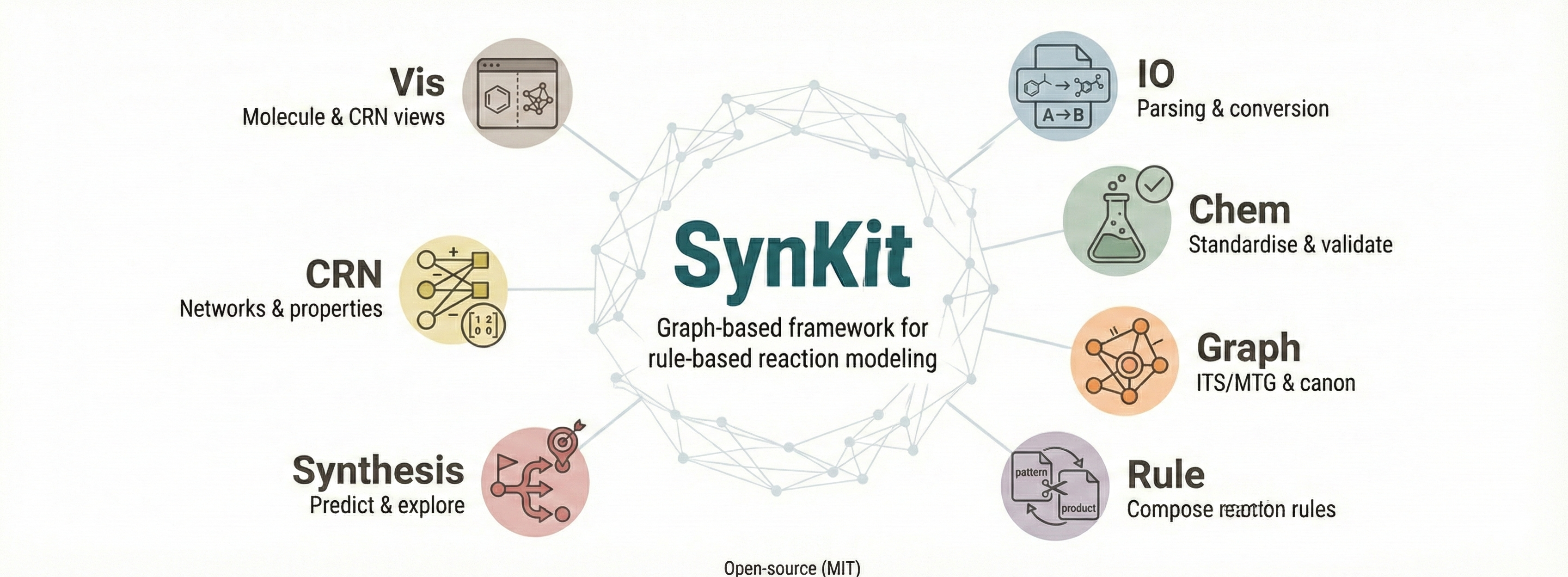
Utility for reaction modeling using graph grammar
Project description
SynKit
Toolkit for Synthesis Planning
SynKit is a collection of tools designed to support the planning and execution of chemical synthesis.
Our tools are tailored to assist researchers and chemists in navigating complex chemical reactions and synthesis pathways, leveraging the power of modern computational chemistry. Whether you're designing novel compounds or optimizing existing processes, synkit aims to provide the critical tools you need.
For more details on each utility within the repository, please refer to the documentation provided in the respective folders.
Table of Contents
Installation
-
Python Installation: Ensure that Python 3.11 or later is installed on your system. You can download it from python.org.
-
Creating a Virtual Environment (Optional but Recommended): It's recommended to use a virtual environment to avoid conflicts with other projects or system-wide packages. Use the following commands to create and activate a virtual environment:
python -m venv synkit-env
source synkit-env/bin/activate
Or Conda
conda create --name synkit-env python=3.11
conda activate synkit-env
- Install from PyPi: The easiest way to use SynTemp is by installing the PyPI package synkit.
pip install synkit
Optional if you want to install full version
pip install synkit[all]
-
Install via Docker
Pull the image:docker pull tieulongphan/synkit:latest # or a specific version: docker pull tieulongphan/synkit:0.1.0
Run a container (sanity check):
docker run --rm tieulongphan/synkit:latest
Contribute
We're welcoming new contributors to build this project better. Please not hesitate to inquire me via [email][tieu@bioinf.uni-leipzig.de].
