A plugin for batch processing of confocal and whole-slide microscopy images of biological tissues
Project description
napari-tmidas
This napari plugin consists of a growing collection of pipelines for fast batch processing of confocal and whole slide microscopy images of biological tissues. This is a WIP and based on the T-MIDAS terminal.
Features
Currently, napari-tmidas provides pipelines as widgets for batch image conversion and processing, object cropping, label image inspection and ROI colocalization (cf. usage below). You can request new batch image processing features in issues.
Installation
(Video installation guides: https://www.youtube.com/@macromeer/videos)
First, install Napari in a virtual environment:
mamba create -y -n napari-tmidas -c conda-forge python=3.11
mamba activate napari-tmidas
python -m pip install "napari[all]"
Now you can install napari-tmidas via pip:
pip install napari-tmidas
For deep learning features (Batch Crop Anything with SAM2, Spotiflow, Careamics, Trackastra), also install:
pip install 'napari-tmidas[deep-learning]'
Or install everything at once:
pip install 'napari-tmidas[all]'
It is recommended though to install the latest development version. Please also execute this command from time to time in the activated environment to benefit from newly added features:
pip install git+https://github.com/MercaderLabAnatomy/napari-tmidas.git
Additional Setup for Batch Crop Anything
To use the Batch Crop Anything pipeline with SAM2, you only need to install ffmpeg manually. SAM2 and its model checkpoints will be automatically installed when you first run the pipeline:
mamba install -c conda-forge ffmpeg
If you want to batch compress image data using Zstandard, use the package manager of your operating system to install it:
sudo apt-get install zstd # Pre-installed on Linux :man_shrugging:
brew install zstd # for macOS (requires Homebrew)
pip install zstandard # Windows with Python >= 3.7
And you are done!
Usage
To use the plugin, start napari in the activated virtual environment with this terminal command:
mamba run -n napari-tmidas napari
You can then find the installed plugin in the Plugins tab.
Microscopy Image Conversion
Converts .lif, .nd2, .czi, .ndpi and Acquifer data to TIF or OME-Zarr formats. Scan a folder, select files, and export with preserved spatial metadata.
Supported Formats:
- TIF - Standard format for compatibility
- OME-Zarr - Recommended for large datasets, spec v0.5 compliant with automatic physical metadata extraction (voxel sizes, spacing)
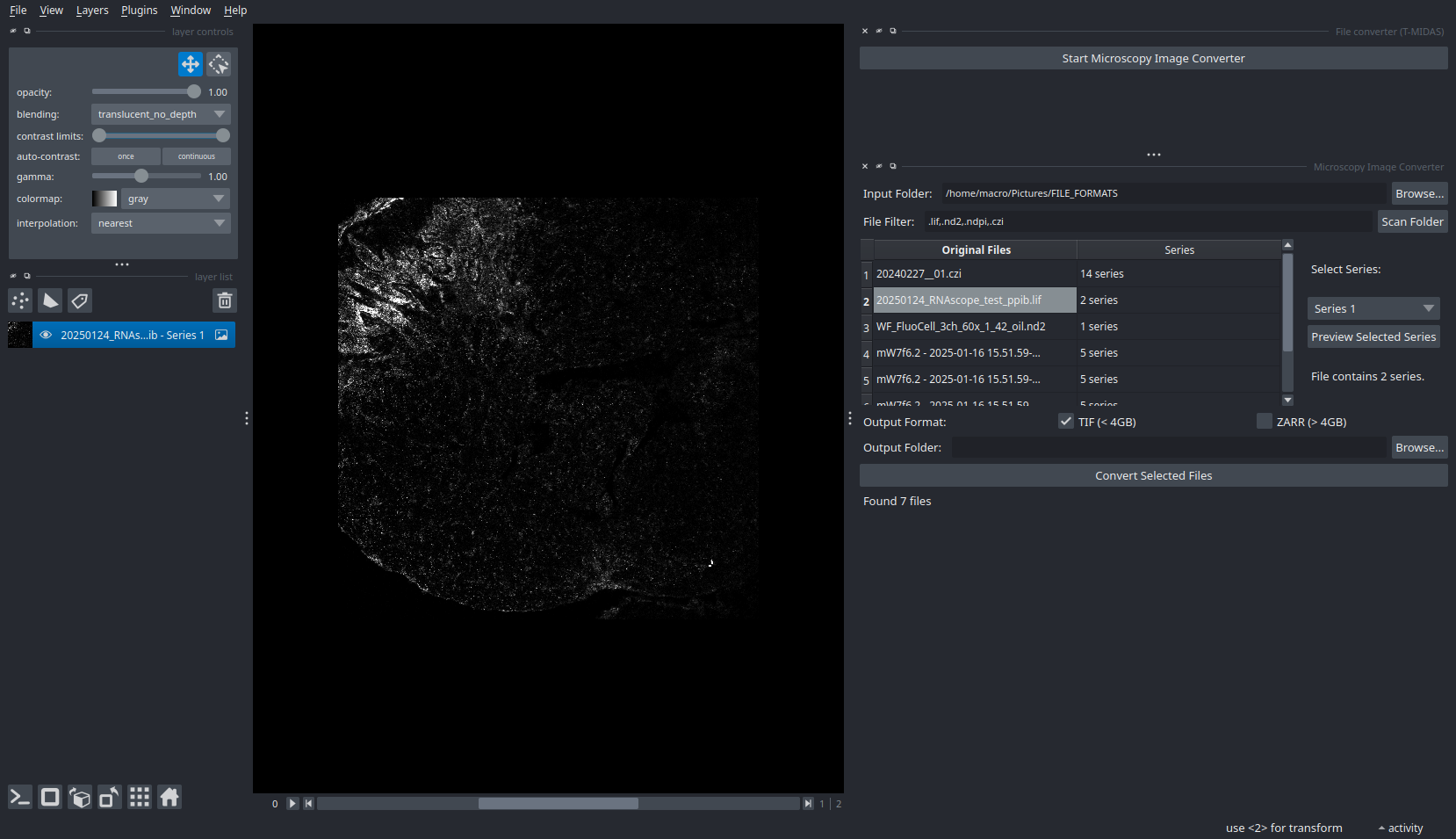
Image Processing
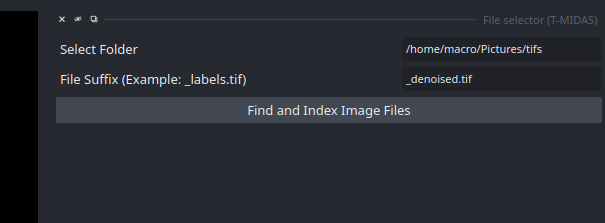
- You start with entering the path to the folder containing the images to be processed (currently supports TIF, later also ZARR) and optionally a filter for filename suffix
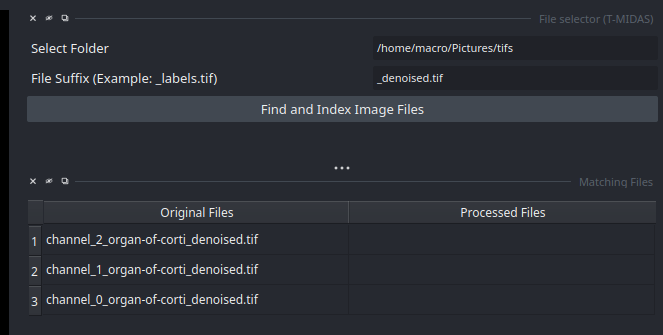
- After indexing the files, a table appears with the found images. You can click on them to inspect them in the viewer.
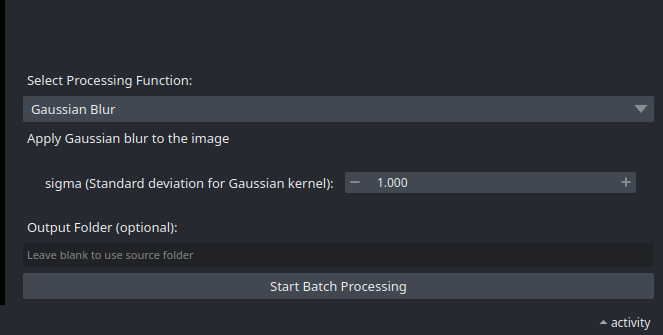
- Next, select a processing function, set parameters if applicable and
Start Batch Processing.
