A modular framework for evolutionary strategies and neuroevolution.
Project description
EvoLib – A Modular Framework for Evolutionary Computation

EvoLib is a lightweight and transparent framework for evolutionary computation, focusing on simplicity, modularity, and clarity — aimed at experimentation, teaching, and small-scale research rather than industrial-scale applications.
Key Features
- Transparent design: configuration via YAML, type-checked validation, and clear module boundaries.
- Modularity: mutation, selection, crossover, and parameter representations can be freely combined.
- Educational value: examples and a clean API make it practical for illustrating evolutionary concepts.
- Neuroevolution support: structural mutations (adding/removing neurons and connections) and evolvable networks via EvoNet.
- Type-checked: PEP8 compliant, and consistent code style.
EvoLib is currently in beta. The core API and configuration format are stable, but some features are still under development.
Directory Structure
evolib/
├── core/ # Individual, Population
├── config/ # Typed component configuration (Vector, EvoNet, etc.)
├── interfaces/ # Enums, types, helper protocols
├── initializers/ # Initializer registry and implementations
├── operators/ # Mutation, crossover, selection, etc.
├── registry/ # Strategy and operator registries
├── representation/ # ParaBase + Vector, EvoNet, Composite etc.
├── utils/ # Logging, plotting, math, config loader
└── examples/ # Educational examples and test runs
Installation
pip install evolib
Requirements: Python 3.10+ and packages in requirements.txt.
Example Usage
from evolib import Pop
def my_fitness(indiv):
# Custom fitness function (example: sum of vector)
indiv.fitness = sum(indiv.para["main"].vector)
pop = Pop(config_path="config/my_experiment.yaml",
fitness_function=my_fitness)
# Run the evolutionary process
pop.run()
For full examples, see 📁examples/ – including adaptive mutation, controller evolution, and network approximation.
Configuration Example (YAML)
parent_pool_size: 20
offspring_pool_size: 60
max_generations: 100
num_elites: 2
max_indiv_age: 0
stopping:
target_fitness: 0.01
patience: 20
min_delta: 0.0001
minimize: true
evolution:
strategy: mu_comma_lambda
modules:
controller:
type: vector
dim: 8
initializer: normal_vector
bounds: [-1.0, 1.0]
mutation:
strategy: adaptive_individual
probability: 1.0
strength: 0.1
brain:
type: evonet
dim: [4, 6, 2]
activation: [linear, tanh, tanh]
initializer: normal_evonet
mutation:
strategy: constant
probability: 1.0
strength: 0.05
# Optional fine-grained control
activations:
probability: 0.01
allowed: [tanh, relu, sigmoid]
structural:
add_neuron: 0.01
add_connection: 0.05
remove_connection: 0.02
recurrent: local # none | direct | local | all
keep_connected: true
ℹ️ Multiple parameter types (e.g. vector + evonet) can be combined in a single individual. Each component evolves independently, using its own configuration.
Use Cases
EvoLib is developed for clarity, modularity, and exploration in evolutionary computation.
It can be applied to:
- Illustrating concepts: simple, transparent examples for teaching and learning.
- Neuroevolution: evolve weights and network structures using EvoNet.
- Multi-module evolution: combine different parameter types (e.g. controller + brain).
- Strategy comparison: benchmark and visualize mutation, selection, and crossover operators.
- Function optimization: test behavior on benchmark functions (Sphere, Ackley, …).
- Showcases: structural XOR, image approximation, and other demo tasks.
- Rapid prototyping: experiment with new evolutionary ideas in a lightweight environment.
Learn EvoLib in 5 Steps
EvoLib includes a small set of examples that illustrate the core concepts step by step:
- Hello Evolution – minimal run with a custom fitness function and visible improvement over generations.
- Strategies in Action – (μ + λ) evolution step by step.
- Function Approximation – evolve support points to match a sine curve.
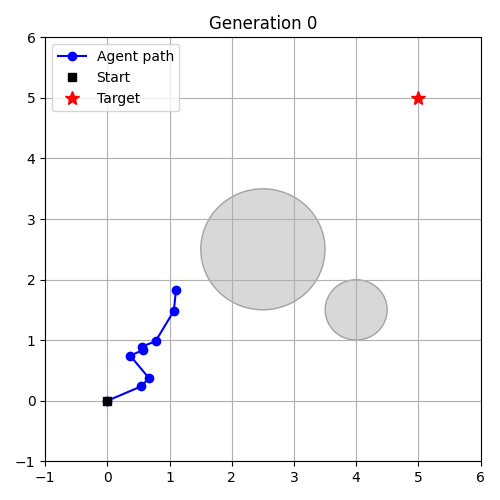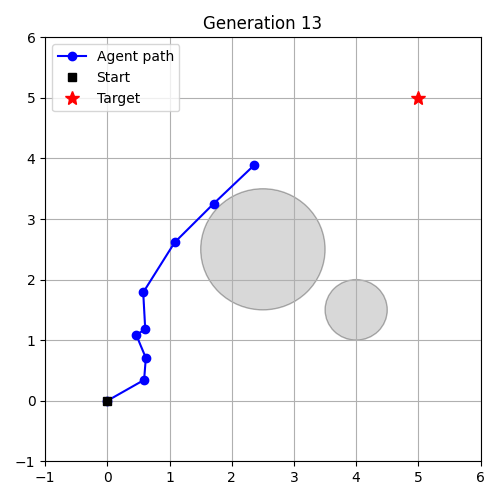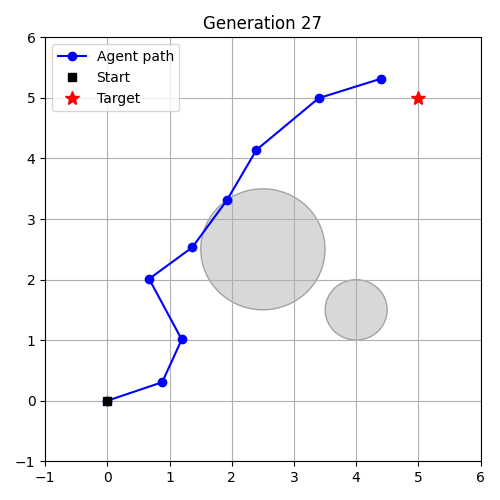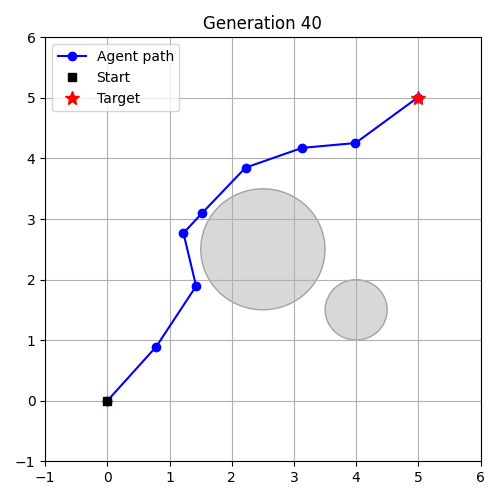
- Evolution as Control – evolve a controller in an environment.
- Neuroevolution with Structural Growth – evolve networks with growing topology.
For deeper exploration, see the full examples directory
Roadmap
- Adaptive Mutation (global, individual, per-parameter)
- Flexible Crossover Strategies (BLX, intermediate, none)
- Structured Neural Representations (EvoNet)
- Composite Parameters (multi-module individuals)
- Neuroevolution
- Topological Evolution (neurons, edges)
- Co-Evolution & Speciation Support
- Advanced Visualization
- Game Environment Integration (pygame, PettingZoo - early prototypes)
- Ray Support for Parallel Evaluation (early prototypes)
Documentation
Documentation for EvoLib is available at: 👉 https://evolib.readthedocs.io/en/latest/
License
MIT License – see MIT License.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file evolib-0.2.0b1.tar.gz.
File metadata
- Download URL: evolib-0.2.0b1.tar.gz
- Upload date:
- Size: 117.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
031077a610d30a50105123675f590621924dc169c9a97c94cd405ced4a5c327e
|
|
| MD5 |
57b14053ed30a5c8e2635a6352053db4
|
|
| BLAKE2b-256 |
50c8fabf3788a6e66e2ec3d591b8208df26989c82713435214a01f9818b0300c
|
File details
Details for the file evolib-0.2.0b1-py3-none-any.whl.
File metadata
- Download URL: evolib-0.2.0b1-py3-none-any.whl
- Upload date:
- Size: 155.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
99088a68468fce1f2d9c0ba3731f68feaf30d70f7605694553c00727124a4b00
|
|
| MD5 |
a37188ed821ada51541b6886b5078332
|
|
| BLAKE2b-256 |
f0cc2712fab2997a4852f4646522fef46dd029f96a1b66aa4d090afcc90b1462
|
