Before you start, ensure your local development environment is set up correctly. Pull the latest version of the main branch to start with the most recent stable code.
git checkout main
git pull
Working on New Features
-
Create a New Branch:
For every new feature or bug fix, create a new branch from themainbranch. Name your branch meaningfully, related to the feature or fix you are working on.git checkout -b feature/your-feature-name
-
Develop and Commit Changes:
Make your changes locally, commit them to your branch. Keep your commits small and focused; each should represent a logical unit of work.git commit -m "Describe the change"
-
Run Quality Checks:
Before finalizing your feature, run the following commands to ensure your code meets our formatting standards and passes all tests:./lint.sh # Check code format pytest Test # Run tests
Fix any issues or errors highlighted by these checks.
Integrating Changes
-
Rebase onto Staging:
Once your feature is complete and tests pass, rebase your changes onto thestagingbranch to prepare for integration.git fetch origin git rebase origin/staging
Carefully resolve any conflicts that arise during the rebase.
-
Push to Your Feature Branch: After successfully rebasing, push your branch to the remote repository.
git push origin feature/your-feature-name
-
Create a Pull Request: Open a pull request from your feature branch to the
stagingbranch. Ensure the pull request description clearly describes the changes and any additional context necessary for review.
Contributing
Publication
License
This project is licensed under MIT License - see the License file for details.
Acknowledgments
This project has received funding from the European Unions Horizon Europe Doctoral Network programme under the Marie-Skłodowska-Curie grant agreement No 101072930 (TACsy -- Training Alliance for Computational)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file synkit-0.0.13.tar.gz.
File metadata
- Download URL: synkit-0.0.13.tar.gz
- Upload date:
- Size: 2.2 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7224879466ce975a4f8ee3aee06e2b41b9fc1e763b1ab20fcf75959e89b06ced
|
|
| MD5 |
ce35c1795c47952a3c207da4af324fc7
|
|
| BLAKE2b-256 |
3958f44349cefeb89ed9adec604c793677e2aac5d91a53511a9295e1df4c49e6
|
Provenance
The following attestation bundles were made for synkit-0.0.13.tar.gz:
Publisher:
publish-package.yml on TieuLongPhan/SynKit
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
synkit-0.0.13.tar.gz -
Subject digest:
7224879466ce975a4f8ee3aee06e2b41b9fc1e763b1ab20fcf75959e89b06ced - Sigstore transparency entry: 312239970
- Sigstore integration time:
-
Permalink:
TieuLongPhan/SynKit@dc1ecc0177ba6f6c09fd30dc8b552184c9eaa5b3 -
Branch / Tag:
refs/tags/v0.0.13 - Owner: https://github.com/TieuLongPhan
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-package.yml@dc1ecc0177ba6f6c09fd30dc8b552184c9eaa5b3 -
Trigger Event:
release
-
Statement type:
File details
Details for the file synkit-0.0.13-py3-none-any.whl.
File metadata
- Download URL: synkit-0.0.13-py3-none-any.whl
- Upload date:
- Size: 265.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a6ad535a2101580cf89095585ea083bd404dfed333cdb12d4d58d16acfdbc773
|
|
| MD5 |
c45143c27d00ad885a09a22df98d3dea
|
|
| BLAKE2b-256 |
44c5bb9be1d628bf0e2111e3b252a3139f6ddd6502605b40dabb144f42427f9c
|
Provenance
The following attestation bundles were made for synkit-0.0.13-py3-none-any.whl:
Publisher:
publish-package.yml on TieuLongPhan/SynKit
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
synkit-0.0.13-py3-none-any.whl -
Subject digest:
a6ad535a2101580cf89095585ea083bd404dfed333cdb12d4d58d16acfdbc773 - Sigstore transparency entry: 312239983
- Sigstore integration time:
-
Permalink:
TieuLongPhan/SynKit@dc1ecc0177ba6f6c09fd30dc8b552184c9eaa5b3 -
Branch / Tag:
refs/tags/v0.0.13 - Owner: https://github.com/TieuLongPhan
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-package.yml@dc1ecc0177ba6f6c09fd30dc8b552184c9eaa5b3 -
Trigger Event:
release
-
Statement type: