A lightweight generative model that extends SMILES fragments into syntactically valid molecules
Project description
Chempleter

Molecular autocomplete




Chempleter is a lightweight generative sequence model based on a multi-layer gated recurrent units (GRU) to predict syntactically valid extensions of a provided molecular fragment or bridge two molecules/molecular fragments. It operates on SELFIES token sequences, ensuring syntactically valid molecular generation and accepts SMILES notation as input. Due to its simple recurrent architecture and small vocabulary, the model runs efficiently on both CPUs and GPUs.
-
What can Chempleter do?
-
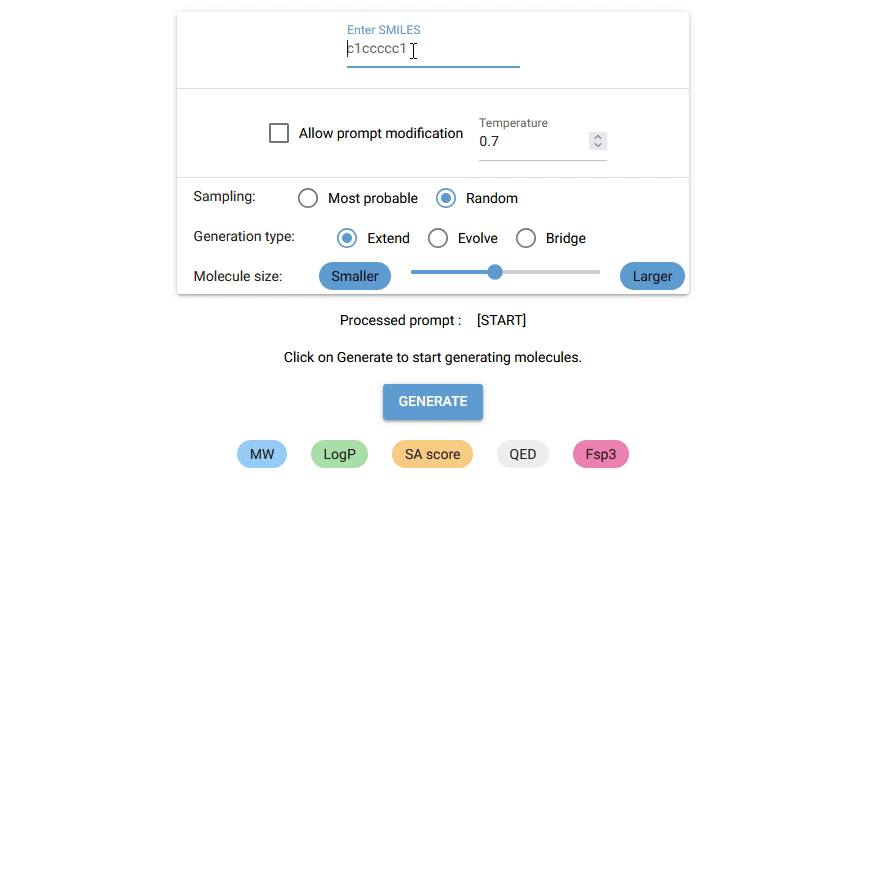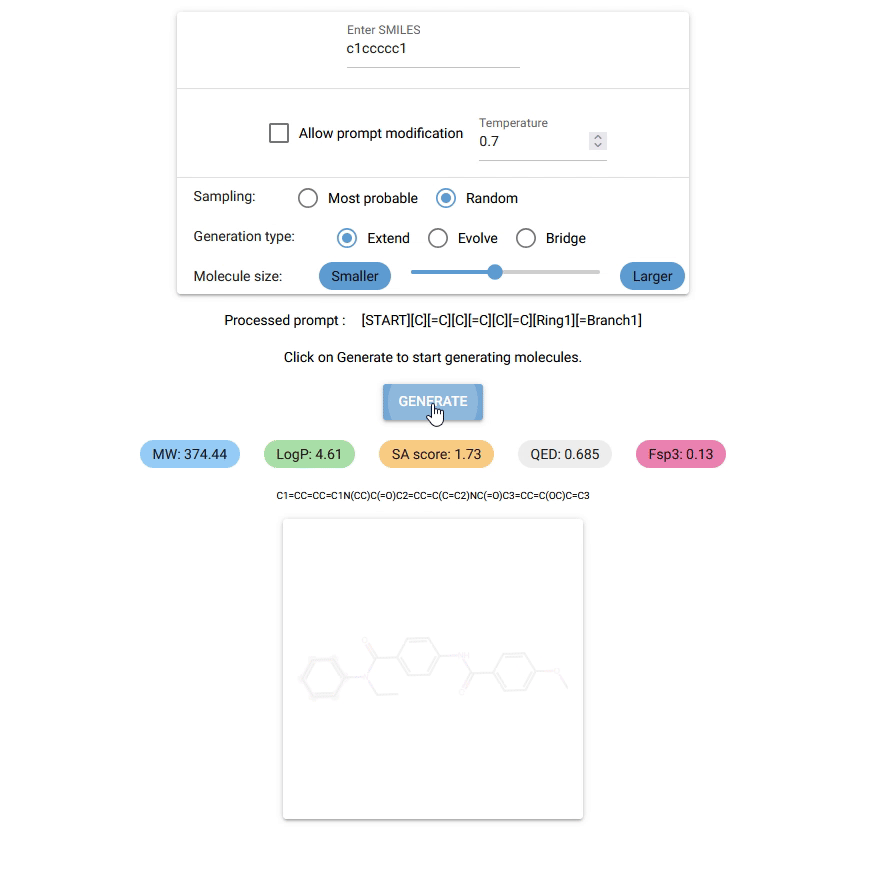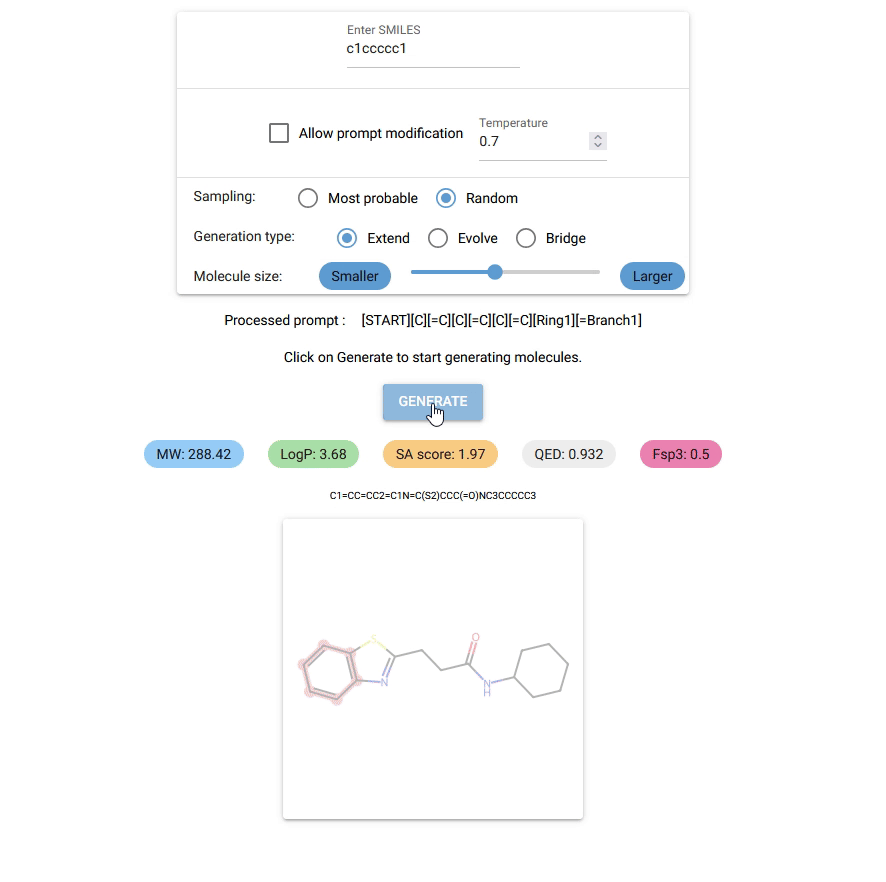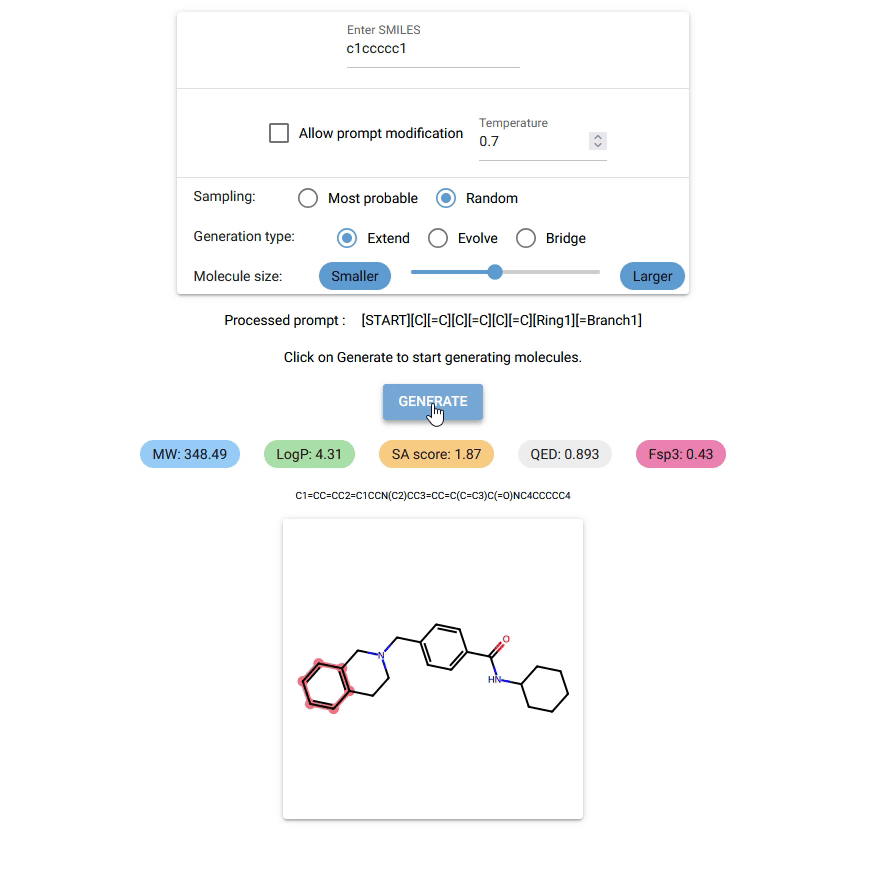
Currently, Chempleter accepts an intial molecule/molecular fragment in SMILES format and generates a larger molecule with that intial structure included, while respecting chemical syntax. It also shows some interesting descriptors.
