Generated from aind-library-template
Project description
aind-analysis-arch-result-access
APIs to access analysis results in the AIND behavior pipeline.
Installation
pip install aind-analysis-arch-result-access
Usage
Access pipeline v1.0 (Han's pipeline)
Fetch the session master table in Streamlit
from aind_analysis_arch_result_access.han_pipeline import get_session_table
df_master = get_session_table(if_load_bpod=False) # `if_load_bpod=True` will load additional 4000+ old sessions from bpod
Fetch logistic regression results
- Get logistic regression results from one session
from aind_analysis_arch_result_access.han_pipeline import get_logistic_regression df_logistic = get_logistic_regression( df_sessions=pd.DataFrame( { "subject_id": ["769253"], "session_date": ["2025-03-12"], } ), model="Su2022", )
- Get logistic regression results in batch (from any dataframe with
subject_idandsession_datecolumns)df_logistic = get_logistic_regression( df_master.query("subject_id == '769253'"), # All sessions from a single subject (query from the `df_master` above) model="Su2022", if_download_figures=True, # Also download fitting plots download_path="./tmp", )
Fetch trial table (🚧 under development)
Fetch analysis figures (🚧 under development)
Access pipeline v2.0 (AIND analysis architecture)
Fetch dynamic foraging MLE model fitting results
-
Get all MLE fitting results from one session
from aind_analysis_arch_result_access.han_pipeline import get_mle_model_fitting df = get_mle_model_fitting(subject_id="730945", session_date="2024-10-24") print(df.columns) print(df[["agent_alias", "AIC", "prediction_accuracy_10-CV_test"]])
output
Query: {'analysis_spec.analysis_name': 'MLE fitting', 'analysis_spec.analysis_ver': 'first version @ 0.10.0', 'subject_id': '730945', 'session_date': '2024-10-24'} Found 5 MLE fitting records! Found 5 successful MLE fitting! Get latent variables from s3: 100%|██████████| 5/5 [00:00<00:00, 58.01it/s] Index(['_id', 'nwb_name', 'status', 'agent_alias', 'log_likelihood', 'AIC', 'BIC', 'LPT', 'LPT_AIC', 'LPT_BIC', 'k_model', 'n_trials', 'prediction_accuracy', 'prediction_accuracy_test', 'prediction_accuracy_fit', 'prediction_accuracy_test_bias_only', 'params', 'prediction_accuracy_10-CV_test', 'prediction_accuracy_10-CV_test_std', 'prediction_accuracy_10-CV_fit', 'prediction_accuracy_10-CV_fit_std', 'prediction_accuracy_10-CV_test_bias_only', 'prediction_accuracy_10-CV_test_bias_only_std', 'latent_variables'], dtype='object') agent_alias AIC prediction_accuracy_10-CV_test 0 QLearning_L1F1_CK1_softmax 239.519051 0.898151 1 QLearning_L1F0_epsi 403.621460 0.762075 2 QLearning_L2F1_CK1_softmax 236.265381 0.903280 3 WSLS 4051.958064 0.636196 4 QLearning_L2F1_softmax 236.512476 0.888611Now the latent variables also contain the
rpe.df.latent_variables.iloc[0].keys()
output
