7779b49fa383db52a8b823d1f4d8576ee1ac317f24307678
|
Provenance
The following attestation bundles were made for squidpy-1.8.1.tar.gz:
Publisher:
release.yaml on scverse/squidpy
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
squidpy-1.8.1.tar.gz -
Subject digest:
d692581ffe1d897605d0825eb40b786482bdf16a4c5f7d5e71ebe3fb46dfc7fd - Sigstore transparency entry: 930419251
- Sigstore integration time:
-
Permalink:
scverse/squidpy@4a018a373206b693b229ed50ab8a3fc75e7497bb -
Branch / Tag:
refs/tags/v1.8.1 - Owner: https://github.com/scverse
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yaml@4a018a373206b693b229ed50ab8a3fc75e7497bb -
Trigger Event:
release
-
Statement type:
File details
Details for the file squidpy-1.8.1-py3-none-any.whl.
File metadata
- Download URL: squidpy-1.8.1-py3-none-any.whl
- Upload date:
- Size: 193.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6ce56c96d640c719f566311af83ff5b91731a7020330499537f007e47e916fe3
|
|
| MD5 |
7f1cbb02d0d6f20c99384c22cb5cde36
|
|
| BLAKE2b-256 |
4a9981e01d2df1e0ce4e1b2f373527eddaabe3eac5a5ffc18bf3f3c4a2960f2c
|
Provenance
The following attestation bundles were made for squidpy-1.8.1-py3-none-any.whl:
Publisher:
release.yaml on scverse/squidpy
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
squidpy-1.8.1-py3-none-any.whl -
Subject digest:
6ce56c96d640c719f566311af83ff5b91731a7020330499537f007e47e916fe3 - Sigstore transparency entry: 930419253
- Sigstore integration time:
-
Permalink:
scverse/squidpy@4a018a373206b693b229ed50ab8a3fc75e7497bb -
Branch / Tag:
refs/tags/v1.8.1 - Owner: https://github.com/scverse
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yaml@4a018a373206b693b229ed50ab8a3fc75e7497bb -
Trigger Event:
release
-
Statement type: