A simple plugin to use models developped for flowering, fruitlet and fruit
Project description
DeepPhenoTree
Herearii Metuarea, Abdoul-Djalil Ousseini Hamza, Walter Guerra†, Andrea Patocchi, Lidia Lozano, Shauny Van Hoye, Francois Laurens, Jeremy Labrosse, Pejman Rasti, David Rousseau†.
† project lead
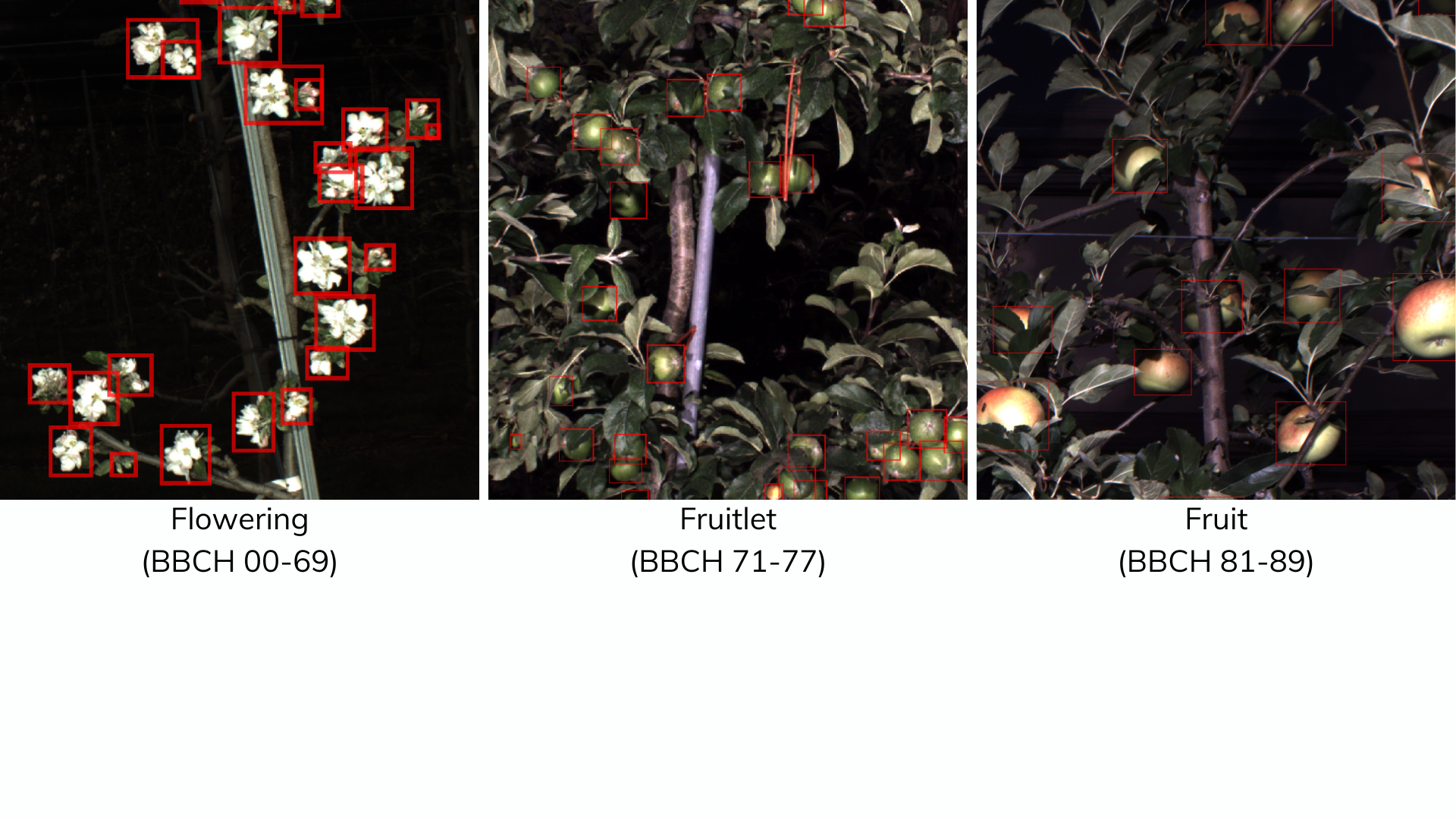


DeepPhenoTree is though as a tool to enable automatic detection of phenological stages associated with flowering, fruitlet, and fruit in harvest time from images using deep learning–based object detection models.
This napari plugin was generated with copier using the napari-plugin-template (None).