dict_keys(['q_value', 'choice_kernel', 'choice_prob', 'rpe']) -
Also download figures
df = get_mle_model_fitting( subject_id="730945", session_date="2024-10-24", if_download_figures=True, download_path="./mle_figures", ) !ls ./mle_figures
output
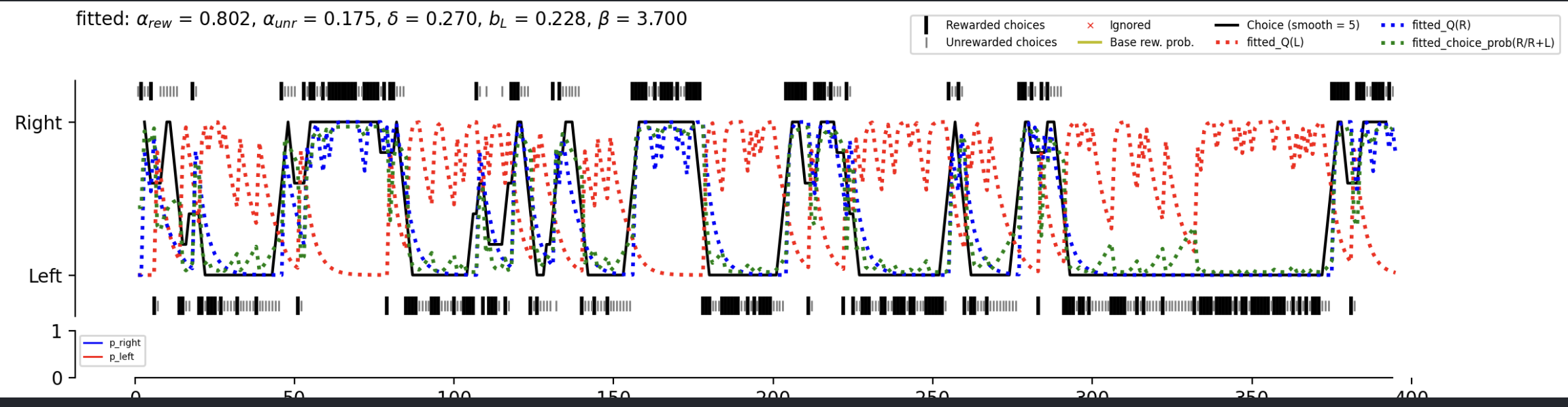
Query: {'analysis_spec.analysis_name': 'MLE fitting', 'analysis_spec.analysis_ver': 'first version @ 0.10.0', 'subject_id': '730945', 'session_date': '2024-10-24'} Found 5 MLE fitting records! Found 5 successful MLE fitting! Get latent variables from s3: 100%|██████████| 5/5 [00:00<00:00, 85.87it/s] Download figures from s3: 100%|██████████| 5/5 [00:00<00:00, 86.45it/s] 730945_2024-10-24_17-38-06_QLearning_L1F0_epsi_58cc5b6f6e.png 730945_2024-10-24_17-38-06_QLearning_L1F1_CK1_softmax_3ffdf98012.png 730945_2024-10-24_17-38-06_QLearning_L2F1_CK1_softmax_5ce7f1f816.png 730945_2024-10-24_17-38-06_QLearning_L2F1_softmax_ec59be40c0.png 730945_2024-10-24_17-38-06_WSLS_7c61d01e0f.pngExample figure:
-
Get fittings from all sessions of a mouse for a specific model
df = get_mle_model_fitting( subject_id="730945", agent_alias="QLearning_L2F1_CK1_softmax", if_download_figures=False, ) print(df.iloc[:10][["nwb_name", "agent_alias"]])
output
Query: {'analysis_spec.analysis_name': 'MLE fitting', 'analysis_spec.analysis_ver': 'first version @ 0.10.0', 'subject_id': '730945', 'analysis_results.fit_settings.agent_alias': 'QLearning_L2F1_CK1_softmax'} Found 32 MLE fitting records! Found 32 successful MLE fitting! Get latent variables from s3: 100%|██████████| 32/32 [00:00<00:00, 80.81it/s] nwb_name agent_alias 0 730945_2024-08-27_16-07-16.nwb QLearning_L2F1_CK1_softmax 1 730945_2024-09-05_16-47-58.nwb QLearning_L2F1_CK1_softmax 2 730945_2024-10-23_15-33-07.nwb QLearning_L2F1_CK1_softmax 3 730945_2024-09-19_17-26-54.nwb QLearning_L2F1_CK1_softmax 4 730945_2024-09-04_16-04-38.nwb QLearning_L2F1_CK1_softmax 5 730945_2024-08-30_15-55-05.nwb QLearning_L2F1_CK1_softmax 6 730945_2024-08-29_15-50-57.nwb QLearning_L2F1_CK1_softmax 7 730945_2024-10-24_17-38-06.nwb QLearning_L2F1_CK1_softmax 8 730945_2024-09-12_17-21-58.nwb QLearning_L2F1_CK1_softmax 9 730945_2024-09-03_15-49-53.nwb QLearning_L2F1_CK1_softmax -
(for advanced users) Use your own docDB query
df = get_mle_model_fitting( from_custom_query={ "analysis_results.fit_settings.agent_alias": "QLearning_L2F1_CK1_softmax", "analysis_results.n_trials" : {"$gt": 600}, }, if_include_latent_variables=False, if_download_figures=False, )
output
Query: {'analysis_spec.analysis_name': 'MLE fitting', 'analysis_spec.analysis_ver': 'first version @ 0.10.0', 'analysis_results.fit_settings.agent_alias': 'QLearning_L2F1_CK1_softmax', 'analysis_results.n_trials': {'$gt': 600}} Found 807 MLE fitting records! Found 807 successful MLE fitting!
Contributing
Installation
To use the software, in the root directory, run
pip install -e .
