A plugin based on scikit-image for segmenting nuclei and cells based on fluorescent microscopy images with high intensity in nuclei and/or membranes
Project description
napari-segment-blobs-and-things-with-membranes (nsbatwm)
This napari-plugin is based on scikit-image and allows segmenting nuclei and cells based on fluorescence microscopy images with high intensity in nuclei and/or membranes.
Usage
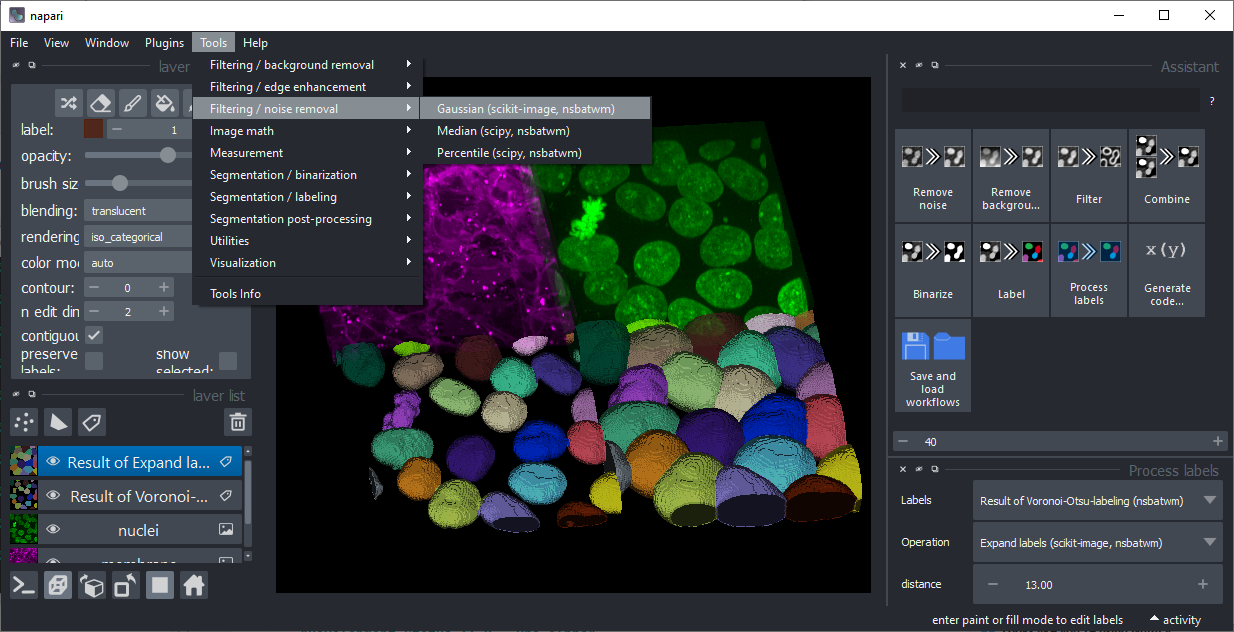
This plugin populates image processing operations to the Tools menu in napari.
You can recognize them with their suffix (nsbatwm) in brackets.
Furthermore, it can be used from the napari-assistant graphical user interface.
Therefore, just click the menu Tools > Utilities > Assistant (na) or run naparia from the command line.
You can also call these functions as shown in the demo notebook.
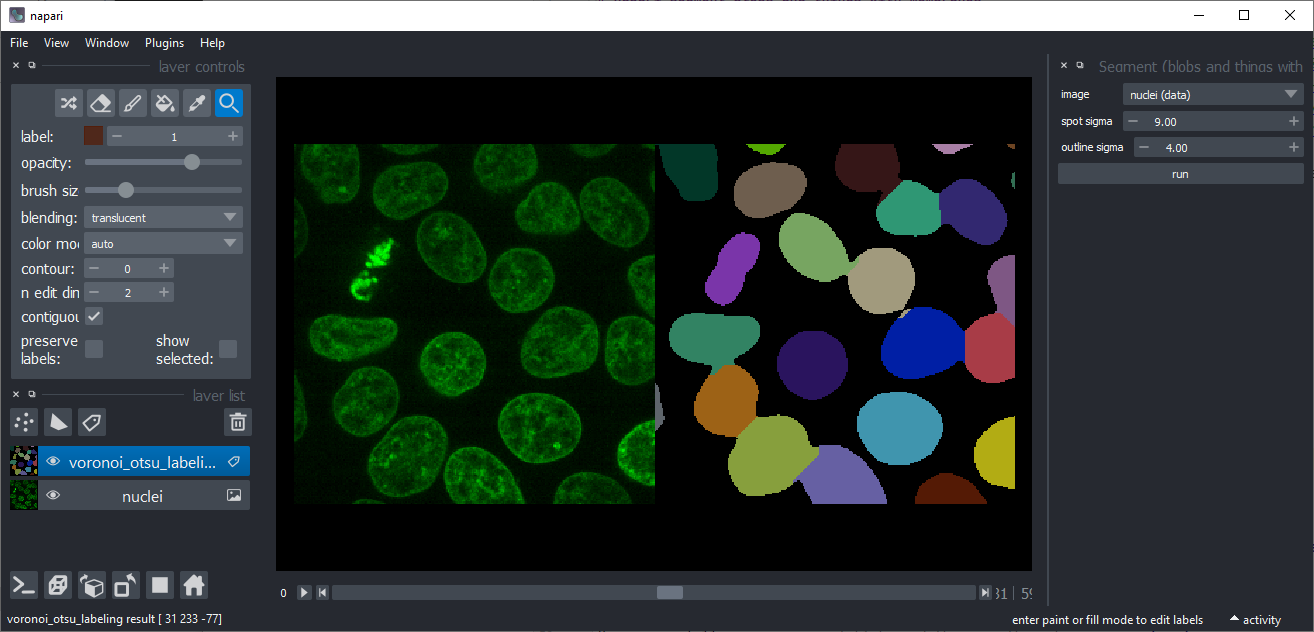
Voronoi-Otsu-Labeling
This algorithm uses Otsu's thresholding method in combination with
Gaussian blur and a
Voronoi-Tesselation
approach to label bright objects such as nuclei in an intensity image. The alogrithm has two sigma parameters which allow
you to fine-tune where objects should be cut (spot_sigma) and how smooth outlines should be (outline_sigma).
This implementation aims to be similar to Voronoi-Otsu-Labeling in clesperanto.
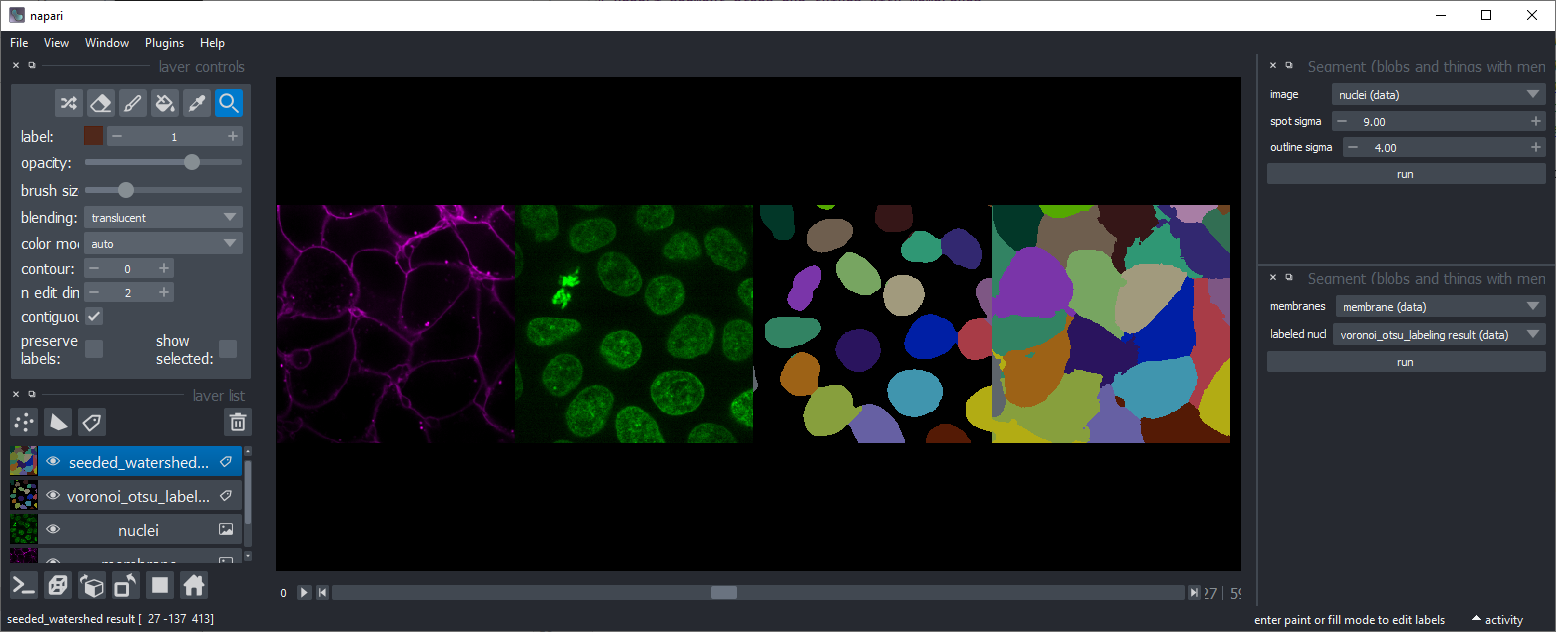
Seeded Watershed
Starting from an image showing high-intensity membranes and a seed-image where objects have been labeled (e.g. using Voronoi-Otsu-Labeling), objects are labeled that are constrained by the membranes.