Contribution
Article (under review)
DeepPhenoTree – Apple Edition: a Multi-site apple phenology RGB annotated dataset with deep learning baseline models. Herearii Metuarea, Abdoul djalil Ousseni hamza, Walter Guerra, Andrea Patocchi, Lidia Lozano, Shauny Van Hoye, Francois Laurens, Jeremy Labrosse, Pejman Rasti, David Rousseau.
Dataset
Herearii Metuarea; Abdoul djalil Ousseni hamza; Lou Decastro; Jade Marhadour; Oumaima Karia; Lorène Masson; Marie Kourkoumelis-Rodostamos; Walter Guerra; Francesca Zuffa; Francesco Panzeri; Andrea Patocchi; Lidia Lozano; Shauny Van Hoye; Marijn Rymenants; François Laurens; Jeremy Labrosse; Pejman Rasti; David Rousseau, 2026, "DeepPhenoTree - Apple Edition", https://doi.org/10.57745/NORPF1, Recherche Data Gouv, V5, UNF:6:FyJNuJx4BVZxWuG8hI4gEw== [fileUNF]
Installation
You can install deepphenotree via pip:
pip install deepphenotree
If napari is not already installed, you can install deepphenotree with napari and Qt via:
pip install "deepphenotree[all]"
To install latest development version :
pip install git+https://github.com/hereariim/deepphenotree.git
GPU is mandatory for time processing and models running (especially RT-DETR). Please visit the official PyTorch website to get the appropriate installation command: 👉 https://pytorch.org/get-started/locally
Exemple : GPU (CUDA 12.1)
pip install torch torchvision torchaudio --index-url https://download.pytorch.org/whl/cu121
Getting started
Running from Python
1. Load sample image
from deepphenotree._sample_data import DeepPhenoTreeData
# Flowering data
data_flower = DeepPhenoTreeData('Flowering')
images = data_flower.data # Dimension : (5120, 5120, 3, 4)
country = data_flower.names # ['Belgium', 'Italy', 'Spain', 'Switzerland']
# Fruitlet data
data_fruitlet = DeepPhenoTreeData('Fruitlet')
# Fruit data
data_fruit = DeepPhenoTreeData('Fruit')
2. Run inference
from deepphenotree.inference import YoloInferencer
image = # Your RGB image
# Flowering task
infer = YoloInferencer("Flowering")
bbx = infer.predict_boxes(image)
# Fruitlet task
infer = YoloInferencer("Fruitlet")
bbx = infer.predict_boxes(image)
# Fruit task
infer = YoloInferencer("Fruit")
bbx = infer.predict_boxes(image)
Running from Napari
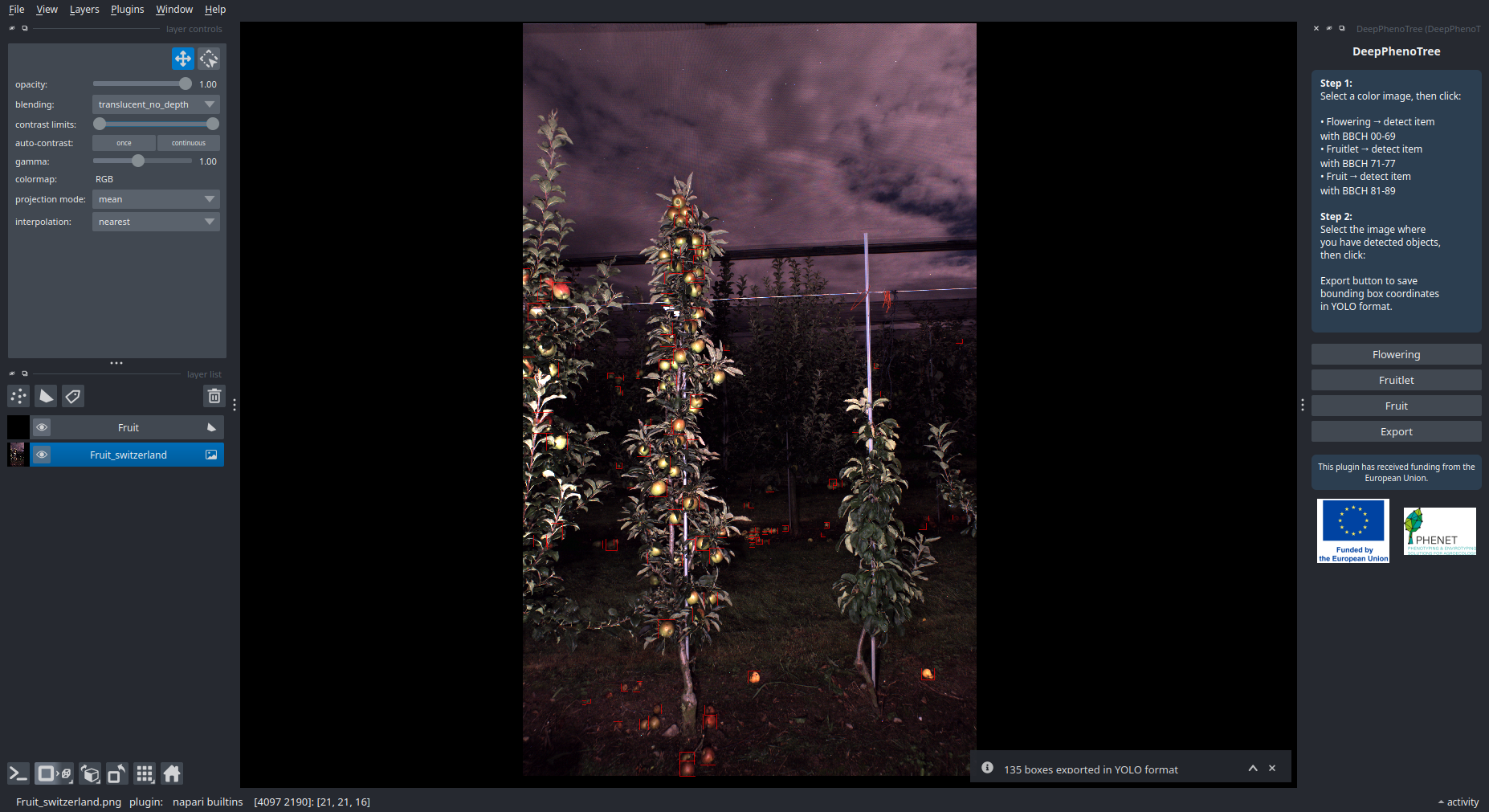
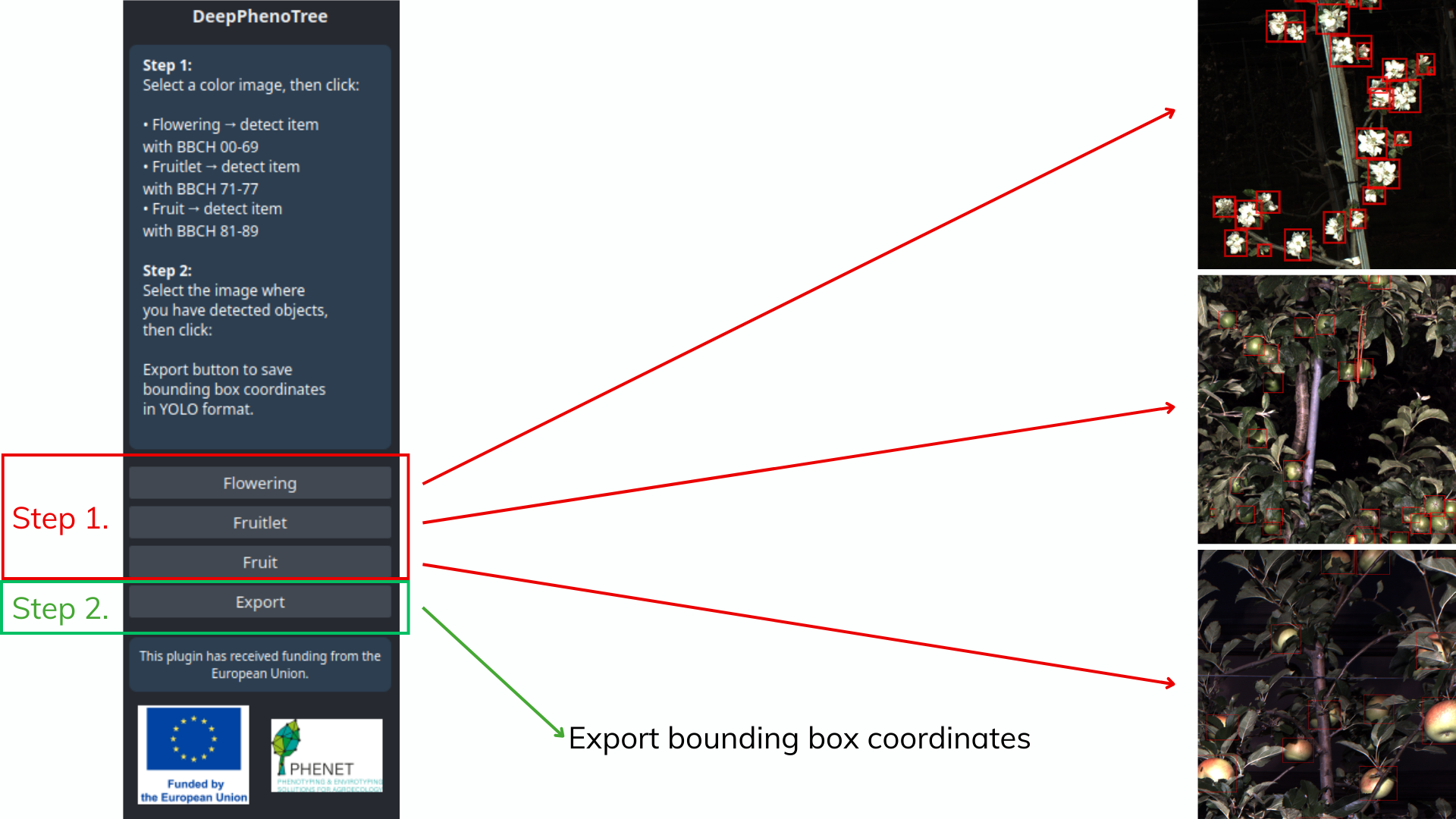
This plugin is a tool to perform targeted image inference on user-provided images. Users can run three specific detection tasks via dedicated buttons: flowering, fruitlet, and fruit detection. The plugin returns the coordinates of bounding boxes around detected objects, and a message informs the user of the number of detected boxes. Several developments are ongoing—feel free to contact us if you have requests or suggestions.
Scheme
Input
User drag and drop RGB image on napari window. Otherwise, user can select an image among suggested images from the plugin :
File > Open Sample > DeepPhenoTree > images
Note : The images available in Open Sample > DeepPhenoTree correspond to the test data associated with the models provided in this plugin.
Process
User click to make inference in image :
- Flowering : Detect all objects (from BBCH 00 to BBCH 69) from bud developpement to flowering.
- Fruitlet : Detect fruit in developement (from BBCH 71 to 77)
- Fruit : Detect all fruit in harvest time (from BBCH 81 to 89)
Output
Bounding box displayed in layer Flowering for flowering, Fruitlet for fruitlet and Fruit for fruit.
Model
DeepPhenoTree consists of a RT-DETR trained on DeepPhenoTree dataset.
The trained models used in this project are not publicly available. They are part of ongoing research and collaborative projects, and therefore cannot be distributed at this time.
