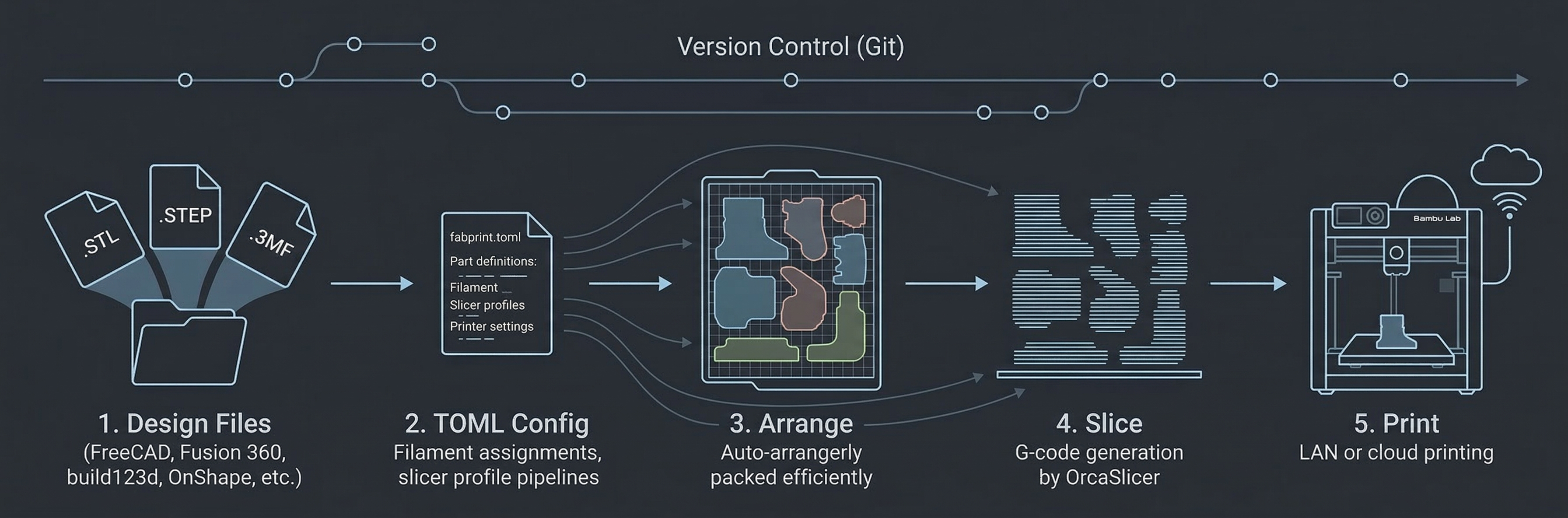
The build system for reproducible 3D prints. Define parts, slicer settings, and printer targets in code.
Project description

estampo
The build system for reproducible 3D prints.
3D printing workflows are trapped in GUIs. Slicer settings get lost between sessions, printer configs drift across machines, and there's no way to version, diff, or review a print job. None of this is accessible to automation — or to AI assistants.
estampo takes a different approach: define parts, slicer settings, and pipeline stages in a single TOML file — text that's git-friendly, diffable, and committable alongside your CAD files. Because the entire workflow is text, it's naturally accessible to AI coding assistants, CI systems, and code review. Same repo, same config → same G-code, locally or in CI.
Warning: estampo is in active early development. We are moving fast and breaking things — config format, CLI flags, and Python APIs may change between minor versions without deprecation. Pin your version if stability matters to you.
Safety: estampo generates G-code from your configuration but does not verify that settings are safe for your specific printer. Incorrect temperatures, speeds, or missing supports can damage your printer or create a fire hazard. Always review sliced output before sending to a printer.
estampo validatechecks config structure and setting names — it does not check print safety. Use at your own risk.
Note: This project was previously called fabprint. If you have an existing install, run
pip install estampoto upgrade — config files and credentials are migrated automatically.
Works with STL, STEP, and 3MF files, and pairs naturally with code-CAD tools like build123d, OpenSCAD, and cadquery.
# estampo.toml — a multi-part print with slicer overrides
[[parts]]
file = "enclosure_base.step"
orient = "flat"
filament = 1 # AMS slot 1: PETG-CF
[[parts]]
file = "enclosure_lid.step"
orient = "upright"
filament = 1
[[parts]]
file = "button_cap.stl"
copies = 4
filament = 2 # AMS slot 2: PLA
[slicer]
engine = "orca"
version = "2.3.1" # pinned for reproducibility
[slicer.orca]
printer = "Bambu Lab P1S 0.4 nozzle"
process = "0.20mm Standard @BBL X1C"
filaments = ["Generic PETG-CF @base", "Generic PLA @base"]
[slicer.orca.overrides]
sparse_infill_density = "25%"
enable_support = 1
brim_type = "auto_brim"
[pipeline]
stages = ["load", "arrange", "plate", "slice", "gcode-info", "pack"]
[pack]
command = "bambox repack {sliced_3mf}"
output = "{sliced_3mf}"
estampo run # arrange → slice → pack, one command
Output → estampo_output/enclosure
✔ Loaded 3 parts
✔ Arranged 3 parts onto plate (256×256mm)
✔ Plate exported → plate.3mf
✔ Preview exported → plate_preview.3mf
✔ Sliced with OrcaSlicer 2.3.1 in 48s
✔ Print time: 3h 42m, 24.6g filament
✔ Packed → plate_sliced.gcode.3mf
What estampo does
- Define parts + settings in
estampo.toml - Arrange — bin-packs models onto the build plate
- Slice — using a pinned slicer version (via Docker) for identical G-code across machines
- Post-process — run command stages (pack for Bambu, template resolution, etc.)
Everything is declared in a single TOML file. Lock the slicer version, pin the profiles, and the output is reproducible on any machine or in CI.
Why not just use OrcaSlicer CLI?
OrcaSlicer CLI is great for slicing a prepared plate. estampo builds a reproducible pipeline around it:
- Arrangement — bin-packs multiple STLs onto the build plate (OrcaSlicer CLI has no arrange step)
- Multi-part filament mapping — per-part filament slot assignment and paint color preservation, injected into the 3MF metadata
- Reproducible builds — pin slicer profiles into your repo + lock OrcaSlicer version in Docker = identical gcode on any machine
- Partial execution —
--until plateto inspect layout,--only sliceto re-slice,--dry-runto test everything - Command stages — run external tools (e.g.
bambox pack) as pipeline stages with variable substitution - Headless Docker slicing — no GUI, no display server, works in CI, uses a specific OrcaSlicer version
Why not just use CuraEngine CLI?
CuraEngine CLI (CuraEngine slice -j definition.json -s KEY=VALUE -l model.stl -o output.gcode) is powerful but requires assembling the full invocation yourself:
| What you need to do | CuraEngine CLI | estampo |
|---|---|---|
| Load printer definition | -j printer.def.json + -d search paths for inheritance |
printer = "Ultimaker 2" in TOML |
| Set material/quality | Chain of -j files or -s overrides for every setting |
[slicer.cura.overrides] in TOML |
| Arrange multiple parts | Not supported — manual positioning | Automatic bin-packing |
| Multi-extruder mapping | -g -e0 -l a.stl -g -e1 -l b.stl per mesh group |
filament = 1 per part |
| Reproducible builds | Track all definition JSONs yourself | version = "5.12.0" + Docker |
| Version control | Shell scripts + scattered JSON definitions | Single TOML file — git-diffable and reviewable |
| Run in CI | Install CuraEngine + definitions manually | uses: estampo/estampo/action@v1 |
| Pack for Bambu printers | Separate manual step | [pack] command stage |
With CuraEngine CLI, your build config ends up spread across shell scripts, -s flag lists, and JSON definition files — hard to diff, review, or commit as a coherent unit. estampo replaces all of that with a single declarative TOML file: git-friendly, diffable, and accessible to AI assistants and code review.
Works with AI assistants
GUI-based slicers require point-and-click interaction that AI assistants cannot drive. estampo's text-based config is different — an AI assistant can read your TOML, understand your print setup, suggest overrides, create CI workflows, and validate the result, all without leaving the conversation.
We provide a ready-to-paste AI setup prompt that gives any AI assistant (Claude, ChatGPT, Copilot) enough context to add estampo to a project from scratch — it scans the repo for model files, asks you a few questions, and generates the config. This isn't an AI feature bolted on — it's an architectural property of using text instead of a GUI.
Best fit
estampo is best suited to:
- Hardware teams keeping CAD and manufacturing inputs in Git
- Engineers who want deterministic slicing in CI
- Teams using AI coding assistants to manage their build and print workflows
- Makers who want a declarative print workflow instead of slicer click-ops
If you mostly want interactive print setup in a GUI, use OrcaSlicer or Cura directly.
Status
Stable
- Declarative print config in
estampo.toml - Multi-part arrangement
- Docker-based slicing with pinned slicer versions (OrcaSlicer, CuraEngine)
- Slicing for any printer supported by OrcaSlicer or CuraEngine
- Profile pinning into your repository
- CI slicing and artifact generation
- Network print initiation via Bambu Cloud
Experimental
- Bambu LAN printing
- Moonraker printing
Quick start
Prerequisites: Python 3.11+ and Docker. Docker is central to estampo — it runs the slicer (OrcaSlicer or CuraEngine) in a container with a pinned version so every machine produces identical G-code. A locally installed slicer can be used as a fallback but is not recommended.
pip install estampo
# or, to install as an isolated CLI tool:
pipx install estampo
