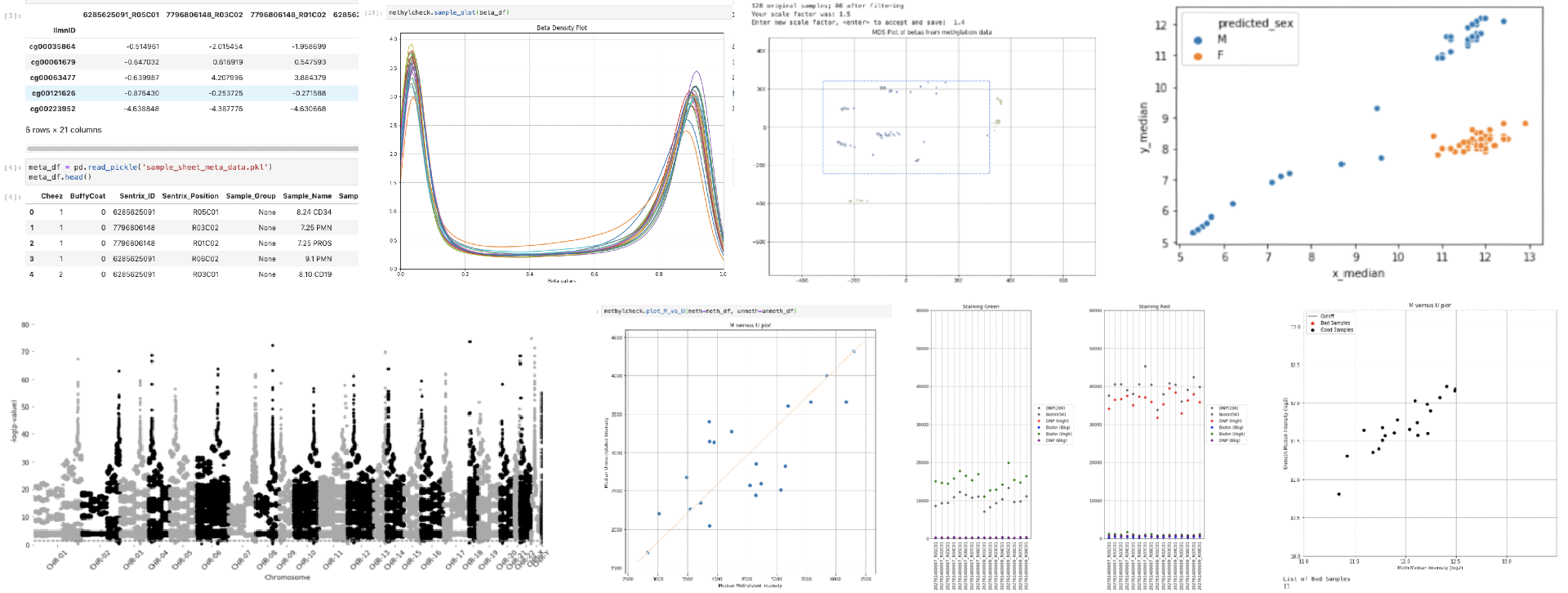
Quality Control (QC), Visualization/plotting, and postprocessing software for Illumina methylation array data. See https://life-epigenetics-methylcheck.readthedocs-hosted.com/en/latest/ for full documentation and examples.
Project description
methylcheck is a Python-based package for filtering and visualizing Illumina methylation array data. The focus is on quality control.
methylcheck Package
This package contains high-level APIs for filtering processed data from local files. 'High-level' means that the details are abstracted away, and functions are designed to work with a minimum of knowledge and specification required. But you can always override the "smart" defaults with custom settings if things don't work. Before starting you must first download processed data from the NIH GEO database or process a set of idat files with methylprep. Refer to methylprep for instructions on this step.
Installation
This package is available in PyPi.
pip install methylcheck or pip3 install methylcheck if your OS defaults to python2x. This package only works in python3.6+.
