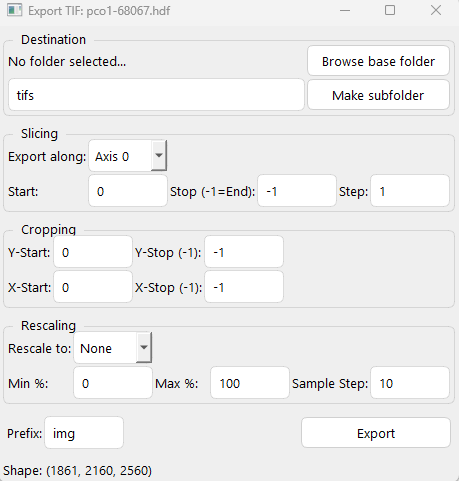
GUI software for viewing images, text, cine, and HDF files.
Project description
DatView
(Dat)a (View)er software
Python GUI software for folder browsing and viewing text, image, Cine, and HDF files
Motivation
For synchrotron-based tomography, users need convenient tools to view their data, typically in TIF, HDF, or CINE format during experiments, along with basic assessment tools such as contrast adjustment, zooming, line-profile viewing, histograms, image statistics, or percentile density. However, at synchrotron facilities, where Linux OS and open-source software are the primary tools, users often need to switch between multiple GUI applications for these tasks such as Nautilus for folder browsing, NeXpy or HDFView for HDF files, Gedit for text files, and ImageJ for image viewing.
This separation of tools is inconvenient, especially since many users are not familiar with the Linux OS. DatView provides a unified GUI for all these tasks, improving efficiency and user experience. DatView runs across operating systems.