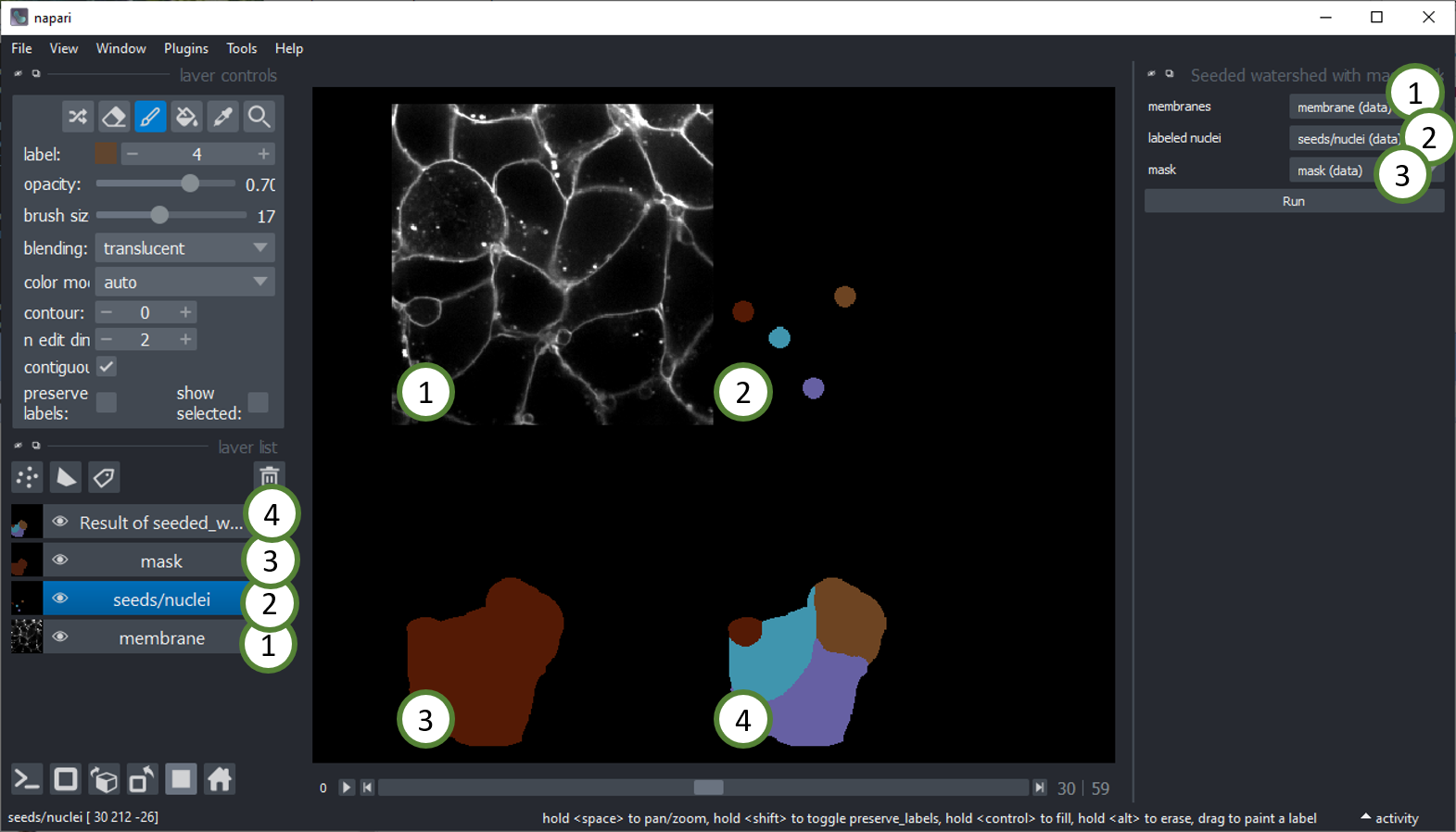
Seeded Watershed with mask
If there is additionally a mask image available, one can use the Seeded Watershed with mask, to constraint the flooding
on a membrane image (1), starting from nuclei (2), limited by a mask image (3) to produce a cell segmentation within the mask (4).
