FluxFEM: A weak-form-centric differentiable finite element framework in JAX
Project description
FluxFEM
A weak-form-centric differentiable finite element framework in JAX, where variational forms are treated as first-class, differentiable programs.
Examples and Features
| Example 1: Diffusion | Example 2: Neo Neohookean Hyper Elasticity |
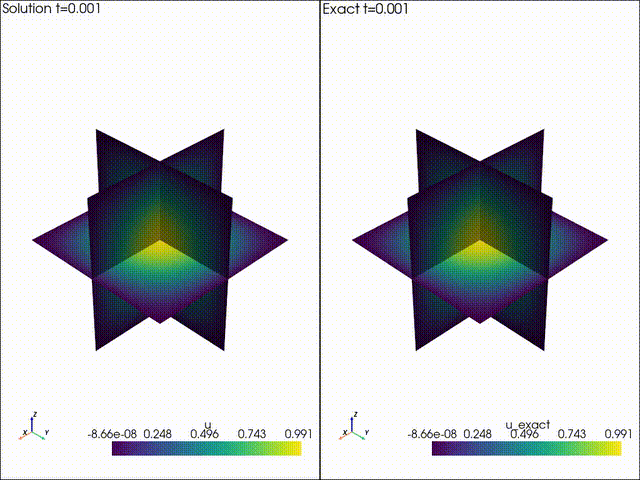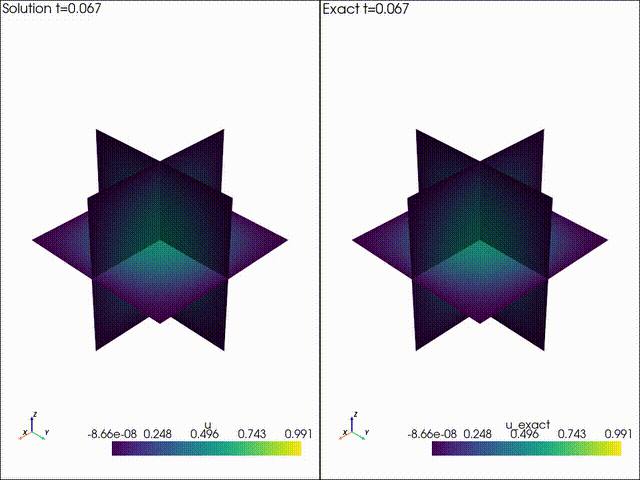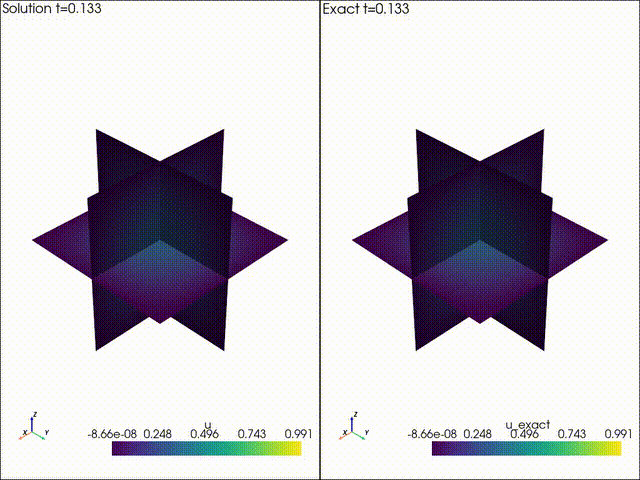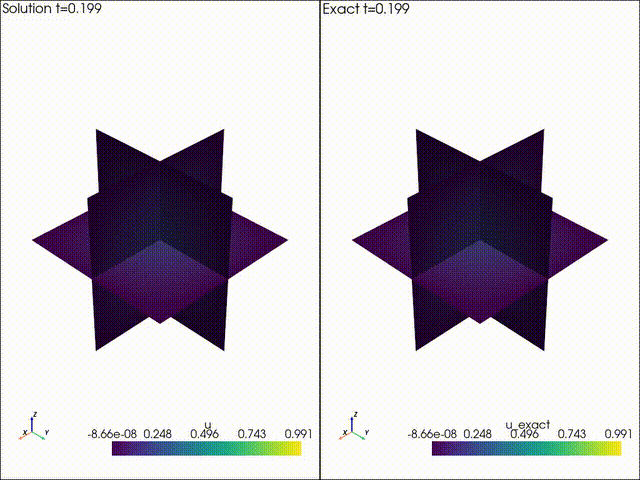
|
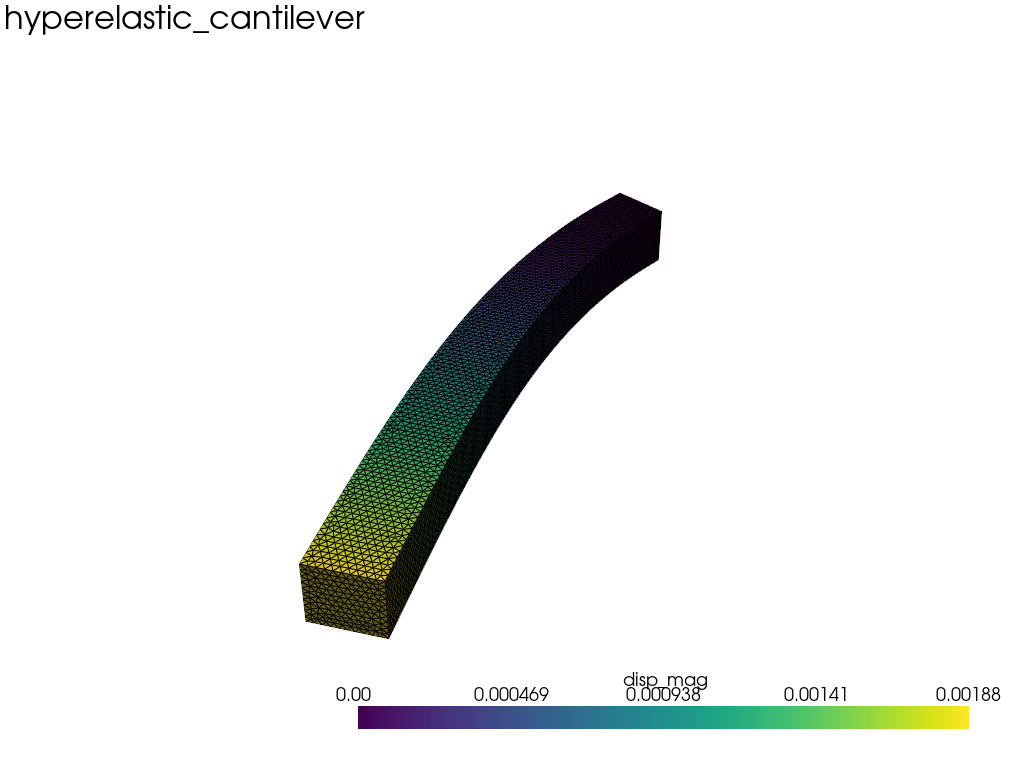
|
Features
- Built on JAX, enabling automatic differentiation with grad, jit, vmap, and related transformations.
- Weak-form–centric API that keeps formulations close to code; weak forms are represented as expression trees and compiled into element kernels, enabling automatic differentiation of residuals, tangents, and objectives.
- Two assembly approaches: tensor-based (scikit-fem–style) assembly and weak-form-based assembly.
- Handles both linear and nonlinear analyses with AD in JAX.
- Optional PETSc/PETSc-shell solvers via
petsc4pyfor scalable linear solves (addfluxfem[petsc]). - Contact interface support for penalty/constraint contact formulations, including role-explicit contact specs (
ContactSpaces,ContactGroupSpaces,OneSidedContactSpaces) and KKT assembly utilities (assemble_contact_coupling_matrices,assemble_contact_kkt,solve_contact_kkt).
Usage
This library provides two assembly approaches.
- A tensor-based assembly, where trial and test functions are represented explicitly as element-level tensors and assembled accordingly (in the style of scikit-fem).
- A weak-form-based assembly, where the variational form is written symbolically and compiled before assembly.
The two approaches are functionally equivalent and share the same element-level execution model,
but they differ in how you author the weak form. The example below mirrors the paper's diffusion
case and makes the distinction explicit with jnp.
Assembly Flow
All expressions are first compiled into an element-level evaluation plan, which operates on quadrature-point–major tensors. This plan is then executed independently for each element during assembly.