Design Philosophy
DatView has been developed following two key guidelines:
- Minimize dependencies and the codebase.
- Maximize functionality and maintainability.
For distributing the software through Pip and Conda, the software is structured based on the RUI (Rendering-Utilities-Interactions) concept, which is a user-friendly adaptation of the MVC design pattern.
For the easiest usage, a monolithic codebase (datview.py) is provided, allowing users to simply copy the file and run it without needing to install the software through Pip or Conda, provided that their Python environment includes H5py, Pillow, and Matplotlib.
Features
-
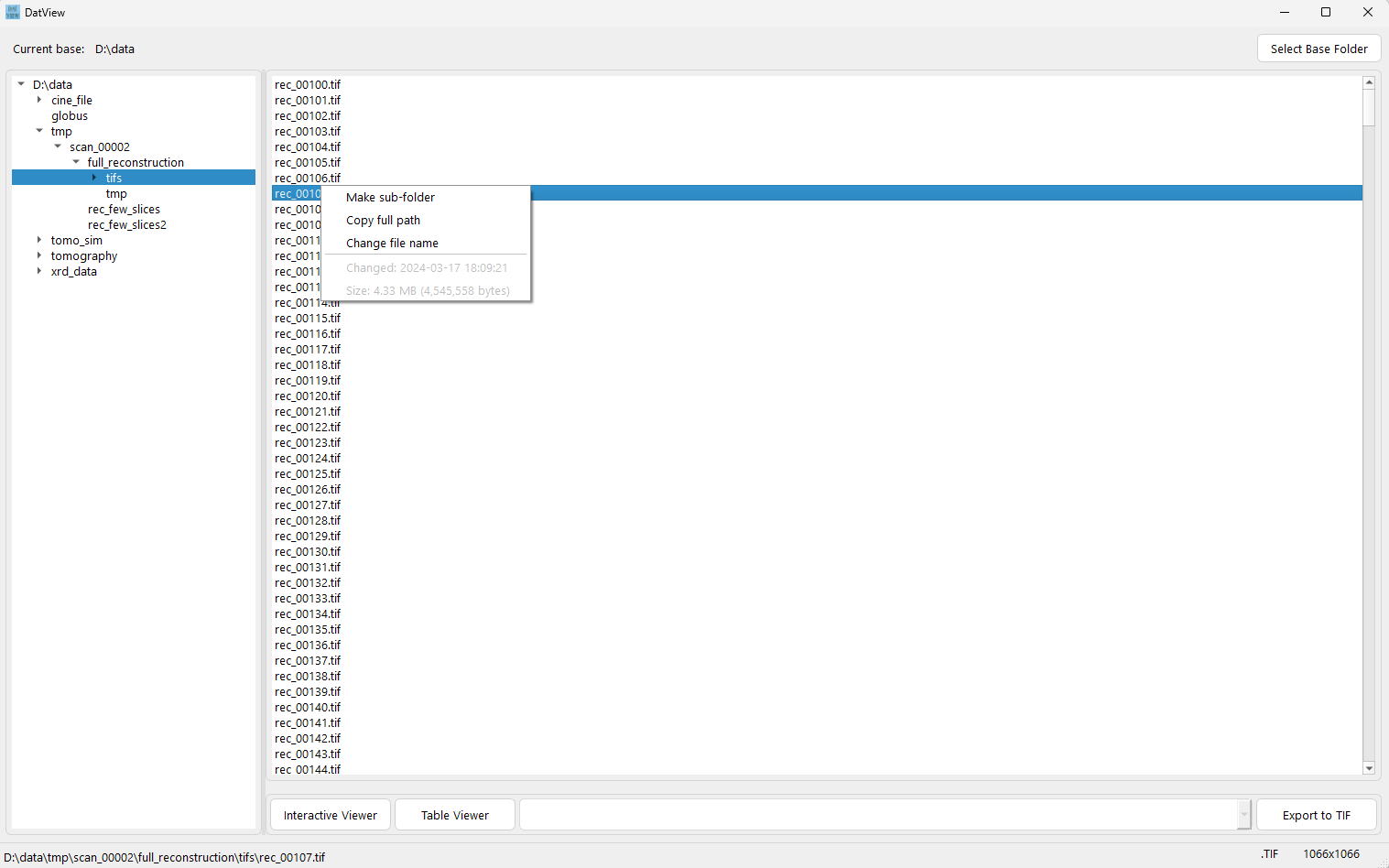
Fast folder browsing and file listing.
-
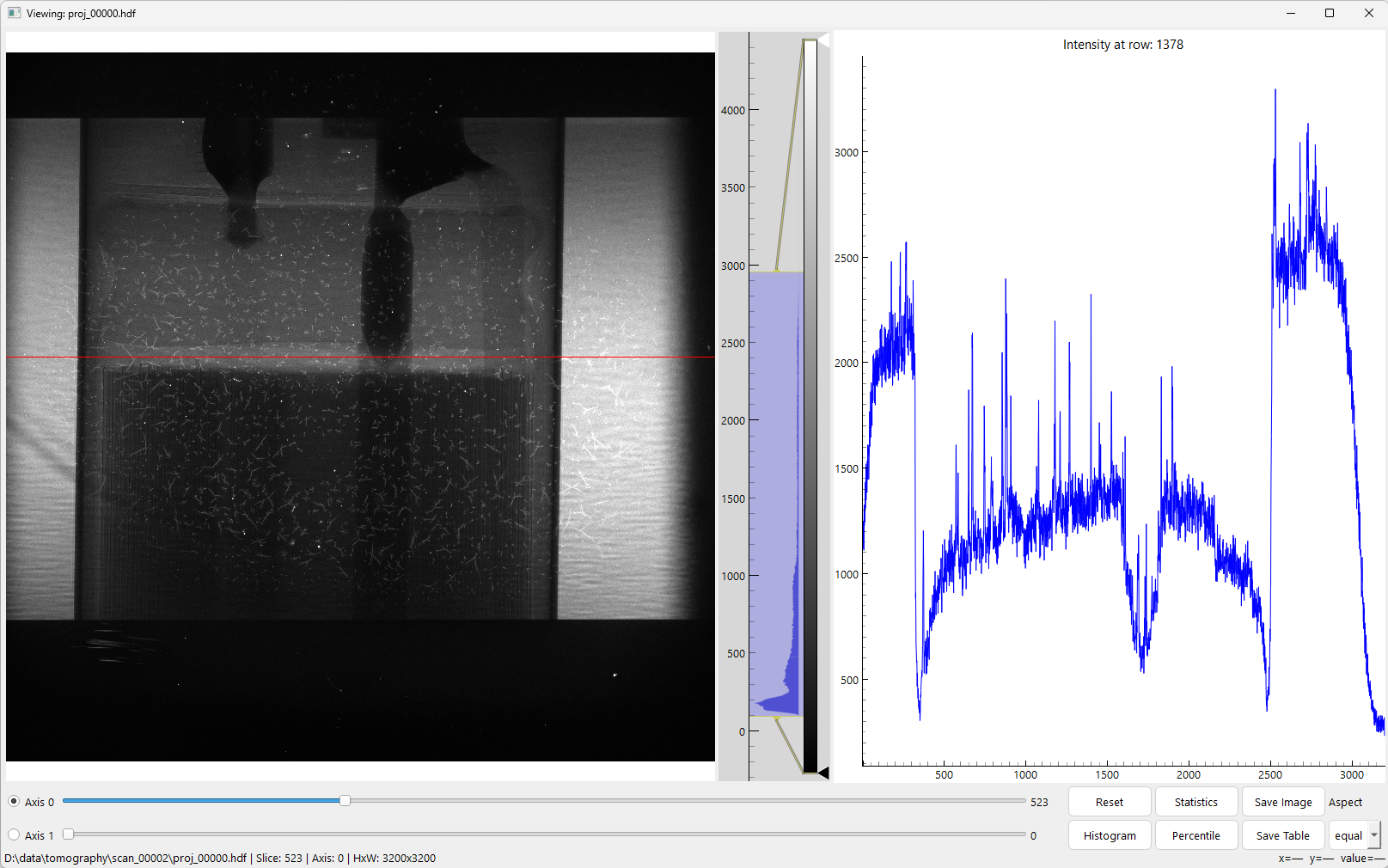
Interactive viewing 1D, 2D, or 3D datasets in an HDF file. Supports ROI zooming, horizontal/vertical line-profile selection, contrast adjustment, and slicing along axis 0 and 1.
