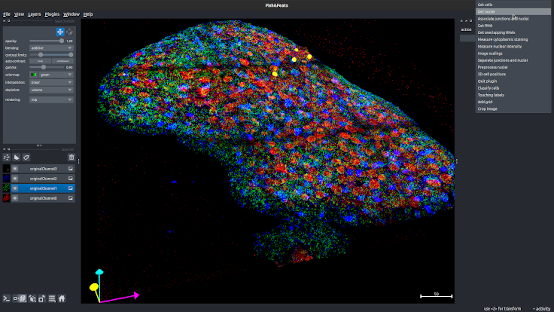
Napari plugin for RNA-Fish+cells analysis pipeline
Project description
Fish&Feats 
Napari plugin to quantify 3D cells in a tissue and their smRNA-Fish or other RNA contents.
Fish&Feats offers several flexible options to analyse 3D cells and RNA counts, from segmentation of apical cells and nuclei to hierarchical clustering of cells based on their RNA contents. Installation/Usage are all described in the documentation.
Installation
Please refer to our documentation page for more details on the installation.
FishFeats is distributed as a pip module on pypi.
It can be installed by typing in a python virtual environement:
pip install fishfeats
Usage
You can launch fishfeats in Napari by going to Plugins>fishfeats>Start.
It will open a file dialog box asking you to select the image that you want to analyze.
Refer to the documentation for presentation of the different steps possible in the pipeline.
Test dataset
You can find in this zenodo repository https://zenodo.org/records/17048217 freely available images that can be used to test our pipeline. All the steps are fully documented in the online documentation and can be performed with one of these test images.
Example of analysis you can do with FishFeats are detailled step-by-step here and can be followed with the test image.
License
Fishfeats is distributed freely under the BSD-3 license.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file fishfeats-1.4.1.tar.gz.
File metadata
- Download URL: fishfeats-1.4.1.tar.gz
- Upload date:
- Size: 133.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.0.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9cf3348f8d6799ef716048db564de42645ef9e1b0135d3cd58aff7111861dd2a
|
|
| MD5 |
00308d50f852a2a3dfd07aee4dc5a527
|
|
| BLAKE2b-256 |
192cd7740b9f627e1575d347c49851fda54a8d5d2c40710aefbf2e6782164b37
|
File details
Details for the file fishfeats-1.4.1-py3-none-any.whl.
File metadata
- Download URL: fishfeats-1.4.1-py3-none-any.whl
- Upload date:
- Size: 139.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.0.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
02ee6a0f340d6384f9ee17b00292a586ee43ce8e5f3138b6463a6709136225c8
|
|
| MD5 |
38986e3c7c5bc29e3fcb05d6ef619f2b
|
|
| BLAKE2b-256 |
3418a84848c825501bf1fd8508afa11cea4e70b29228a67931ac718d1d4315e2
|