- You can click on the images in the table to show them in the viewer. For example first click on one of the
Original Files, and then the correspondingProcessed Fileto see an overlay.
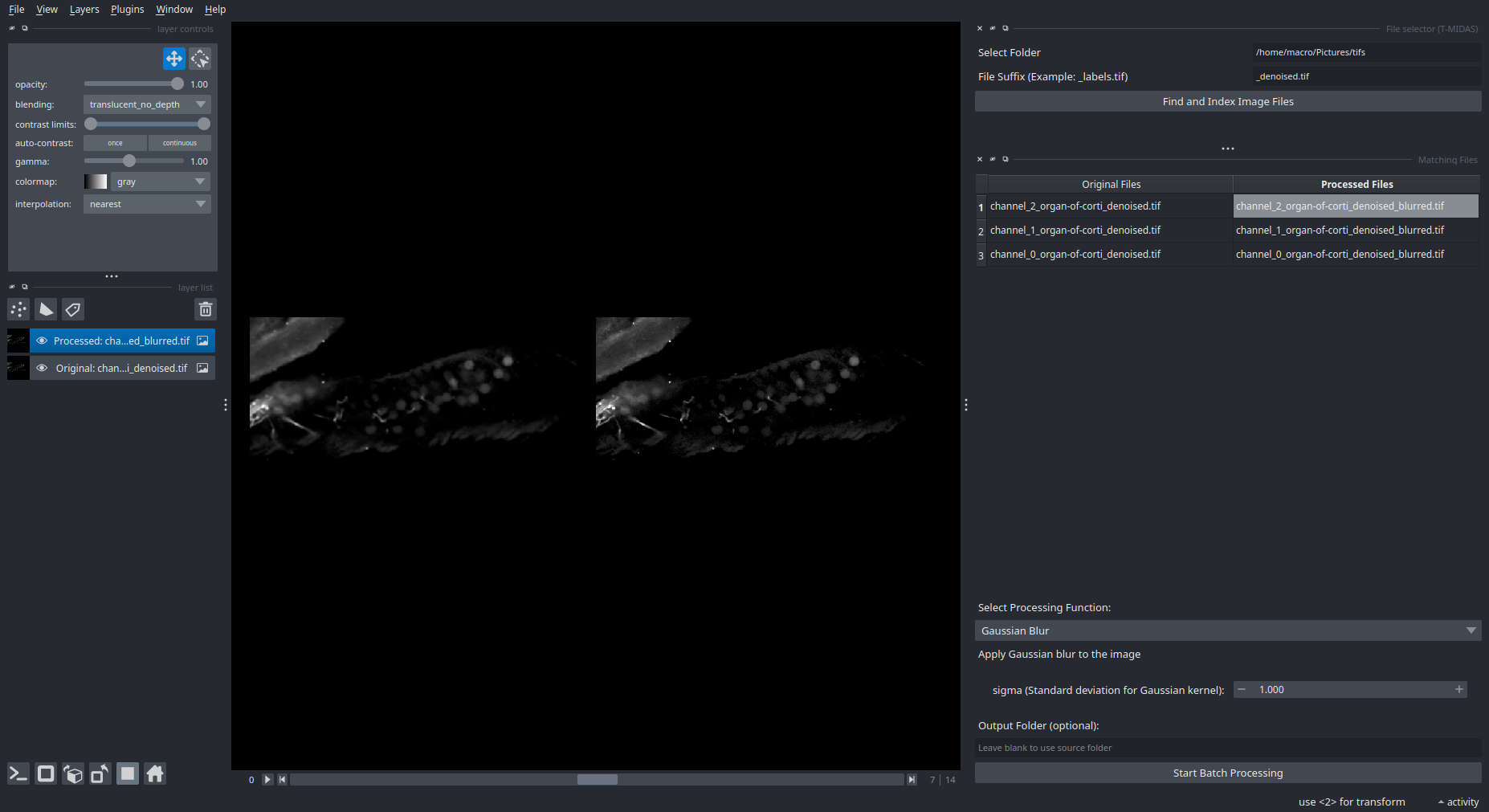
Note that whenever you click on an Original File or Processed File in the table, it will replace the one that is currently shown in the viewer. So naturally, you'd first select the original image, and then the processed image to correctly see the image pair that you want to inspect.