Importing your data
Methylcheck is designed to accept the output from the methylprep package. If you have a bunch of idat samples, methylprep will return a single pickled pandas dataframe containing all the beta values for probes.
Load your data in a Jupyter Notebook like this:
mydata = pandas.read_pickle('beta_values.pkl')
If you processed a large batch of samples using the batch_size option in methylprep process, there's a convenience function in methylcheck (methylcheck.load) that will load and combine a bunch of output files in the same folder:
import methylize
df = methylcheck.load('<path to folder with methylprep output>')
# or
df,meta = methylcheck.load_both('<path to folder with methylprep output>')
This conveniently loads a dataframe of all meta data associated with the samples, if you are using public GEO data. Some analysis functions require specifying which samples are part of a treatment group (vs control) and the meta dataframe object can be used for this.
For more, check out our examples of loading data into methylcheck
GEO
Alternatively, you can import public GEO datasets directly, if they are processed data containing either probe beta values for samples or methylated/unmethylated signal intensities. If you have idat files, process them first with methylprep, or use the methylprep download -i <GEO_ID> option to download and process public data.
In general, the best way to import data is to use methylprep and run
run_pipeline(data_folder, betas=True)
# or from the command line:
python -m methylprep process -d <filepath to idats> --all
collect the beta_values.pkl file it returns/saves to disk, and load that in a Jupyter notebook. From there, each data transformation is a single line of code using Pandas DataFrames. methylcheck will keep track of the data format/structures for you, and you can visualize the effect of each filter as you go. You can also export images of your charts for publication.
Refer to the Jupyter notebooks on readthedocs for examples of filtering probes from a batch of samples, removing outlier samples, and generating plots of data.
Quality Control (QC)
The simplest way to generate a battery of plots about your data is to run this function in a Jupyter notebook:
import methylcheck
methylcheck.run_qc('<path to your methylprep processed files>')
Other functions
methylcheck provides functions to
- predict the sex of samples (
.get_sex) - detect probes that differ between two sets of samples within a batch (
.diff_meth_probes) - remove sex-chromosome-linked probes and control probes
- remove "sketchy" probes, deemed unreliable by researchers
- filter sample outliers based on multi-dimensional scaling
- combine datasets for analysis
- plot sample beta or m-value distributions, or raw uncorrected probe channel intensities
Authors
Parts of this package were ported from minfi, an R package, and extended/developed by the team at Foxo Bioscience, who maintains it. You can write to info@LifeEgx.com to give feedback, ask for help, or suggest improvements. For bugs, report issues on our github repo page.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file methylcheck-0.5.9.tar.gz.
File metadata
- Download URL: methylcheck-0.5.9.tar.gz
- Upload date:
- Size: 37.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.14.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/45.1.0.post20200127 requests-toolbelt/0.9.1 tqdm/4.42.0 CPython/3.7.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cc5e8dc821e0e247d344a8b58ddcf753b9b6fe03a99e5f869ffdb28571b88d2f
|
|
| MD5 |
d027e2f53c28fb9b8af12cb7062d6eea
|
|
| BLAKE2b-256 |
fbb6b9faecf103ba9daa44393663f71eae61fbf00b655b59302a66ba4e3786eb
|
File details
Details for the file methylcheck-0.5.9-py3-none-any.whl.
File metadata
- Download URL: methylcheck-0.5.9-py3-none-any.whl
- Upload date:
- Size: 8.8 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.14.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/45.1.0.post20200127 requests-toolbelt/0.9.1 tqdm/4.42.0 CPython/3.7.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
462462ea0dd76da007173e5936690cfa0ab45a8154f739e1b529135cdad6a529
|
|
| MD5 |
4c7fa0ad7e72bea0f3f8cec9a94549bd
|
|
| BLAKE2b-256 |
9080765462c336ed867ad23b34ce95f8af9962bd480465b3833b1eed5e619df2
|