As a result, both assembly approaches:
- use the same quadrature-major (q, a, i) data layout,
- perform element-local tensor contractions,
- and are fully compatible with JAX transformations such as
jit,vmap, and automatic differentiation.
kernel-based assembly (explicit JIT units)
If you want to control JIT boundaries explicitly, build a JIT-compiled element kernel
and pass it to space.assemble. The kernel must return the integrated element
contribution (not the quadrature integrand). For untagged raw kernels, pass kind=.
import fluxfem as ff
import jax
import jax.numpy as jnp
space = ff.make_hex_space(mesh, dim=1, intorder=2)
# bilinear: kernel(ctx) -> (n_ldofs, n_ldofs)
ker_K = ff.make_element_bilinear_kernel(ff.diffusion_form, 1.0, jit=True)
K = space.assemble(ff.diffusion_form, 1.0, kernel=ker_K)
# linear: kernel(ctx) -> (n_ldofs,)
def linear_kernel(ctx):
integrand = ff.scalar_body_force_form(ctx, 2.0)
wJ = ctx.w * ctx.test.detJ
return (integrand * wJ[:, None]).sum(axis=0)
ker_F = jax.jit(linear_kernel)
F = space.assemble(ff.scalar_body_force_form, 2.0, kernel=ker_F)
tensor-based vs weak-form-based (diffusion example)
tensor-based assembly
The tensor-based assembly provides an explicit, low-level formulation with element kernels written using jax.numpy.(jnp).
import fluxfem as ff
import jax.numpy as jnp
@ff.kernel(kind="bilinear", domain="volume")
def diffusion_kernel(ctx: ff.FormContext, kappa):
# ctx.test.gradN / ctx.trial.gradN: (n_qp, n_nodes, dim)
# output tensor: (n_qp, n_nodes, n_nodes)
return kappa * jnp.einsum("qia,qja->qij", ctx.test.gradN, ctx.trial.gradN)
space = ff.make_hex_space(mesh, dim=3, intorder=2)
params = ff.Params(kappa=1.0)
K_ts = space.assemble(diffusion_kernel, params=params.kappa)
weak-form-based assembly
In the weak-form-based assembly, the variational formulation itself is the primary object. The expression below defines a symbolic weak form. FluxFEM compiles it internally during assembly, and you can still request an explicit compiled object when you want reuse/control.
import fluxfem as ff
import fluxfem.helpers_wf as h_wf
space = ff.make_hex_space(mesh, dim=3, intorder=2)
params = ff.Params(kappa=1.0)
U = ff.NamedSpace("U", space)
V = ff.NamedSpace("V", space)
# u, v are symbolic trial/test fields (weak-form DSL objects).
# u.grad / v.grad are symbolic nodes (expression tree), not numeric arrays.
form_wf = ff.BilinearForm.volume(
lambda u, v, p: p.kappa * (v.grad @ u.grad) * h_wf.dOmega()
)
# Shortest path for standard Galerkin assembly (U == V == space).
K_wf = space.assemble(form_wf, params=params)
# Equivalent explicit-role form when you want U/V to appear in the code.
K_wf_roles = ff.BilinearSpaces(test=V, trial=U).assemble(form_wf, params=params)
If you want to compile once and reuse explicitly:
compiled = ff.BilinearForm.volume(
lambda u, v, p: p.kappa * (v.grad @ u.grad) * h_wf.dOmega()
).get_compiled()
K_wf = space.assemble(compiled, params=params)
Linear Elasticity assembly (weak-form based assembly)
import fluxfem as ff
import fluxfem.helpers_wf as h_wf
space = ff.make_hex_space(mesh, dim=3, intorder=2)
D = ff.isotropic_3d_D(1.0, 0.3)
form_wf = ff.BilinearForm.volume(
lambda u, v, D: h_wf.ddot(v.sym_grad, D @ u.sym_grad) * h_wf.dOmega()
)
K = space.assemble(form_wf, params=D)
Neo-Hookean residual assembly (weak-form DSL)
Below is a Neo-Hookean hyperelasticity example written in weak form. The residual is expressed symbolically and compiled into element-level kernels executed per element. No manual derivation of tangent operators is required; consistent tangents (Jacobians) for Newton-type solvers are obtained automatically via JAX AD.
def neo_hookean_residual_wf(v, u, params):
mu = params["mu"]
lam = params["lam"]
F = h_wf.I(3) + h_wf.grad(u) # deformation gradient
C = h_wf.matmul(h_wf.transpose(F), F)
C_inv = h_wf.inv(C)
J = h_wf.det(F)
S = mu * (h_wf.I(3) - C_inv) + lam * h_wf.log(J) * C_inv
dE = 0.5 * (h_wf.matmul(h_wf.grad(v), F) + h_wf.transpose(h_wf.matmul(h_wf.grad(v), F)))
return h_wf.ddot(S, dE) * h_wf.dOmega()
res_form = ff.ResidualForm.volume(neo_hookean_residual_wf)
R = space.assemble_residual(res_form, u, ff.Params(mu=1.0, lam=1.0))
J = space.assemble_jacobian(res_form, u, ff.Params(mu=1.0, lam=1.0))
autodiff + jit compile
You can differentiate through the solve and JIT compile the hot path. The inverse diffusion tutorial shows this pattern:
def loss_theta(theta):
kappa = jnp.exp(theta)
u = solve_u_jit(kappa, traction_true)
diff = u[obs_idx_j] - u_obs[obs_idx_j]
return 0.5 * jnp.mean(diff * diff)
solve_u_jit = jax.jit(solve_u)
loss_theta_jit = jax.jit(loss_theta)
grad_fn = jax.jit(jax.grad(loss_theta))
FESpace vs FESpacePytree
Use FESpace for standard workflows with a fixed mesh. When you need to carry
the space through JAX transformations (e.g., shape optimization where mesh
coordinates are part of the computation), use FESpacePytree via
make_*_space_pytree(...). This keeps the mesh/basis in the pytree so
jax.jit/jax.grad can see geometry changes.
In other words:
- fixed-geometry solve/assembly:
FESpace - geometry-sensitive differentiation:
FESpacePytree
The mesh-move example in tutorials/diffusion_3d_mesh_proxy.py
computes jax.grad(...) with respect to node coordinates on top of
make_hex_space_pytree(...).
Current boundary:
- geometry-dependent objectives and residual-style quantities can be differentiated in JAX when the geometry is carried through a pytree space
- there is not yet a dedicated public "shape derivative" API layer; shape sensitivity is currently expressed as ordinary JAX differentiation through assembly/solve code
backend="numpy"is not part of this differentiable path
For same-space Galerkin assembly, space.assemble(...) and space.assemble_*
remain the shortest paths.
When you want the roles to be explicit, prefer
LinearSpaces(...).assemble(...), BilinearSpaces(...).assemble(...),
ResidualSpaces(...).assemble(...), and JacobianSpaces(...).assemble(...).
For contact interfaces, prefer contact.assemble_* methods over top-level
ff.assemble_contact_* helpers. The top-level assemble_* helpers remain
available as compatibility entrypoints.
Mixed systems
Mixed systems use two kinds of names:
- field name: the global block name in the mixed vector, such as
"u"or"p" - FE space: the discrete space where that field lives
Start from the mathematical picture:
- unknown
ulives in FE spaceV - unknown
plives in FE spaceQ
V = ff.make_hex_space(mesh, dim=3, intorder=2)
Q = ff.make_hex_space(mesh, dim=1, intorder=2)
# "u in V" (the name "u" is the mixed field name)
u_field = ff.NamedSpace("u", V)
# "p in Q"
p_field = ff.NamedSpace("p", Q)
mixed = ff.MixedSpace(u_field, p_field)
Read that as:
"u"and"p"are the field names in the global mixed vectorVandQare the actual FE spaces for those fields
So ff.NamedSpace("u", V) means:
- this mixed field is called
"u" - it lives in FE space
V
In the common mixed API, field names are also the default symbolic names used inside weak forms. That keeps the notation close to the mathematical picture.
If you want to make the per-field roles explicit, MixedSpace(...) can also
reuse the existing single-field role specs:
U = ff.NamedSpace("u", V)
Vt = ff.NamedSpace("v_u", V)
P = ff.NamedSpace("p", Q)
Qt = ff.NamedSpace("v_p", Q)
mixed = ff.MixedSpace(
u=ff.ResidualSpaces(test=Vt, unknown=U),
p=ff.BilinearSpaces(test=Qt, trial=P),
)
That reads as:
- the
ufield usesVfor both test and unknown, with explicit symbolic namesv_uandu - the
pfield usesQfor both test and trial, with explicit symbolic namesv_pandp
Next, define one residual function per equation. Inside those functions, use the
field names when you want to refer to another unknown symbolically.
Here unknown_ref("p") means "the symbolic mixed unknown stored under field name p".
The test function is usually just the first local argument (v, q), so a separate
test-side lookup is often unnecessary.
def res_u(v, u, p):
p_ref = ff.unknown_ref("p")
return (...) * h_wf.dOmega()
def res_p(q, p_field, p):
u_ref = ff.unknown_ref("u")
return (...) * h_wf.dOmega()
For the common case where residual names and field names match, keep it short:
residuals = ff.make_mixed_residuals(
u=res_u,
p=res_p,
)
If you need to route an equation to a different field name explicitly, use
bind_mixed_residual(...).
Once the field layout and residual bindings are defined, assembly follows the same object-centered flow as the single-space API:
res_form = ff.ResidualForm.mixed(residuals)
R = mixed.assemble_residual(res_form, u0, ff.Params(alpha=1.0))
J = mixed.assemble_jacobian(res_form, u0, ff.Params(alpha=1.0))
If you prefer a higher-level problem object, MixedProblem(...) is still available.
Contact weak forms
Contact follows the same broad assembly pattern as the volume and surface APIs:
- define a contact spec
- prepare a reusable interface object
- optionally update a contact state at the current iterate
- assemble interface operators or contact contributions
The main difference from standard volume assembly is that contact has a state-dependent geometric part. In FluxFEM terms:
ContactPairSpec/ContactGroupSpec/OneSidedContactSpecare declarative specsprepare(...)performs the heavy reusable setupinitialize_state()/update_state(...)expose the state-explicit nonlinear path
FluxFEM contact assembly is expressed in residual/jacobian form. When comparing
against references written directly as K u = f bilinear/linear forms, a global
sign flip may be required to match conventions; this is expected if the physics
is otherwise identical.
In the current implementation, prepare(...) is the preferred public alias for
the older to_contact_surface_space(...) setup step.
Pair Contact Quickstart
Start by defining a contact side on each body, then combine them into a contact spec and prepare a reusable interface:
master_side = ff.ContactSideSpec.from_facets(master_mesh, master_facets, master_space)
slave_side = ff.ContactSideSpec.from_facets(slave_mesh, slave_facets, slave_space)
contact = ff.ContactPairSpec(
master=master_side,
slave=slave_side,
).prepare(
quad_order=1,
backend="jax",
)
Internally, this setup step prepares reusable interface data such as pairing, supermesh geometry, facet-to-element maps, and quadrature caches.
Once you have a prepared contact interface, bilinear weak forms follow the same object-centered pattern:
contact_form = ff.BilinearForm.contact(a_contact)
B = contact.assemble_bilinear_form(contact_form, params)
If you want an explicit compiled object for reuse:
compiled = ff.BilinearForm.contact(a_contact).get_compiled()
B = contact.assemble_bilinear_form(compiled, params)
State-Explicit Nonlinear Contact
For curved surfaces, load stepping, or nonlinear contact updates, expose the state-dependent part explicitly:
contact_state = contact.initialize_state()
contact_state = contact.update_state(
state={"a": u_master, "b": u_slave},
contact_state=contact_state,
geometry="current",
active_set="update",
)
The current ContactState object is a lightweight public skeleton for this
workflow. It records the interface kind, iteration, geometry mode, and a shape
summary of the current field state. This is the intended public boundary for
future state-dependent contact updates.
Penalty Contact
Penalty-family contact assembly uses the same prepared contact object:
ops_penalty = contact.assemble_penalty(
weak_form=contact_residual_form,
state={"a": u_master, "b": u_slave},
params=params,
backend="jax",
)
Multiplier Contact
Constraint-family contact adds a multiplier spec on top of the same interface:
lm_space = ff.MultiplierSpec.from_contact(
contact,
family="p0",
side="master",
)
ops_constraint = contact.assemble_multiplier(
rho=1.0,
multiplier=lm_space,
backend="numpy",
)
So the usual flow is:
- create
ContactSideSpecobjects from mesh facets - build a spec with
ContactPairSpec(...) - call
prepare(...)to build reusable interface setup - optionally call
initialize_state()/update_state(...)in nonlinear loops - choose penalty or multiplier assembly on that prepared contact object
Mixed weak-form naming follows this convention:
- simple single-space code:
ctx.test/ctx.trial - named mixed field lookup:
ctx.bindings["u"] - explicit space-key lookup:
ctx.spaces["V"] - explicit residual-to-field routing:
bind_mixed_residual(...)
Use the explicit forms only where they help readability or avoid ambiguity. Examples:
Backend notes
backend="jax" is the primary path for differentiation and Jacobian assembly.
backend="numpy" is available mainly for forward assembly/evaluation and comparison/debug workflows.
Today, the practical split is:
jax: bilinear/linear/residual assembly, autodiff-based Jacobians, geometry-sensitive differentiationnumpy: bilinear/linear/residual/functional forward assembly in many paths, plus several contact/coupled utilitiesnumpyJacobian assembly is not generally implemented; for exampleassemble_jacobian(..., backend="numpy")is not available
For contact/supermesh code, backend="numpy" is also used in places where the Jacobian is approximated by finite differences rather than differentiated symbolically.
Block assembly
For constraints like contact problems (e.g., adding Lagrange multipliers), build a block matrix explicitly:
from fluxfem import solver as ff_solver
# Example blocks from contact coupling
K_uu = ...
K_cc = ...
K_uc = ...
blocks = ff_solver.make_block_matrix(
diag=ff_solver.block_diag(order=("u", "c"), u=K_uu, c=K_cc),
rel={("u", "c"): K_uc},
symmetric=True,
transpose_rule="T",
)
# Lazy container; assemble when you need the global matrix.
K = blocks.assemble()
FluxFEM also provides high-level contact utilities:
# Pair contact
side_master = ff.ContactSideSpec.from_surfaces(surf_master, elem_conn=conn_master, value_dim=3)
side_slave = ff.ContactSideSpec.from_surfaces(surf_slave, elem_conn=conn_slave, value_dim=3)
contact = ff.ContactPairSpec(master=side_master, slave=side_slave).prepare(
quad_order=4,
backend="jax",
)
# One-to-many contact
contact_group = ff.ContactGroupSpec(
master=side_master,
slaves=[side_slave],
).prepare(
quad_order=4,
backend="jax",
)
# One-sided contact
floor_contact = ff.OneSidedContactSpec(side=side_slave).prepare(
quad_order=4,
)
# Optional nonlinear/state-explicit update
contact_state = contact.initialize_state()
contact_state = contact.update_state(
state={"a": u_master, "b": u_slave},
contact_state=contact_state,
geometry="current",
active_set="update",
)
# 1) Assemble constraint operators (B, Kuu, ...)
lm_space = ff.MultiplierSpec.from_contact(
contact_group,
family="p0_supermesh",
side="master",
)
ops: ff.ContactOperators = contact_group.assemble_multiplier(
rho=1.0,
multiplier=lm_space,
backend="numpy",
# Optional: also evaluate and store residual/jacobian metadata on the same ContactOperators.
weak_form=contact_residual_form,
state={"a": u_master, "b": u_slave},
params=params,
)
# 2) Penalty-family path: user weak form -> residual/jacobian operators
ops_nitsche: ff.ContactOperators = contact.assemble_penalty(
weak_form=contact_residual_form,
state={"a": u_master, "b": u_slave},
params=params,
backend="jax",
)
# 3) Unified coupled API (Penalty Family)
builder = ff.CoupledSystemBuilder.from_structural(K_u, F_u)
builder.register_blocks([
("master", space_master, {"value_dim": 1}),
("slave", space_slave, {"value_dim": 1}),
])
builder.add_contact(
ops_nitsche,
master="master",
slave="slave",
value_dim=1,
)
system = builder.build()
u = system.solve(dirichlet_dofs=dir_dofs, dirichlet_vals=0.0, format="csr")
# 4) Unified coupled API (Constraint Family): KKT assembly is internal to builder
builder_mortar = ff.CoupledSystemBuilder.from_structural(K_u, F_u)
builder_mortar.register_blocks([
("master", space_master, {"value_dim": 1}),
("slave", space_slave, {"value_dim": 1}),
])
builder_mortar.add_contact(
ops,
master="master",
slave="slave",
value_dim=1,
)
system_mortar = builder_mortar.build()
# Advanced: law/formulation can be set explicitly when needed.
# - law="one_sided_normal_frictionless"
# - formulation="multiplier" | "penalty_consistent"
Contact API boundaries (fixed terms):
spec: declarative contact definition (ContactPairSpec,ContactGroupSpec,OneSidedContactSpec).contact: prepared interface geometry/pairing/supermesh/quadrature.contact_state: current-iterate contact state for nonlinear workflows.multiplier: LM discretization (family,side,value_dim).formulation: enforcement variant used for routing.ops: assembled bundle passed toCoupledSystemBuilder.
Notes:
- Multiple contacts can be added with different settings per
builder.add_contact(...)call. prepare(...)is the preferred public name for the reusable setup step;to_contact_surface_space(...)remains available as a compatibility entrypoint.ContactStatecurrently provides the public skeleton for state-explicit contact workflows; future nonlinear contact updates should build on this boundary.MultiplierSpecis the preferred public name for multiplier discretization;ContactMultiplierSpaceremains available as a compatibility name.ContactMultiplierSpace/MultiplierSpecp0-like families ("p0","p0_active","p0_supermesh") currently supportside="master"only.- See docs:
Usage -> Contact API Boundaries.
Documentation
Full documentation, tutorials, and API reference are hosted at this site.
Tutorials
tutorials/linearelastic_tensile_bar.py(linear elasticity, weak-form assembly)tutorials/neo_hookean_cantilever.py(nonlinear hyperelasticity)tutorials/thermoelastic_bar_1d.py/tutorials/thermoelastic_bar_1d_mixed.py(thermoelastic coupling)tutorials/contact_supported_box_by_pillars.py(large box supported by multiple small boxes via penalty contact + Dirichlet supports)tutorials/petsc_shell_poisson_demo.py(PETSc shell solver integration; see alsotutorials/petsc_shell_poisson_pmat_demo.py)
Setup
You can install FluxFEM either via pip or Poetry.
Supported Python Versions
FluxFEM supports Python 3.11–3.13:
Choose one of the following methods:
Using pip
pip install fluxfem
pip install "fluxfem[cuda12]" -f https://storage.googleapis.com/jax-releases/jax_cuda_releases.html
Using poetry
poetry add fluxfem
poetry add fluxfem[cuda12]
PETSc Integration
Optional PETSc-based solvers are available via petsc4py. Enable with the extra:
pip install "fluxfem[petsc]"
or
pip install "fluxfem[petsc,cuda12]" -f https://storage.googleapis.com/jax-releases/jax_cuda_releases.html
poetry add fluxfem --extras "petsc"
or
poetry add "fluxfem[petsc,cuda12]"
or
poetry add fluxfem --extras "petsc" --extras "cuda12"
Note: you must match the petsc4py version to the PETSc version you have
installed. The current FluxFEM extra pins petsc4py==3.24.4 (see
[project.optional-dependencies]), so make sure your PETSc install is
compatible with that petsc4py release, or override it to match your PETSc
build.
GPU note: this repo currently tests CUDA via the cuda12 extra only. Other CUDA
versions are not covered by CI and may require manual JAX installation.
Acknowledgements
I acknowledge the open-source software, libraries, and communities that made this work possible.
Citation
Reference to cite if you use FluxFEM in a paper:
@software{Watanabe_FluxFEM_2026,
author = {Watanabe, Kohei},
doi = {10.5281/zenodo.18734689},
month = {2},
title = {{FluxFEM}},
url = {https://github.com/kevin-tofu/fluxfem},
year = {2026}
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file fluxfem-0.3.5.tar.gz.
File metadata
- Download URL: fluxfem-0.3.5.tar.gz
- Upload date:
- Size: 251.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.3 CPython/3.11.15 Linux/6.17.0-1008-azure
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2218e15095e604e0435db08fec51b22411507b50e6a37de1e6b144d63dc9a68f
|
|
| MD5 |
9ecbac6b2f667cdbb4f8dbf9c8294423
|
|
| BLAKE2b-256 |
30b1fbe83e3d08ed484cd8412a081eedec4507937c2f97f63247307c7f805064
|
File details
Details for the file fluxfem-0.3.5-py3-none-any.whl.
File metadata
- Download URL: fluxfem-0.3.5-py3-none-any.whl
- Upload date:
- Size: 276.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.3 CPython/3.11.15 Linux/6.17.0-1008-azure
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0d0e32957eb9e87f434b91070b4aef87d8e76b69dbb169f0f0599e05afa6ddb6
|
|
| MD5 |
d32fb51ad6e1c42a3d30c8e0c094f486
|
|
| BLAKE2b-256 |
4f55bc6c4c25fd757b77bad4d6dadc3e733e80607c56ef04cc8a2d3c4219e65c
|

