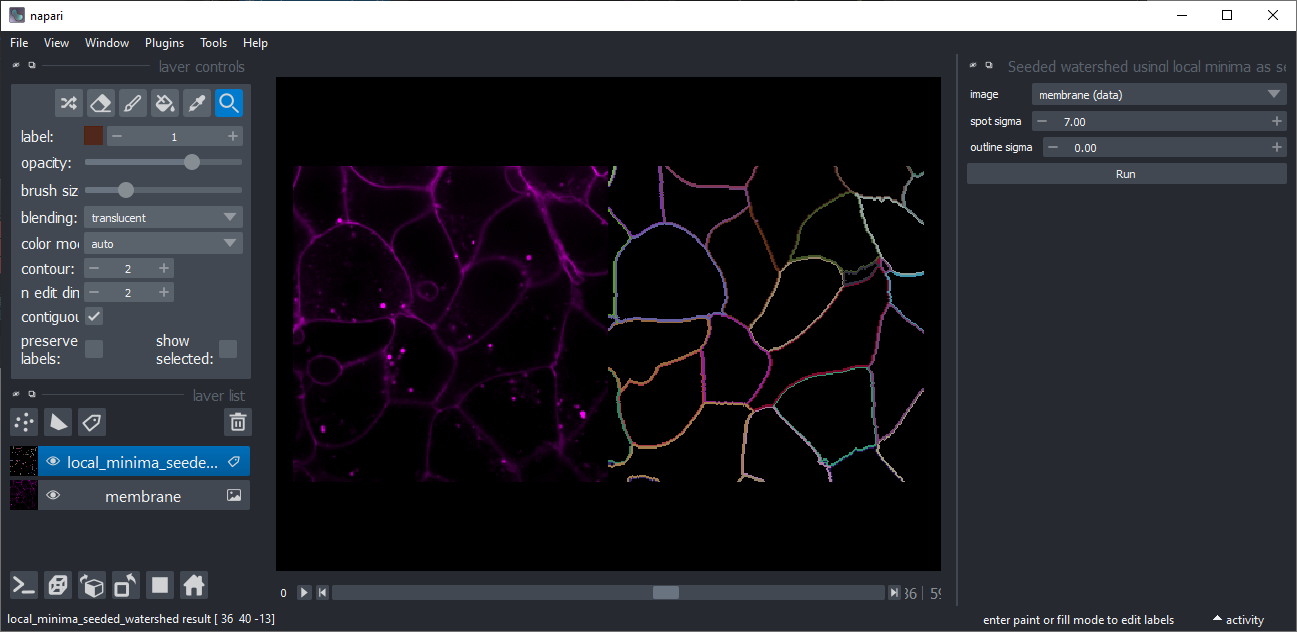
Seeded Watershed using local minima as starting points
Similar to the Seeded Watershed and Voronoi-Otsu-Labeling explained above, you can use this tool to segment an image showing membranes without an additional image showing nuclei. The two sigma parameters allow to fine tune how close objects can be and how precise their boundaries are detected.
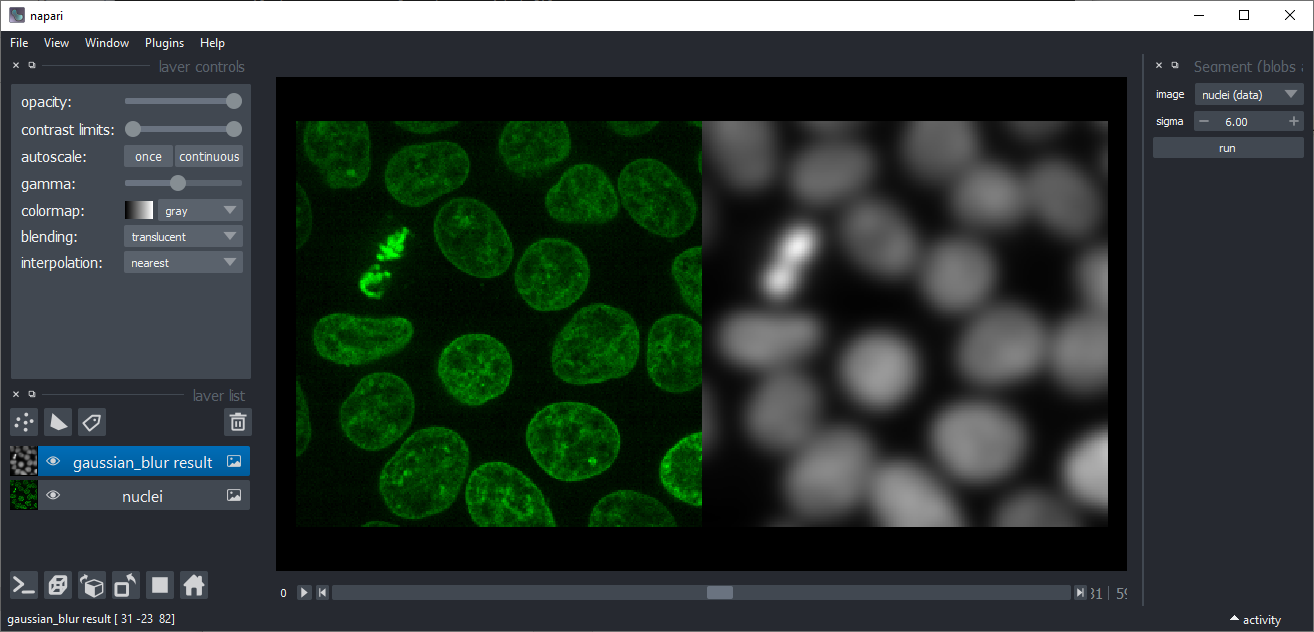
Gaussian blur
Applies a Gaussian blur to an image. This might be useful for denoising, e.g. before applying the Threshold-Otsu method.