Processing Function Credits
The image processing capabilities are powered by several excellent open-source tools:
- Cellpose 4: Advanced cell segmentation
- Trackastra: Cell tracking and analysis
- VisCy: Virtual staining using deep learning
- CAREamics: Content-aware image restoration and enhancement
- Spotiflow: Accurate and efficient spot detection for fluorescence microscopy
Processing Function Documentation
Detailed documentation for specific processing functions:
Core Processing
- Basic Processing Functions - Label and intensity operations, channel splitting/merging, time series
- Cellpose Segmentation - Deep learning cell/nucleus segmentation
- TrackAstra Tracking - Cell tracking across time-lapse data
- VisCy Virtual Staining - Virtual staining of phase/DIC images using deep learning
Analysis and Quality Control
- Grid View: Intensity + Labels Overlay - Visual QC for segmentation results
- Intensity-Based Label Filtering - Filter labels by signal intensity
- Regionprops Analysis - Extract quantitative properties from labels
Advanced Processing
- Advanced Processing Functions - Denoising (CAREamics), spot detection (Spotiflow), SciPy/scikit-image filters, compression, colocalization
Batch Label Inspection
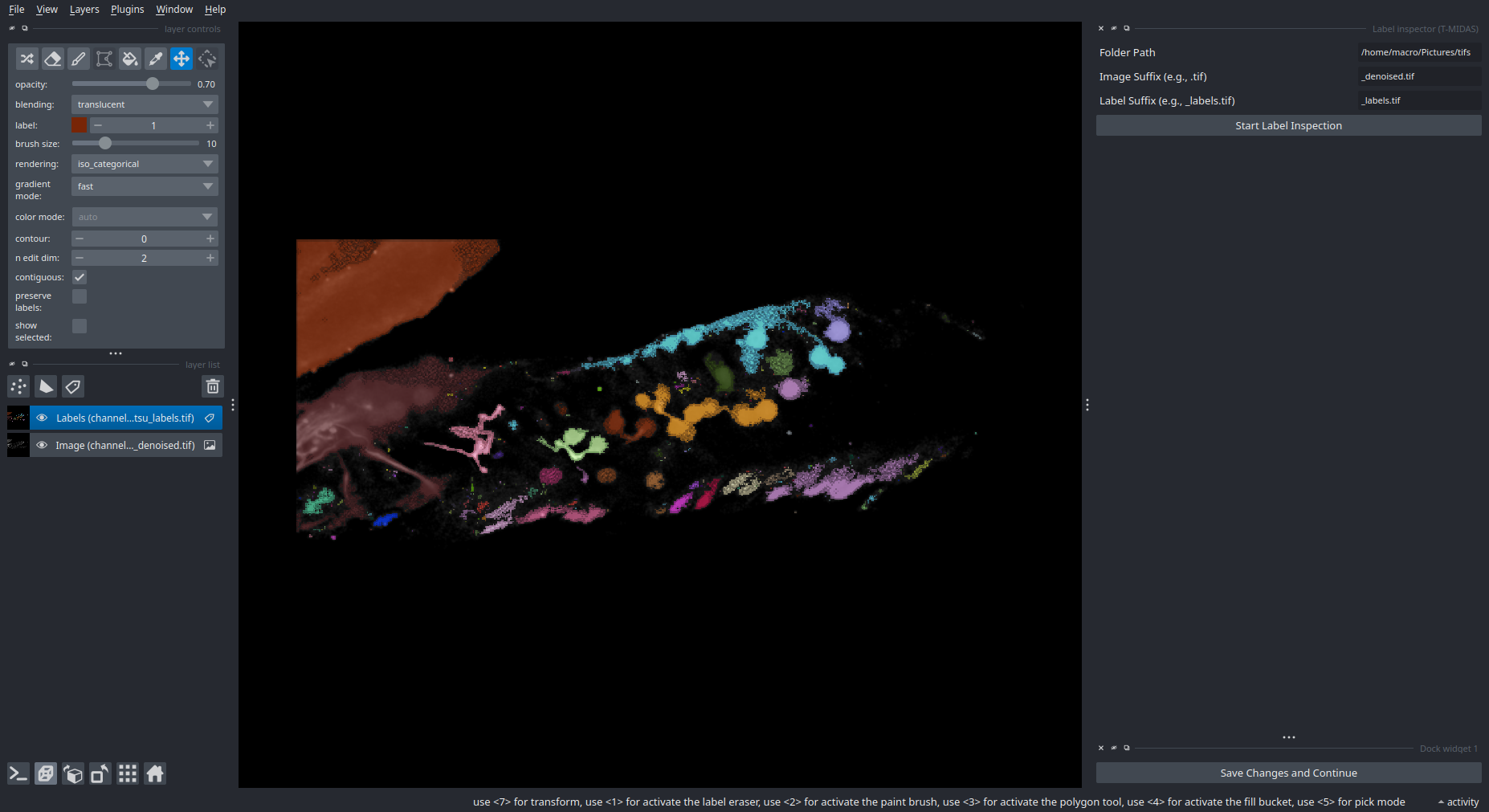
If you have already segmented a folder full of images and now you want to maybe inspect and edit each label image, you can use the Plugins > T-MIDAS > Batch Label Inspection, which automatically saves your changes to the existing label image once you click the Save Changes and Continue button (bottom right).