-
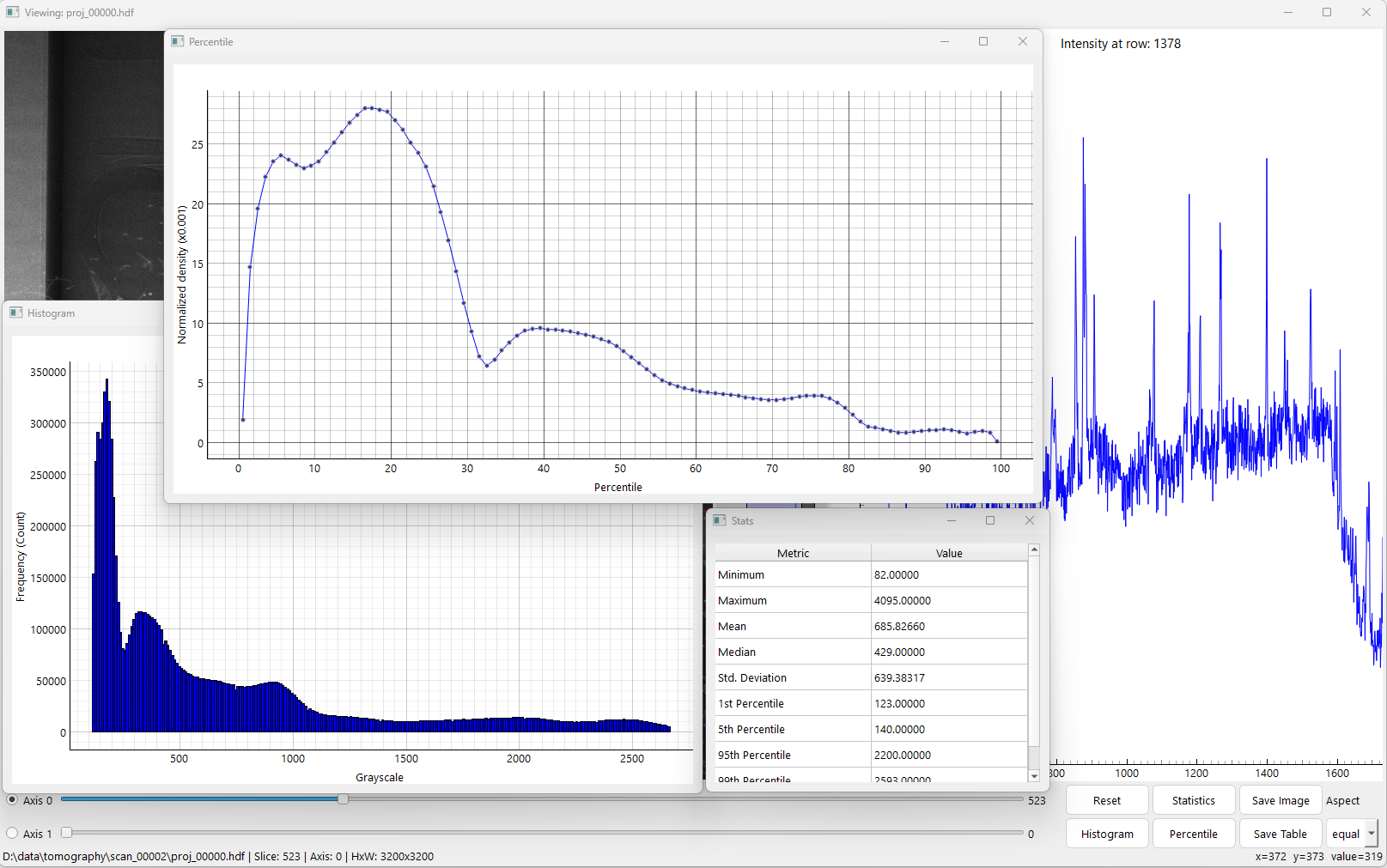
Options to display histogram, percentile density, image statistics.
-
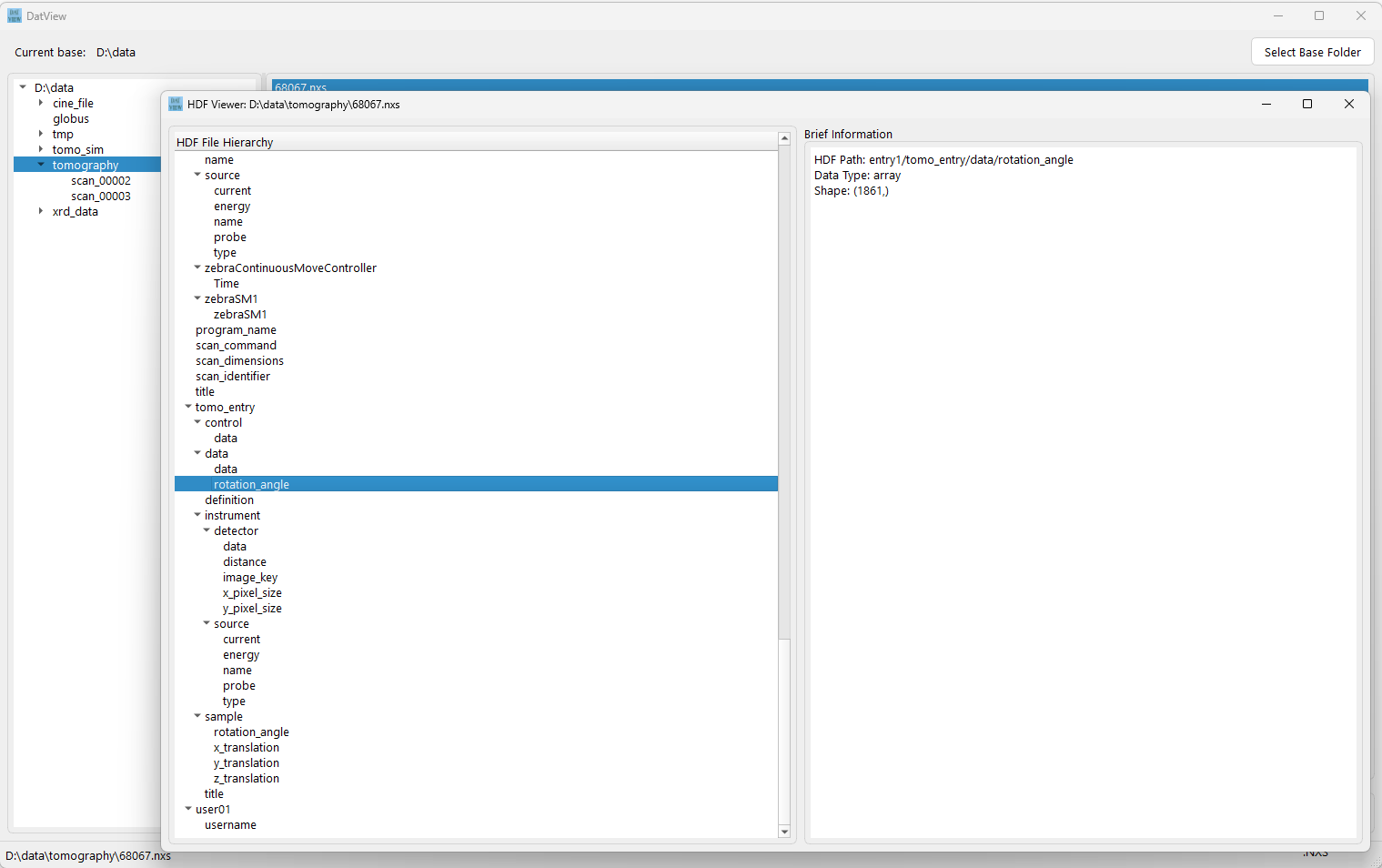
View metadata in HDF or CINE files, and display text-file contents.
-
Export to TIF files from HDF or CINE files.
-
Interactive viewing of TIF files in a folder or frames of a CINE file.
-
Interactive viewing of common image formats (JPG, PNG, TIF, ...).
-
Viewing 1D or 2D datasets of an HDF file in table format.
-
Opening multiple interactive viewers simultaneously.
-
Saving a 2D array in a 3D dataset (HDF or CINE) as an image.
-
Saving a 1D or 2D dataset of an HDF file or the current line profile as a CSV file.
Installation
Install Miniconda, Anaconda or Miniforge, then open a Linux terminal or the Miniconda/Anaconda PowerShell prompt and use the following commands for installation.
Using pip:
pip install datview
Using conda:
conda install -c conda-forge datview
Once installed, launching Datview with