Generate a config with the interactive wizard, or dump a commented template:
estampo init # interactive wizard — discovers profiles and CAD files, creates TOML
estampo init --template # dump a commented template
Or create estampo.toml by hand (see full config reference):
[pipeline]
stages = ["load", "arrange", "plate", "slice", "pack"]
[plate]
size = [256, 256] # build plate dimensions in mm
padding = 5.0
[slicer]
engine = "orca"
version = "2.3.1" # pin OrcaSlicer version for reproducibility
[slicer.orca]
printer = "Bambu Lab P1S 0.4 nozzle"
process = "0.20mm Standard @BBL X1C"
[slicer.orca.overrides] # simple way to define print settings without editing JSON
sparse_infill_density = "30%" # stronger infill
wall_loops = 3 # extra wall strength
enable_support = 1
brim_type = "auto_brim" # help adhesion
curr_bed_type = "Textured PEI Plate"
[[parts]] # define multiple parts using STEP, STL or 3MF inputs
file = "frame.step"
rotate = [180, 0, 0] # flip so mounting plate faces down
filament = "Generic PETG-CF @base"
[[parts]]
file = "wheel.stl"
copies = 5
orient = "upright"
filament = "Generic PETG-CF @base"
Run it (see full CLI reference):
estampo run # arrange, slice and send to printer
estampo run --until slice # stop after slicing
estampo run --dry-run # full pipeline without sending to printer
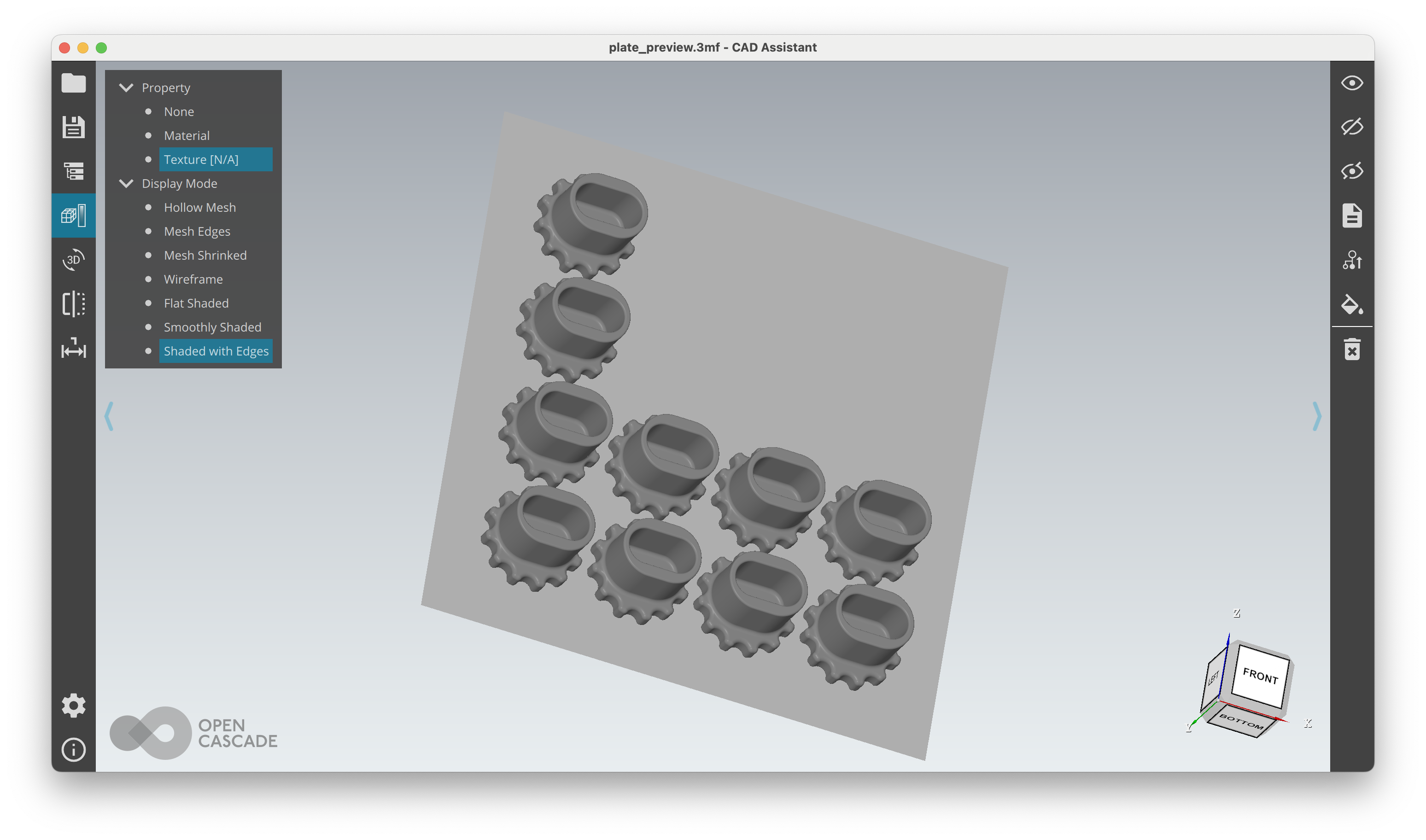
The arrangement (plate) stage generates a plate_preview.3mf — open it in any 3MF viewer to check placement:
Reproducibility
Pin profiles into your repo so builds are identical across machines:
estampo profiles pin # copies slicer profiles into ./profiles/
git add profiles/ # commit to lock them
Combined with a pinned version in [slicer] (which locks the Docker image), the same config always produces the same gcode.