However, the codebase is provided to ensure reproducibility and transparency of the proposed methodology.
Images results
Standard deviation is computed over 5-fold cross-validation. Overall (4 sites) denotes the aggregated evaluation across the four experimental sites (Switzerland, Belgium, Spain, and Italy).
| Dataset | Location | Precision | Recall | mAP@.5 | mAP@.5:.95 |
|---|---|---|---|---|---|
| Overall (4 sites) | 0.69 ± 0.01 | 0.58 ± 0.02 | 0.65 ± 0.02 | 0.37 ± 0.02 | |
| Switzerland | 0.73 ± 0.02 | 0.60 ± 0.04 | 0.68 ± 0.03 | 0.40 ± 0.04 | |
| Flowering | Belgium | 0.72 ± 0.02 | 0.63 ± 0.03 | 0.69 ± 0.03 | 0.40 ± 0.03 |
| Spain | 0.66 ± 0.01 | 0.53 ± 0.05 | 0.60 ± 0.03 | 0.30 ± 0.02 | |
| Italy | 0.69 ± 0.04 | 0.61 ± 0.03 | 0.67 ± 0.04 | 0.40 ± 0.04 | |
| ------------ | ---------------------- | ---------------- | ---------------- | -------------- | -------------- |
| Overall (4 sites) | 0.85 ± 0.02 | 0.73 ± 0.02 | 0.82 ± 0.02 | 0.53 ± 0.01 | |
| Switzerland | 0.86 ± 0.04 | 0.78 ± 0.04 | 0.84 ± 0.06 | 0.56 ± 0.04 | |
| Fruitlet | Belgium | 0.83 ± 0.03 | 0.65 ± 0.04 | 0.77 ± 0.04 | 0.52 ± 0.14 |
| Spain | 0.86 ± 0.02 | 0.72 ± 0.03 | 0.81 ± 0.03 | 0.52 ± 0.03 | |
| Italy | 0.88 ± 0.01 | 0.80 ± 0.01 | 0.88 ± 0.01 | 0.61 ± 0.01 | |
| ------------ | ---------------------- | ---------------- | ---------------- | -------------- | -------------- |
| Overall (4 sites) | 0.87 ± 0.01 | 0.79 ± 0.01 | 0.86 ± 0.01 | 0.57 ± 0.01 | |
| Switzerland | 0.86 ± 0.03 | 0.80 ± 0.02 | 0.87 ± 0.02 | 0.59 ± 0.01 | |
| Fruit | Belgium | 0.90 ± 0.01 | 0.84 ± 0.01 | 0.90 ± 0.01 | 0.63 ± 0.02 |
| Spain | 0.86 ± 0.02 | 0.75 ± 0.02 | 0.84 ± 0.02 | 0.51 ± 0.03 | |
| Italy | 0.88 ± 0.02 | 0.84 ± 0.03 | 0.90 ± 0.02 | 0.66 ± 0.02 |
DeepPhenoTree Dataset
DeepPhenoTree – Apple Edition, a multi-site, multi-variety, RGB image dataset dedicated to the classification of key apple treephenological stages.