datview
Using -h for option usage
datview -h
Installing from source:
- If using a single file:
- Copy the file datview.py. Install python, h5py, pillow, and matplotlib
- Run:
python datview.py
- If using setup.py
- Create conda environment
conda create -n datview python=3.11 conda activate datview
- Clone the source (git needs to be installed)
git clone https://github.com/algotom/datview.git
- Navigate to the cloned directory (having setup.py file)
pip install .
- Create conda environment
Generating the executable application
-
Install the required Python packages in your environment: PyInstaller, H5py, Hdf5plugin, Pillow, Matplotlib
-
Use the build_exe_app.py script and run the following command:
python build_exe_app.py
Usage
- Double-click an HDF or CINE file to display its metadata.
- Click "Interactive Viewer" to view images from a 3D dataset in an HDF file or from a folder of TIFF files.
- Click "Save Image" to save the image you’re currently viewing in the Interactive-Viewer window of an HDF file.
- Click "Save Table" to save a 1D or 2D array from an HDF dataset or a current line-profile in interactive viewer, to a CSV file.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file datview-1.3.0.tar.gz.
File metadata
- Download URL: datview-1.3.0.tar.gz
- Upload date:
- Size: 68.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
87e798525a52cd56218a88e23e9354587f55d7171d20833132c016155a655be6
|
|
| MD5 |
743ecb67465e749a968306378103d1fd
|
|
| BLAKE2b-256 |
78764ec25b323a90cfa0cffeb27f153f35dedecfbe6f7b37d0f978f3bfeedc9c
|
File details
Details for the file datview-1.3.0-py3-none-any.whl.
File metadata
- Download URL: datview-1.3.0-py3-none-any.whl
- Upload date:
- Size: 67.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bc85327389b56545b68bbfeb201a9c004e4511cf24b224a563da7bbdd14b75bb
|
|
| MD5 |
5d781a20edc494551d019a1235eb1522
|
|
| BLAKE2b-256 |
d16a8dabad57787a8e6eab99c42cd0ef0b2d1b3752e99a8769f8ad525f282600
|