CI/CD example
Automate slicing in GitHub Actions — push a commit, get G-code as a build artifact with print metrics on your PR:
# .github/workflows/slice.yml
name: Slice
on: [push, pull_request]
jobs:
slice:
runs-on: ubuntu-latest
steps:
- uses: actions/checkout@v6
- uses: estampo/estampo/action@v1
The action slices your model, uploads G-code as an artifact, and posts print time / filament stats as a PR comment. See action/README.md for all options.
CLI overview
estampo init # interactive config wizard
estampo init --template # dump commented TOML template
estampo init --workflow # wizard + GitHub Actions workflow
estampo validate # check config for issues
estampo run # full pipeline
estampo run --until plate # stop after plating
estampo run --only slice # run just one stage
estampo run --dry-run # everything except sending to printer
estampo profiles list # list available slicer profiles
estampo profiles pin # pin profiles for reproducible builds
Printing
estampo is printer-agnostic — it produces sliced output but does not send files to printers. For Bambu Lab printers, use bambox as a command stage to pack and print. Run bambox login to set up credentials (~/.config/bambox/credentials.toml). See the AI setup prompt for full examples.
Documentation
- CLI reference — all commands, flags, and pipeline stages
- Config reference — complete TOML format
- Developing — setup, testing, architecture
Development
estampo's codebase is authored by Claude, Anthropic's AI assistant, under the direction and review of a human maintainer. Architecture decisions, priorities, and acceptance criteria are set by the maintainer; Claude writes the code, tests, documentation, and CI workflows. Every commit is reviewed before merge.
License
Apache 2.0
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file estampo-0.4.0b1.tar.gz.
File metadata
- Download URL: estampo-0.4.0b1.tar.gz
- Upload date:
- Size: 8.9 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
09b173980a47b4db6501daac8484df00589a11b3a62254b48bb69654bd89668e
|
|
| MD5 |
4fefe4c089780d5045f253618f4df5b1
|
|
| BLAKE2b-256 |
d9a998f9ee965704d30715347c40d6102eeb44ee3205a48986262b1d158edef3
|
Provenance
The following attestation bundles were made for estampo-0.4.0b1.tar.gz:
Publisher:
release.yml on estampo/estampo
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
estampo-0.4.0b1.tar.gz -
Subject digest:
09b173980a47b4db6501daac8484df00589a11b3a62254b48bb69654bd89668e - Sigstore transparency entry: 1338319225
- Sigstore integration time:
-
Permalink:
estampo/estampo@e8716ab4035aa00cb30fd824c4f13fa67b0100d1 -
Branch / Tag:
refs/heads/main - Owner: https://github.com/estampo
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@e8716ab4035aa00cb30fd824c4f13fa67b0100d1 -
Trigger Event:
workflow_dispatch
-
Statement type:
File details
Details for the file estampo-0.4.0b1-py3-none-any.whl.
File metadata
- Download URL: estampo-0.4.0b1-py3-none-any.whl
- Upload date:
- Size: 160.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6741a83a1597483796025c8edc0a7f71b03f3793525aef3e7f2ce265235d28d9
|
|
| MD5 |
b08f3f2c47c8cacc31f5eaa97031cb16
|
|
| BLAKE2b-256 |
b3420e2f5de4bc1d9550cbc816e8657e89bf580b5c3ef5b0013880ef8513c3bc
|
Provenance
The following attestation bundles were made for estampo-0.4.0b1-py3-none-any.whl:
Publisher:
release.yml on estampo/estampo
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
estampo-0.4.0b1-py3-none-any.whl -
Subject digest:
6741a83a1597483796025c8edc0a7f71b03f3793525aef3e7f2ce265235d28d9 - Sigstore transparency entry: 1338319296
- Sigstore integration time:
-
Permalink:
estampo/estampo@e8716ab4035aa00cb30fd824c4f13fa67b0100d1 -
Branch / Tag:
refs/heads/main - Owner: https://github.com/estampo
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@e8716ab4035aa00cb30fd824c4f13fa67b0100d1 -
Trigger Event:
workflow_dispatch
-
Statement type: