nested sampling post-processing
Project description
- Version:
- 2.11.6
- Homepage:
- Documentation:








anesthetic brings together tools for processing nested sampling chains by leveraging standard scientific python libraries.
TLDR: See the quickstart.
Current functionality includes:
Computation of Bayesian evidences, Kullback-Liebler divergences and Bayesian model dimensionalities.
Marginalised 1d and 2d plots.
Dynamic replaying of nested sampling.
This tool was designed primarily for use with nested sampling outputs, although it can be used for normal MCMC chains.
For an interactive view of a nested sampling run, you can use the anesthetic script.
$ anesthetic <ns file root>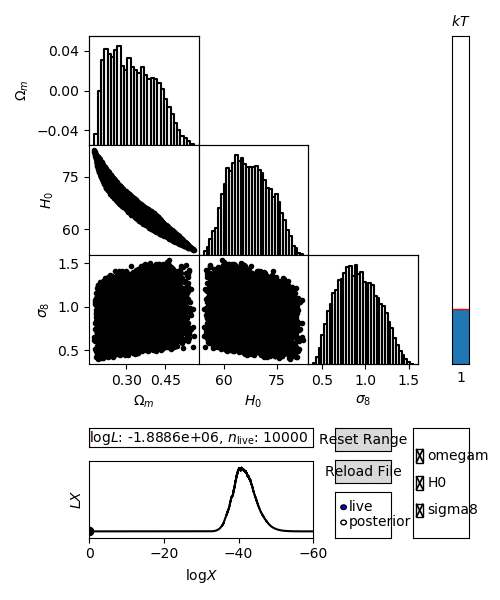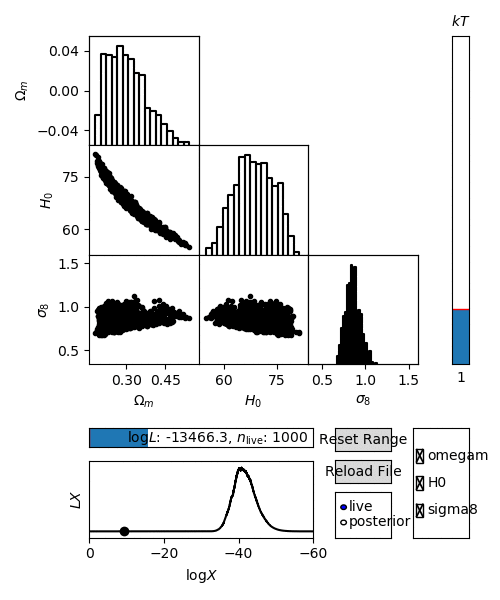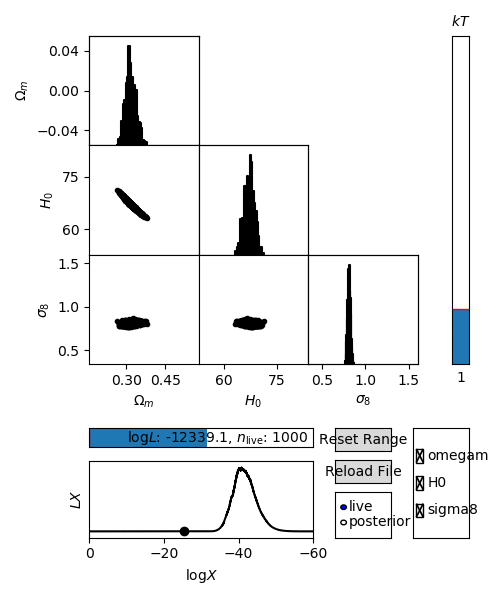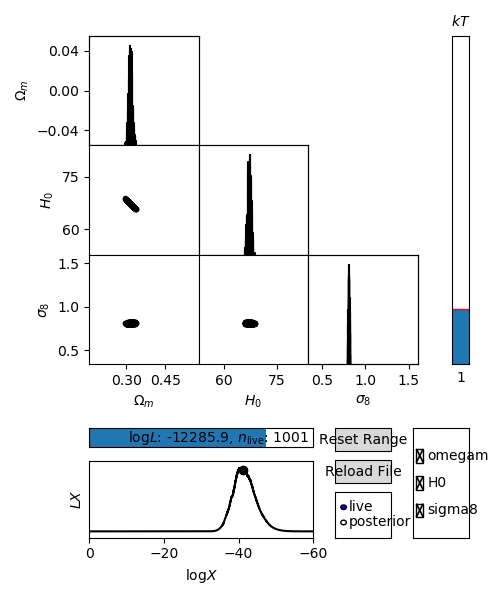
Features
Both samples and plotting axes are stored as a pandas.DataFrame, with parameter names as indices, which makes for easy access and modification.
Sensible color scheme for plotting nearly flat distributions.