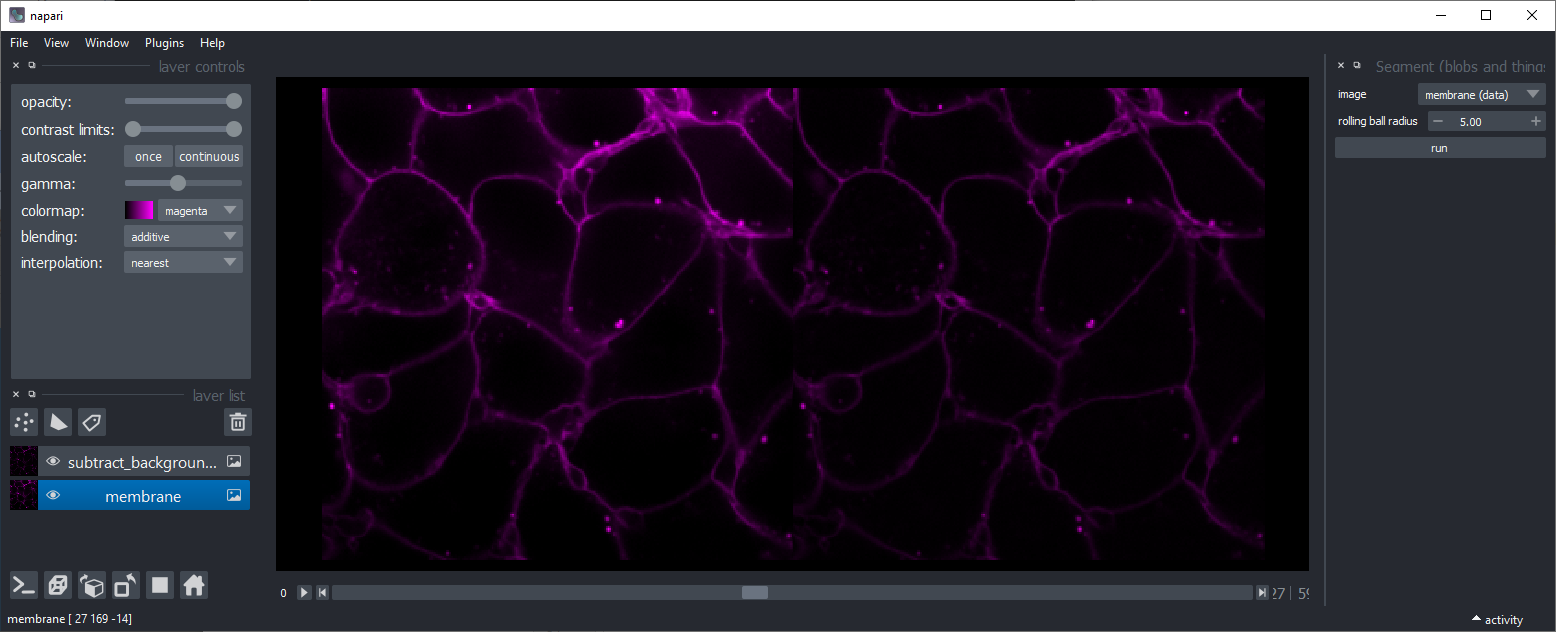
Subtract background
Subtracts background using scikit-image's rolling-ball algorithm. This might be useful, for example to make intensity of membranes more similar in different regions of an image.
