A plugin to automatically count lung organoids using Deep Learning.
Project description
Napari Organoid Counter - Version 0.2.9 is out!
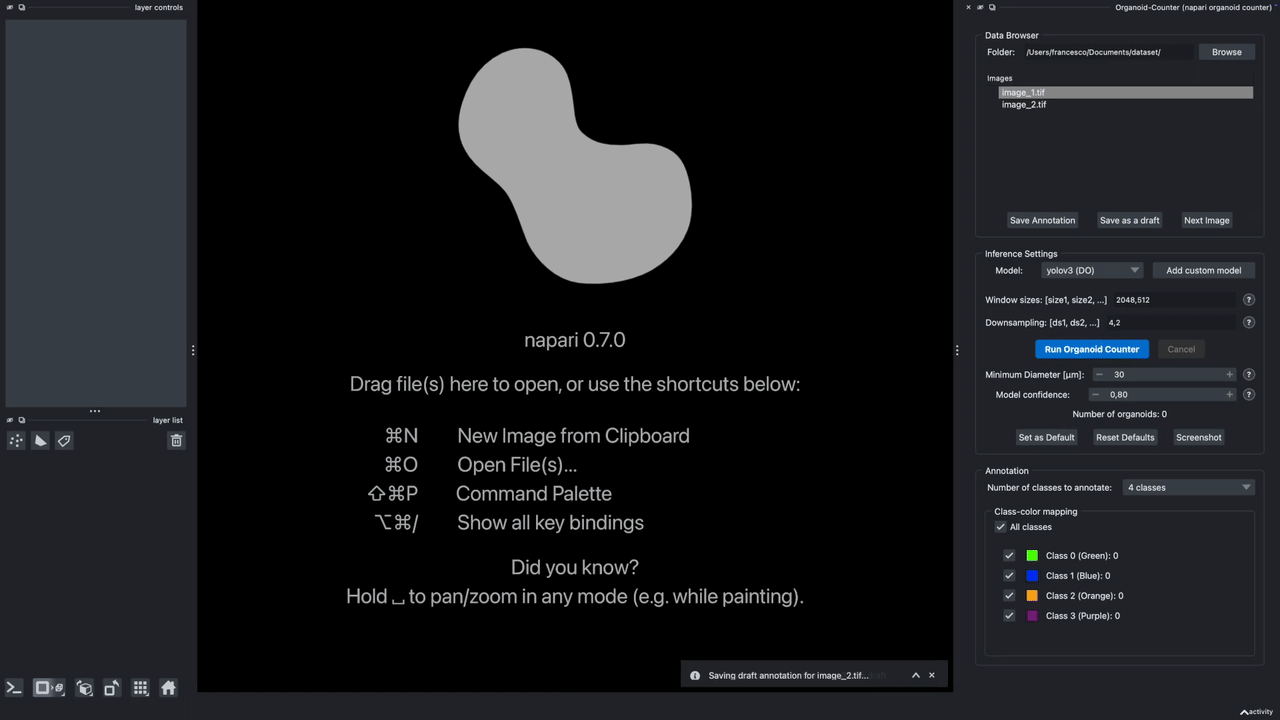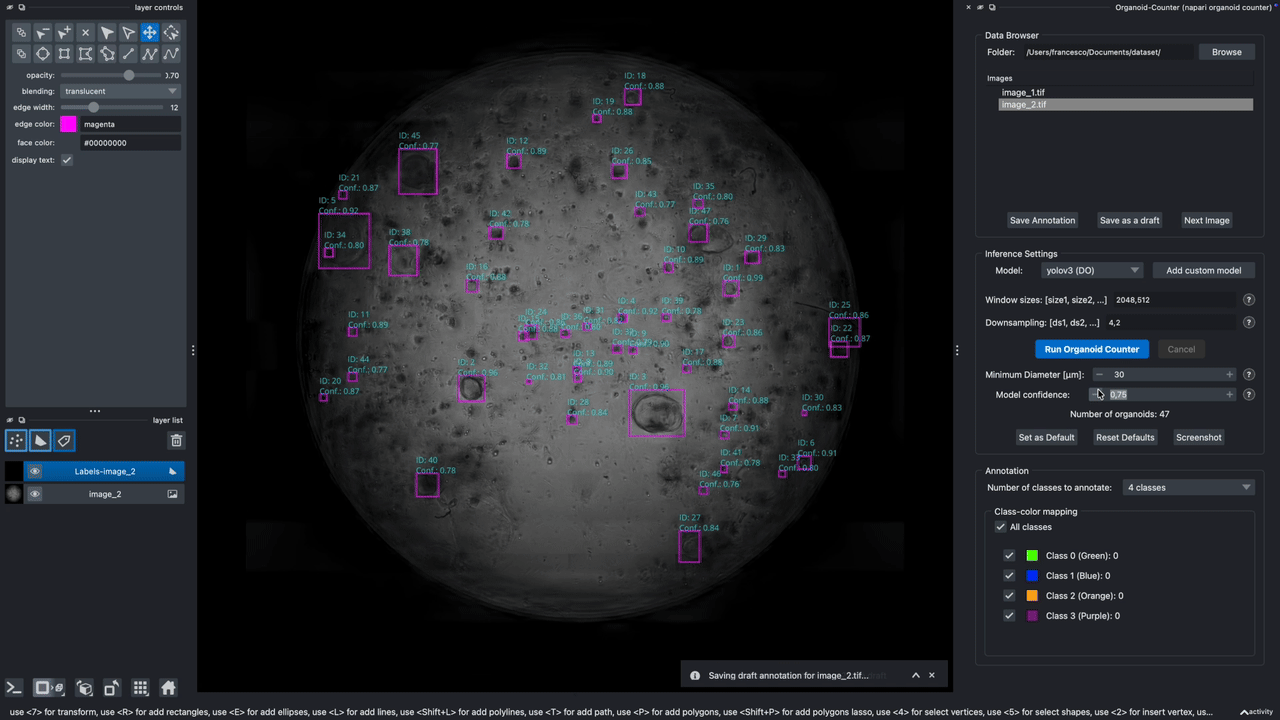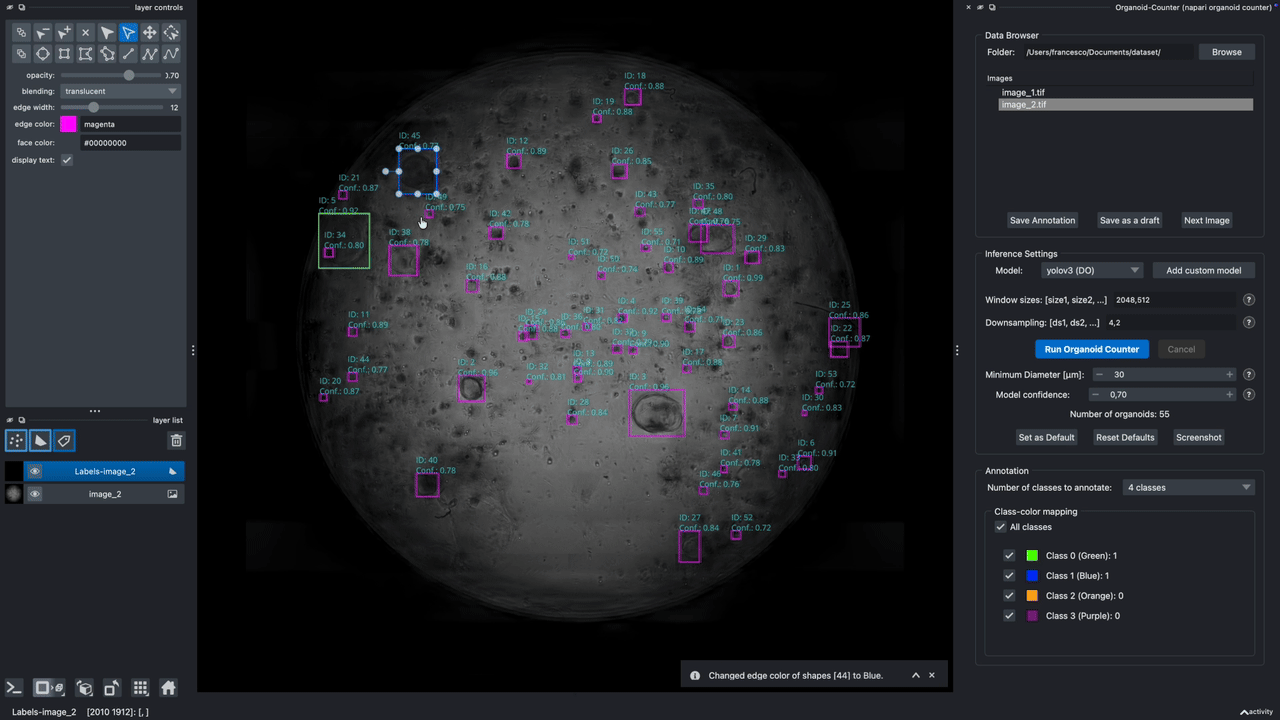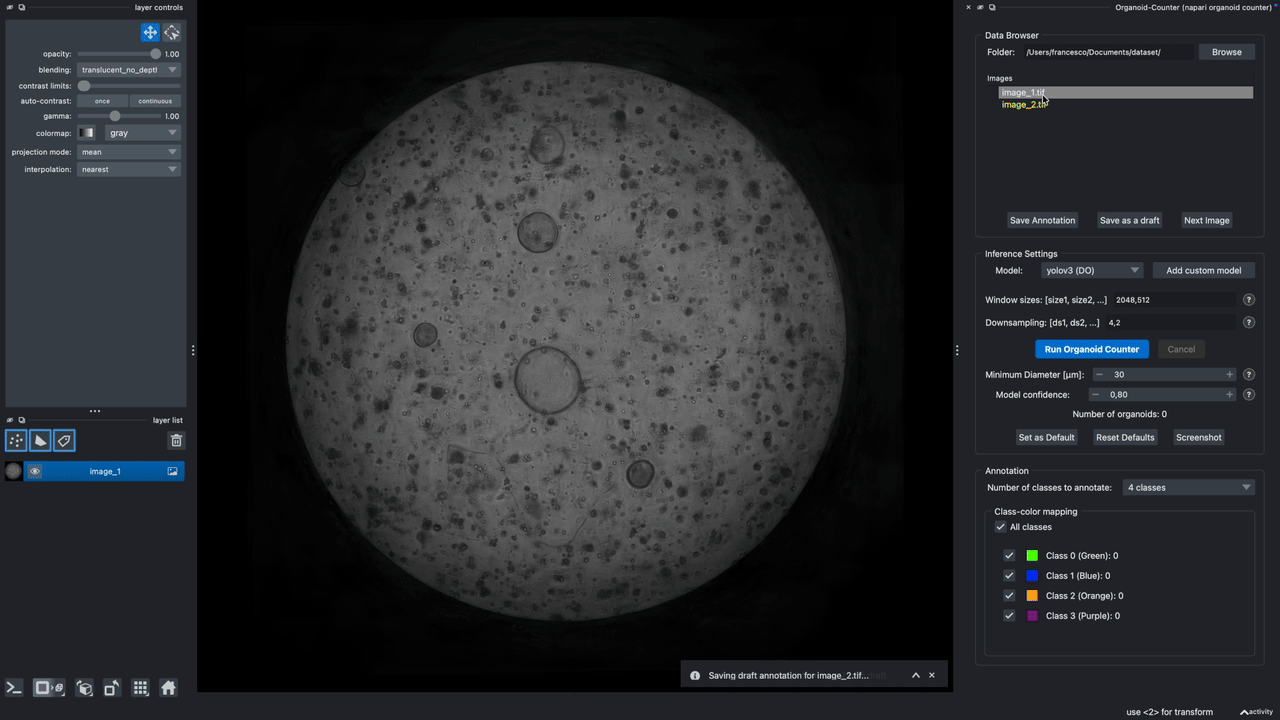
A napari plugin to automatically count lung organoids from microscopy imaging data. Note: this plugin only supports single-channel grayscale images.