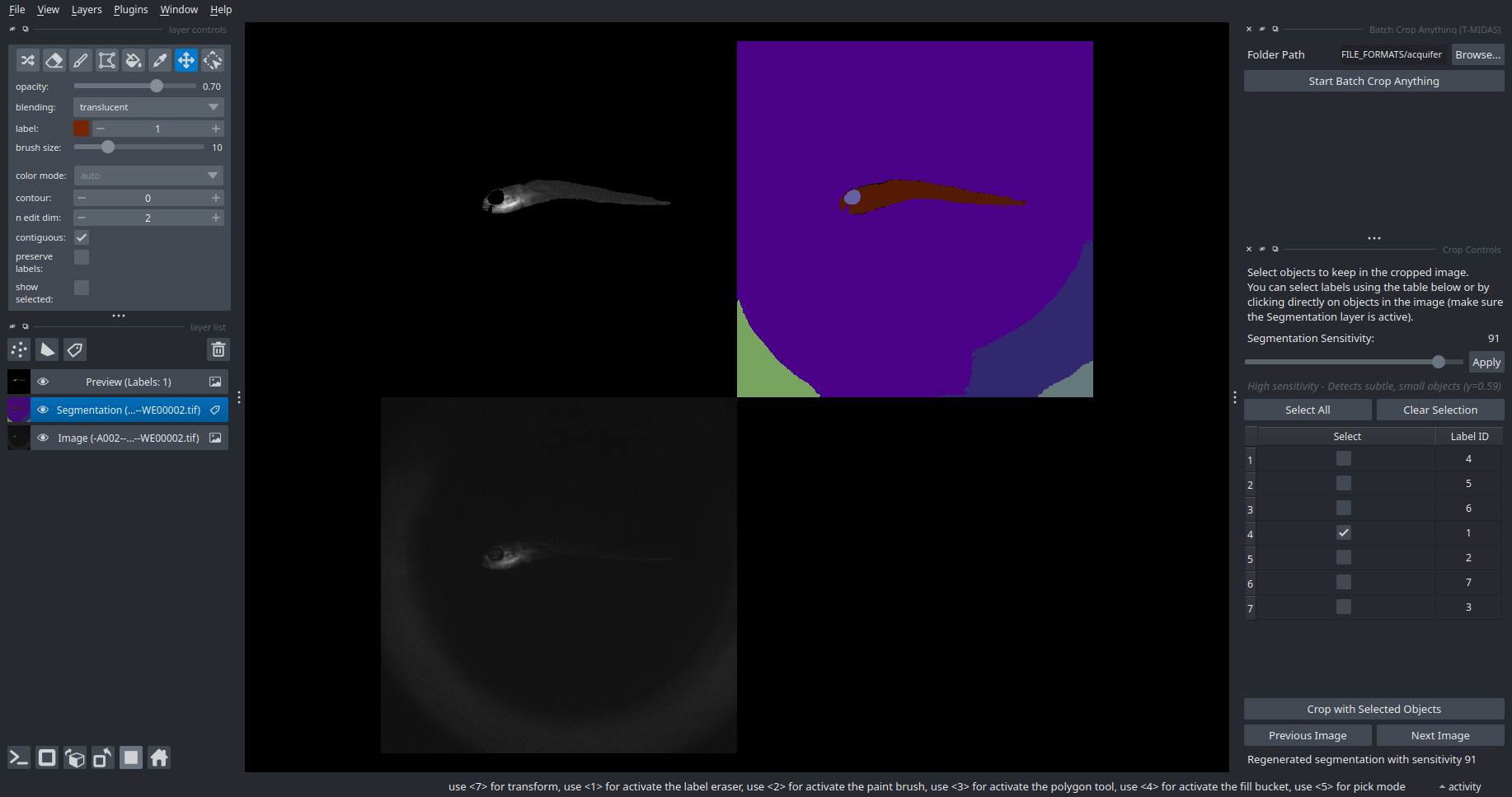
Crop Anything
This pipeline combines the Segment Anything Model (SAM2; supports YX, ZYX and TYX data) for automatic object detection with an interactive interface for selecting and cropping multiple objects from images. To launch the widget, open Plugins > T-MIDAS > Batch Crop Anything. Cropping works like this: Enter 2D view and go to the first z slice where the object to be cropped is appearing. Activate/select the points layer and click on the object. Terminal shows progress. You can then proceed to select another object (always do this in 2D mode)
ROI Colocalization
This pipeline quantifies colocalization between labeled regions of interest (ROIs) across multiple image channels. It determines the extent of overlap between ROIs in a reference channel and those in one or two other channels. The output is a table of colocalization counts. Optionally, the size of reference channel ROIs, as well as the total or median size of colocalizing ROIs in the other channels, can be included. Colocalization is determined using Boolean masking. The number of colocalizing instances is determined by counting unique label IDs within the overlapping regions. Typically, the reference channel contains larger structures, while other channels contain smaller, potentially nested, structures. For example, the reference channel might contain cell bodies, with the second and third channels containing nuclei and sub-nuclear objects, respectively.