-
It can be used to generate a wide range of structural analogs which the share same core structure (by changing the sampling temperature) or decorate a core scaffold iteratively (by increasing generated token lengths)
-
It can be used to bridge two molecules/molecular fragments to explore linker chemistry.
-
In the future, it might be adapated to predict structures with a specific chemical property using a regressor to rank predictions and transition towards more "goal-directed" predictions.
-
Prerequisites
- Python ">=3.12"
- uv (optional but recommended)
Installation
See detailed installation instructions.
Getting started
Visit Chempleter's docs.
Quick start
-
Run the GUI directly without installing (via uv):
-
On windows:
uvx --from chempleter chempleter-gui.exe -
On linux/MacOS:
uvx --from chempleter chempleter-gui -
To know more about using the GUI and various options, see here.
Or
-
-
Install using uv
uv pip install chempleter -
Use the GUI
-
To start the Chempleter GUI after installing, execute in a terminal:
uv run chempleter-gui -
Type in the SMILES notation for the starting structure or leave it empty to generate random molecules. Click on
GENERATEbutton to generate a molecule. -
To know more about using the GUI and various options, see here.
-
-
Run GUI after installation
uv run chempleter-gui -
Use as a python library
-
To use Chempleter as a python library:
from chempleter.inference import extend generated_mol, generated_smiles, generated_selfies = extend(smiles="c1ccccc1") print(generated_smiles) >> C1=CC=CC=C1C2=CC=C(CN3C=NC4=CC=CC=C4C3=O)O2
To draw the generated molecule :
from rdkit import Chem Chem.Draw.MolToImage(generated_mol)
-
For details on available paramenters and inference functions, see generating molecules.
-
Model history and validation
Project structure
- src/chempleter: Contains python modules relating to different functions.
- src/chempleter/processor.py: Contains fucntions for processing csv files containing SMILES data and generating training-related files.
- src/chempleter/dataset.py: ChempleterDataset class
- src/chempleter/model.py: ChempleterModel class
- src/chempleter/inference.py: Contains functions for inference
- src/chempleter/train.py: Contains functions for training
- src/chempleter/gui.py: Chempleter GUI built using NiceGUI
- src/chempleter/data : Contains trained model, vocabulary files
License
MIT License
Copyright (c) 2025-2026 Davis Thomas Daniel
Contributing
Any contribution, improvements, feature ideas or bug fixes are always welcome.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file chempleter-0.1.0b6.tar.gz.
File metadata
- Download URL: chempleter-0.1.0b6.tar.gz
- Upload date:
- Size: 21.8 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.8.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4794f7be8af7454b906ce3d4e2b1de20deb2cb2246405d05646d80953b6b2b3b
|
|
| MD5 |
e598b9432b2e6ee4a7165d2554739079
|
|
| BLAKE2b-256 |
b0d91552b1bdc98e1c93865736d6704284f869227bc40d53df326fffc56438fe
|
File details
Details for the file chempleter-0.1.0b6-py3-none-any.whl.
File metadata
- Download URL: chempleter-0.1.0b6-py3-none-any.whl
- Upload date:
- Size: 21.9 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.8.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ea90e385aa3c9204e4c6ef3aa01b89bd9abcba091702673d61a7dc37f1602c7a
|
|
| MD5 |
1e351ad035d0b0d7d6b54cbd872e6acc
|
|
| BLAKE2b-256 |
72872131368defb1a69ed258bee719a241783f052d4e7cd1b3e3c36681040f4f
|