Hold it for the demo!
This demo showcases the features of Version 0.2.9.
This napari plugin was generated with Cookiecutter using @napari's cookiecutter-napari-plugin template.
Installation
This plugin has been tested with python 3.11 - you may consider using conda or mamba to create your dedicated environment before running the napari-organoid-counter.
-
You can install
napari-organoid-countervia pip:pip install napari-organoid-counterTo install for a developer:
git clone https://github.com/HelmholtzAI-Consultants-Munich/napari-organoid-counterpip install -e .
For installing on a Windows machine directly from within napari, follow the instuctions here.
Latest version features
Checkout Features of the latest version here.
How to use?
- To launch the plugin directly from your terminal:
napari -w napari-organoid-counter
- For convenience you can add a shell alias by adding this to your
~/.zshrc(macOS) or~/.bashrc(Linux):
alias organoid='napari -w napari-organoid-counter'
Then reload your shell config:
source ~/.zshrc # macOS
source ~/.bashrc # Linux
You can now launch the plugin simply by running:
organoid
- Alternatively, you can also start napari manually and select the plugin from the drop down menu.
For more information on this plugin, its intended audience, and a Quickstart guide go to our Quickstart guide.
Contributing
Contributions are very welcome. Tests can be run with [pytest], please ensure the coverage at least stays the same before you submit a pull request.
License
Distributed under the terms of the MIT license, "napari-organoid-counter" is free and open source software
Dependencies
napari-organoid-counter uses the BioIO[1] for reading and processing images.
[1] Eva Maxfield Brown, Dan Toloudis, Jamie Sherman, Madison Swain-Bowden, Talley Lambert, Sean Meharry, Brian Whitney, AICSImageIO Contributors (2023). BioIO: Image Reading, Metadata Conversion, and Image Writing for Microscopy Images in Pure Python [Computer software]. GitHub. https://github.com/bioio-devs/bioio
Issues
If you encounter any problems, please file an issue along with a detailed description.
Citing
If you use this plugin for your work, please cite it using the following:
Christina Bukas, Francesco Campi, Abdulkader Ghandoura, & Piraud, M. (2026). HelmholtzAI-Consultants-Munich/napari-organoid-counter: v0.2.6 (v0.2.6). Zenodo. https://doi.org/10.5281/zenodo.19824587
bibtex:
@software{christina_bukas_2026_19824587,
author = {Christina Bukas and
Francesco Campi and
Abdulkader Ghandoura and
Piraud, Marie},
title = {HelmholtzAI-Consultants-Munich/napari-organoid-
counter: v0.2.6
},
month = apr,
year = 2026,
publisher = {Zenodo},
version = {v0.2.6},
doi = {10.5281/zenodo.19824587},
url = {https://doi.org/10.5281/zenodo.19824587},
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_organoid_counter-0.2.9rc1.tar.gz.
File metadata
- Download URL: napari_organoid_counter-0.2.9rc1.tar.gz
- Upload date:
- Size: 79.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f5accb9cc03092cc8f1e6e0c1e10029319982c3523b72400021d9e6d950ae602
|
|
| MD5 |
81ad52cd94254254bf7792bf08659c06
|
|
| BLAKE2b-256 |
7dd3d7dfa10f6ccba0d4db0b1154824e8012bf722a4047cabc7c416e2949e9c4
|
File details
Details for the file napari_organoid_counter-0.2.9rc1-py3-none-any.whl.
File metadata
- Download URL: napari_organoid_counter-0.2.9rc1-py3-none-any.whl
- Upload date:
- Size: 50.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d24f8984d8a303bf9722f6b41083637e32f0aeb985dfb56fd180dfb794d63982
|
|
| MD5 |
94ecb867679af1f77430e9d8578ff618
|
|
| BLAKE2b-256 |
0c24f794db32c7c1933ca11335e971dca039c625b1a3d09da5015ea776f4d0f5
|