To develop the code, run
pip install -e .[dev]
Linters and testing
There are several libraries used to run linters, check documentation, and run tests.
- Please test your changes using the coverage library, which will run the tests and log a coverage report:
coverage run -m unittest discover && coverage report
- Use interrogate to check that modules, methods, etc. have been documented thoroughly:
interrogate .
- Use flake8 to check that code is up to standards (no unused imports, etc.):
flake8 .
- Use black to automatically format the code into PEP standards:
black .
- Use isort to automatically sort import statements:
isort .
Pull requests
For internal members, please create a branch. For external members, please fork the repository and open a pull request from the fork. We'll primarily use Angular style for commit messages. Roughly, they should follow the pattern:
<type>(<scope>): <short summary>
where scope (optional) describes the packages affected by the code changes and type (mandatory) is one of:
- build: Changes that affect build tools or external dependencies (example scopes: pyproject.toml, setup.py)
- ci: Changes to our CI configuration files and scripts (examples: .github/workflows/ci.yml)
- docs: Documentation only changes
- feat: A new feature
- fix: A bugfix
- perf: A code change that improves performance
- refactor: A code change that neither fixes a bug nor adds a feature
- test: Adding missing tests or correcting existing tests
Semantic Release
The table below, from semantic release, shows which commit message gets you which release type when semantic-release runs (using the default configuration):
| Commit message | Release type |
|---|---|
fix(pencil): stop graphite breaking when too much pressure applied |
|
feat(pencil): add 'graphiteWidth' option |
|
perf(pencil): remove graphiteWidth optionBREAKING CHANGE: The graphiteWidth option has been removed.The default graphite width of 10mm is always used for performance reasons. |
(Note that the BREAKING CHANGE: token must be in the footer of the commit) |
Documentation
To generate the rst files source files for documentation, run
sphinx-apidoc -o docs/source/ src
Then to create the documentation HTML files, run
sphinx-build -b html docs/source/ docs/build/html
More info on sphinx installation can be found here.
Read the Docs Deployment
Note: Private repositories require Read the Docs for Business account. The following instructions are for a public repo.
The following are required to import and build documentations on Read the Docs:
- A Read the Docs user account connected to Github. See here for more details.
- Read the Docs needs elevated permissions to perform certain operations that ensure that the workflow is as smooth as possible, like installing webhooks. If you are not the owner of the repo, you may have to request elevated permissions from the owner/admin.
- A .readthedocs.yaml file in the root directory of the repo. Here is a basic template:
# Read the Docs configuration file
# See https://docs.readthedocs.io/en/stable/config-file/v2.html for details
# Required
version: 2
# Set the OS, Python version, and other tools you might need
build:
os: ubuntu-24.04
tools:
python: "3.13"
# Path to a Sphinx configuration file.
sphinx:
configuration: docs/source/conf.py
# Declare the Python requirements required to build your documentation
python:
install:
- method: pip
path: .
extra_requirements:
- dev
Here are the steps for building docs in Read the Docs. See here for detailed instructions:
- From Read the Docs dashboard, click on Add project.
- For automatic configuration, select Configure automatically and type the name of the repo. A repo with public visibility should appear as you type.
- Follow the subsequent steps.
- For manual configuration, select Configure manually and follow the subsequent steps
Once a project is created successfully, you will be able to configure/modify the project's settings; such as Default version, Default branch etc.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file aind_analysis_arch_result_access-0.6.1.tar.gz.
File metadata
- Download URL: aind_analysis_arch_result_access-0.6.1.tar.gz
- Upload date:
- Size: 26.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0a541f2913f1c5678e3145de29622ee2fc4c696e67bff7f1ccd42dd03e55ea21
|
|
| MD5 |
483f406e25396fcc4237e3083eb41a0b
|
|
| BLAKE2b-256 |
8a47b0f97b862c64e394351a52552ce761cfe15ac13974e17fba10d23b66e40a
|
File details
Details for the file aind_analysis_arch_result_access-0.6.1-py3-none-any.whl.
File metadata
- Download URL: aind_analysis_arch_result_access-0.6.1-py3-none-any.whl
- Upload date:
- Size: 17.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c18ff46459845c8d5059c32ae7399e0e25924f156add85d42d5b71abc47524a8
|
|
| MD5 |
90f6af5edd202e3e20ae8dcc2f0ffb05
|
|
| BLAKE2b-256 |
bcac839f34f8735236ebfa37dc2b536b343eb1677b283aefa618e2258e8885fd
|