Contributing
Contributions are very welcome. Tests can be run with tox, please ensure the coverage at least stays the same before you submit a pull request.
License
Distributed under the terms of the BSD-3 license, "napari-tmidas" is free and open source software
Issues
If you encounter any problems, please file an issue along with a detailed description.
This napari plugin was generated with copier using the napari-plugin-template.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_tmidas-0.2.6.tar.gz.
File metadata
- Download URL: napari_tmidas-0.2.6.tar.gz
- Upload date:
- Size: 232.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
24c14e33bfbdaeef9e6ff995ac2e3aad2b574d5269d3360387627d197f15279f
|
|
| MD5 |
98762d62cfefb2166468435e5b3a60d9
|
|
| BLAKE2b-256 |
69b6d319d808213b9c91073e6301a257c26d528099aa9546e075bfb030263622
|
File details
Details for the file napari_tmidas-0.2.6-py3-none-any.whl.
File metadata
- Download URL: napari_tmidas-0.2.6-py3-none-any.whl
- Upload date:
- Size: 227.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
56087fd4e043e17f9078a4786287de300bf274a726e19f80931621048ebbf5ee
|
|
| MD5 |
924860fa843677744b5bf7f51654ff30
|
|
| BLAKE2b-256 |
b02e2465d6e379be336c8883f749282e89ee17142ce2d76ebc3065f17eeb995a
|