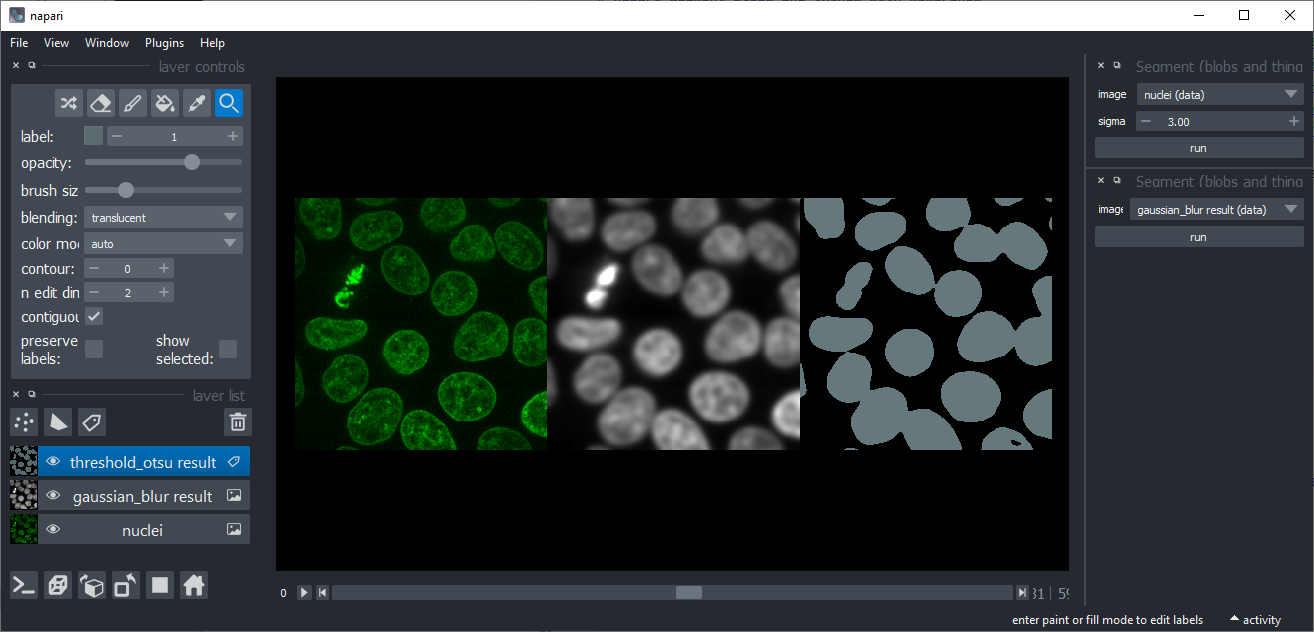
Threshold Otsu
Binarizes an image using scikit-image's threshold Otsu algorithm, also known as Otsu's method.
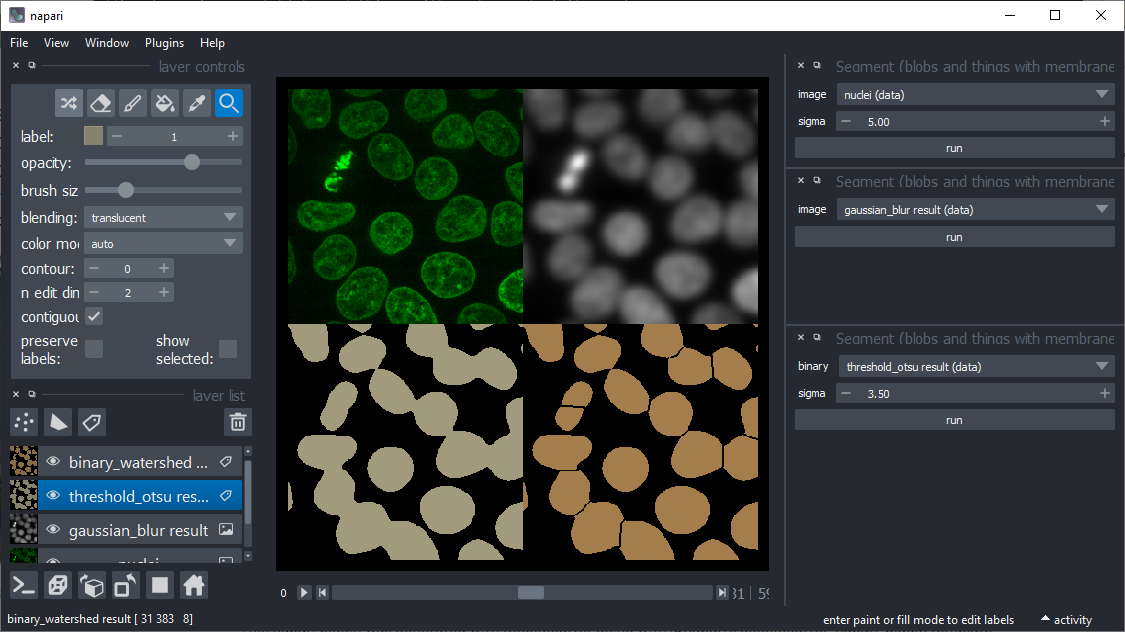
Split touching objects (formerly known as binary watershed).