For easy extension/modification, uses the standard python libraries: numpy, scipy, matplotlib and pandas.
Installation
anesthetic can be installed via pip
pip install anestheticvia conda
conda install -c handley-lab anestheticor from the github repository
git clone https://github.com/handley-lab/anesthetic
cd anesthetic
python -m pip install .You can check that things are working by running the test suite:
export MPLBACKEND=Agg # only necessary for OSX users
python -m pip install ".[test]"
python -m pytest
flake8 anesthetic tests
pydocstyle --convention=numpy anestheticDependencies
Basic requirements:
Python 3.9+
Documentation:
Tests:
Documentation
Full Documentation is hosted at ReadTheDocs. To build your own local copy of the documentation you’ll need to install sphinx. You can then run:
python -m pip install ".[all,docs]"
cd docs
make htmland view the documentation by opening docs/build/html/index.html in a browser. To regenerate the automatic RST files run:
sphinx-apidoc -fM -t docs/templates/ -o docs/source/ anesthetic/Citation
If you use anesthetic to generate plots for a publication, please cite as:
Handley, (2019). anesthetic: nested sampling visualisation. Journal of Open Source Software, 4(37), 1414, https://doi.org/10.21105/joss.01414
or using the BibTeX:
@article{anesthetic,
doi = {10.21105/joss.01414},
url = {http://dx.doi.org/10.21105/joss.01414},
year = {2019},
month = {Jun},
publisher = {The Open Journal},
volume = {4},
number = {37},
pages = {1414},
author = {Will Handley},
title = {anesthetic: nested sampling visualisation},
journal = {The Journal of Open Source Software}
}Contributing
There are many ways you can contribute via the GitHub repository.
You can open an issue to report bugs or to propose new features.
Pull requests are very welcome. Note that if you are going to propose major changes, be sure to open an issue for discussion first, to make sure that your PR will be accepted before you spend effort coding it.
Questions/Comments
Another posterior plotting tool?
This is my posterior plotter. There are many like it, but this one is mine.
There are several excellent tools for plotting marginalised posteriors:
Why create another one? In general, any dedicated user of software will find that there is some functionality that in their use case is lacking, and the designs of previous codes make such extensions challenging. In my case this was:
For large numbers of samples, kernel density estimation is slow, or inaccurate (particularly for samples generated from nested sampling). There are kernel density estimators, such as fastKDE, which ameliorate many of these difficulties.
Existing tools can make it difficult to define new parameters. For example, the default cosmomc chain defines omegabh2, but not omegab. The transformation is easy, since omegab = omegabh2/ (H0/100)**2, but implementing this transformation in existing packages is not so trivial. anesthetic solves this issue by storing the samples as a pandas array, for which the relevant code for defining the above new parameter would be
from anesthetic import read_chains
samples = read_chains(file_root) # Load the samples
label = 'omegab'
tex = '$\Omega_b$'
h = (samples.H0/100)
samples[(label, tex)] = samples.omegabh2/h**2 # Define omegab
samples.plot_1d('omegab') # Simple 1D plotMany KDE plotting tools have conventions that don’t play well with uniformly distributed parameters, which presents a problem if you are trying to plot priors along with your posteriors. anesthetic has a sensible mechanism, by defining the contours by the amount of iso-probability mass they contain, but colouring the fill in relation to the probability density of the contour.
What’s in a name?
There is a convention for naming nested sampling packages with words that have nest in them (nestle and dynesty, nestorflow). Doing a UNIX grep:
grep nest /usr/share/dict/wordsyields a lot of superlatives (e.g. greenest), but a few other cool names for future projects:
amnesty
defenestrate
dishonestly
inestimable
minestrone
rhinestone
I chose anesthetic because I liked the soft ‘th’, and in spite of the US spelling.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file anesthetic-2.11.6.tar.gz.
File metadata
- Download URL: anesthetic-2.11.6.tar.gz
- Upload date:
- Size: 105.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ec4f6af88f49da2a642eb197fed7df28fb786331b26172c868a1564ed88599ef
|
|
| MD5 |
32eaeaa4f63d6fd645b7875c043e1035
|
|
| BLAKE2b-256 |
786f8f814cea4c8bac3c721e6d0c089bf685ffd66ec2c5b3495a9a7db4f93092
|
File details
Details for the file anesthetic-2.11.6-py3-none-any.whl.
File metadata
- Download URL: anesthetic-2.11.6-py3-none-any.whl
- Upload date:
- Size: 79.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f781f6aa29181a4f85e6faebf0c0f3adda0055d57be9251ee66af09de71f822d
|
|
| MD5 |
307d86d182524eb007452e29866e1337
|
|
| BLAKE2b-256 |
ac7f2cbbeffc2715a9a272d3d61c2368bb9a4cdfcf4a479d83291eb775f85af1
|