Acknowlegments
This work is led by IRHS, Université d'Angers, INRAe, Institut Agro. It is supported by :
- The European Union’s Horizon Europe research innovation programme under grant agreement No 101094587 (PHENET project)
- The French National Research Agency (ANR) as part of the Programme d'Investissements d'Avenir under grant agreement ANR-11-INBS-0012 (PHENOME-EMPHASIS)
- The French National Research Agency (ANR) as part of the France 2030 investment plan under grant agreement ANR-22-PEAE-0012 (AGROECOPHEN)
- The HPC resources of IDRIS under the allocation 2024-AD010115553 made by GENCI.
Contact
Imhorphen team, bioimaging research group 42 rue George Morel, Angers, France
- Herearii Metuarea, herearii.metuarea@univ-angers.fr
- Abdoul-Djalil Ousseini Hamza, abdoul-djalil.ousseini-hamza@inrae.fr
- Pr David Rousseau, david.rousseau@univ-angers.fr
Contributing
Contributions are very welcome. Tests can be run with tox, please ensure the coverage at least stays the same before you submit a pull request.
License
Distributed under the terms of the GNU LGPL v3.0 license, "deepphenotree" is free and open source software
Issues
If you encounter any problems, please file an issue along with a detailed description.
Citing
If you use DeepPhenoTree in your research, please use the following BibTeX entry.
@article{metuarea2026deepphenotree,
title={DeepPhenoTree--Apple Edition: a Multi-site apple phenology RGB annotated dataset with deep learning baseline models},
author={Herearii Metuarea and Abdoul-Djalil Ousseini Hamza and Walter Guerra and Francesca Zuffa and Francesco Panzeri and Andrea Patocchi and Lidia Lozano and Shauny Van Hoye and François Laurens and Jeremy Labrosse and Pejman Rasti and David Rousseau},
year={2026}
}
If you use DeepPhenoTree dataset in your research, please use the following BibTeX entry.
@dataset{metuarea2026deepphenotree_dataset,
author = {Metuarea Herearii and Abdoul-Djalil Ousseini Hamza and Decastro Lou and Marhadour Jade and Karia Oumaima and Masson Lorène and Kourkoumelis-Rodostamos Marie and Guerra Walter and Zuffa Francesca and Panzeri Francesco and Patocchi Andrea and Lozano Lidia and Van Hoye Shauny and Rymenants Marijn and Laurens François and Labrosse Jeremy and Rasti Pejman and Rousseau David},
title = {DeepPhenoTree – Apple Edition},
year = {2026},
version = {5},
publisher = {Recherche Data Gouv},
doi = {10.57745/NORPF1}
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file deepphenotree-1.0.8.tar.gz.
File metadata
- Download URL: deepphenotree-1.0.8.tar.gz
- Upload date:
- Size: 13.8 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b2d2ccd223dce6a688b51b2baf20b81fd602c008f293d34bb6459557c6f225ab
|
|
| MD5 |
56042c2f1c8de98941b58caf2d10c15c
|
|
| BLAKE2b-256 |
83886126b5cf29fdc1c8aa183bf0623aec281a0d22d0a44ec448e219997c6949
|
File details
Details for the file deepphenotree-1.0.8-py3-none-any.whl.
File metadata
- Download URL: deepphenotree-1.0.8-py3-none-any.whl
- Upload date:
- Size: 13.8 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a6834d0b449a5d49b9c4011a028ca4d0da50423978e13fc1c62854123cbec4b2
|
|
| MD5 |
69ba5974d52a9d275f460fabf4f49d35
|
|
| BLAKE2b-256 |
bc69df137ce3ed23d7fc507a19e5b0b9fb2f0de5ee09bc95410084d01394ff88
|



