In case objects stick together after thresholding, this tool might help. It aims to deliver similar results as ImageJ's watershed implementation.
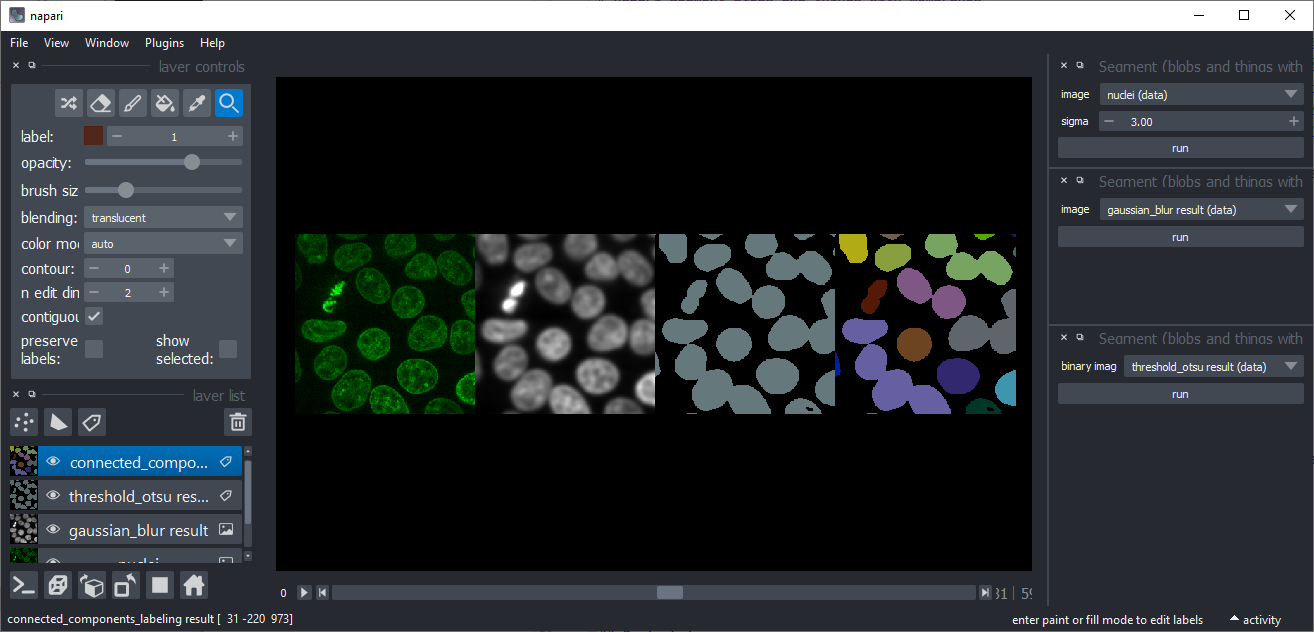
Connected component labeling
Takes a binary image and produces a label image with all separated objects labeled differently. Under the hood, it uses scikit-image's label function.
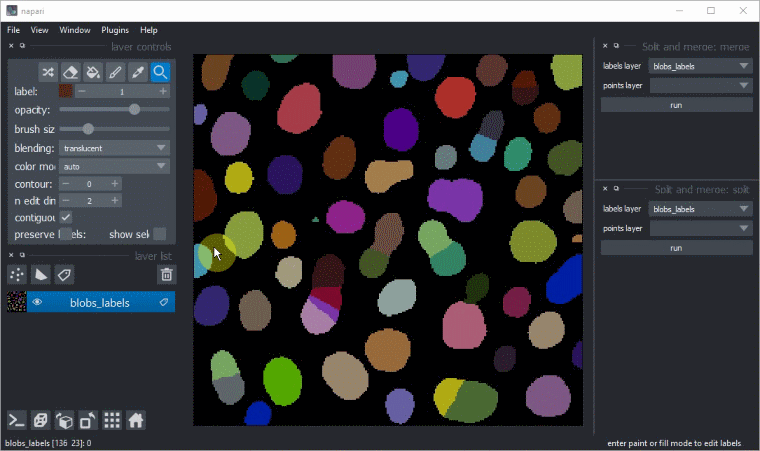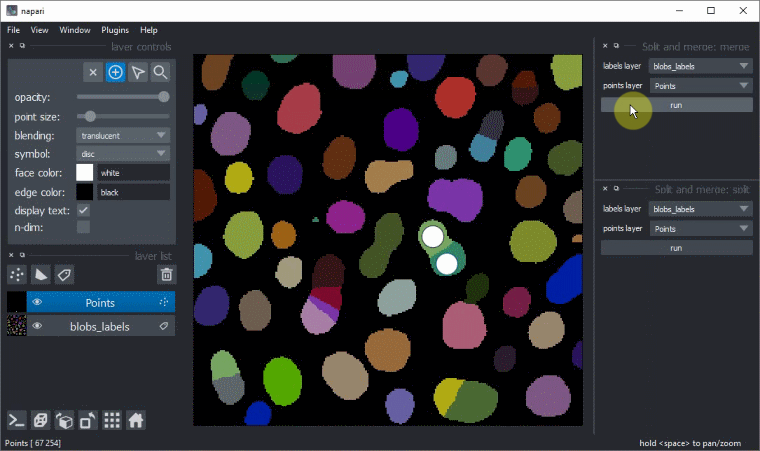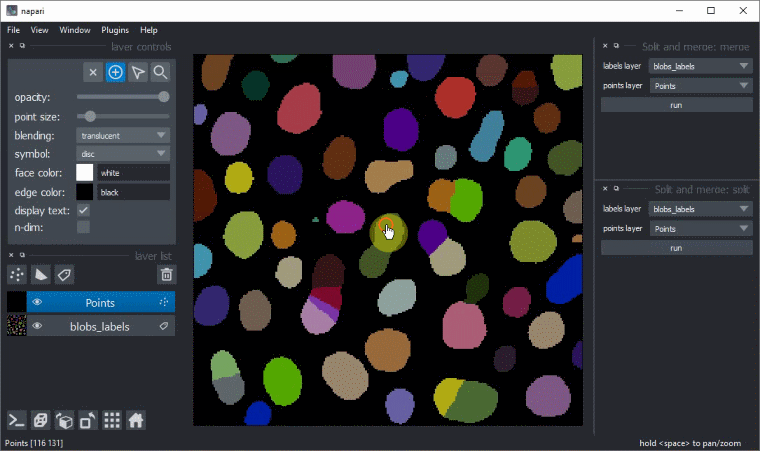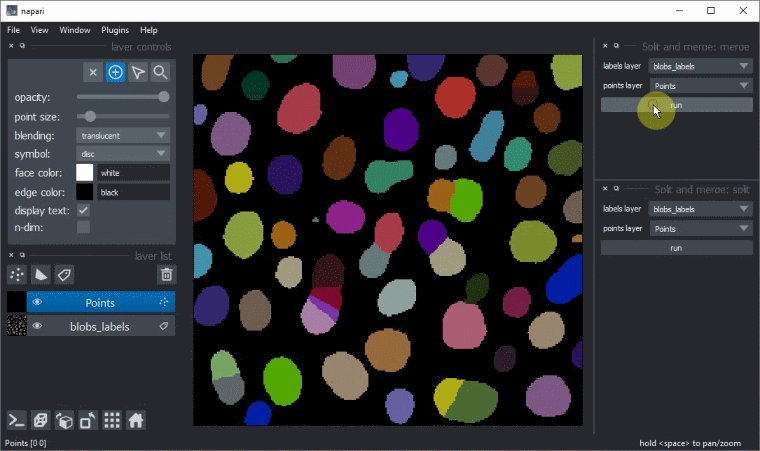
Manual split and merge labels
Split and merge labels in napari manually via the Tools > Utilities menu:
This napari plugin was generated with Cookiecutter using with @napari's cookiecutter-napari-plugin template.
Installation
This plugin is part of devbio-napari. To install it, please follow its installation instructions.
Contributing
Contributions are very welcome. Tests can be run with tox, please ensure the coverage at least stays the same before you submit a pull request.
License
Distributed under the terms of the BSD-3 license, "napari-segment-blobs-and-things-with-membranes" is free and open source software
Issues
If you encounter any problems, please create a thread on image.sc along with a detailed description and tag @haesleinhuepf.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_segment_blobs_and_things_with_membranes-0.3.12.tar.gz.
File metadata
- Download URL: napari_segment_blobs_and_things_with_membranes-0.3.12.tar.gz
- Upload date:
- Size: 17.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4514c6162bd85b98c8850ab83e75295bcda789b27fe322d0a07744e568984259
|
|
| MD5 |
09d8d72477c706ca3d546cc2ba38a821
|
|
| BLAKE2b-256 |
5847e199562d4c03efc5c639fcb85e0020b62f768b1c78e42cb7e314986adc68
|
File details
Details for the file napari_segment_blobs_and_things_with_membranes-0.3.12-py3-none-any.whl.
File metadata
- Download URL: napari_segment_blobs_and_things_with_membranes-0.3.12-py3-none-any.whl
- Upload date:
- Size: 16.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a94e48cf95ef20ddd9268aca01d020af8ba531ba9896cc16731a4702ba775819
|
|
| MD5 |
99a9aaa52dbd815981aff609644b84a8
|
|
| BLAKE2b-256 |
86ce6295b128abec209d003295cb56dbf99a8d0072c107866124ca5ffd466bf6
|