A plugin for batch processing of confocal and whole-slide microscopy images of biological tissues
Project description
napari-tmidas
AI-powered batch processing for microscopy images
Transform, analyze, and quantify microscopy data at scale with deep learning - from file conversion to segmentation, tracking, and analysis.
✨ Key Features
🤖 5 AI Methods Built-In
- Virtual staining (VisCy) • Denoising (CAREamics) • Spot detection (Spotiflow) • Segmentation (Cellpose) • Tracking (Trackastra)
- Auto-install in isolated environments • No dependency conflicts • GPU acceleration
🔄 Universal File Conversion
- Convert LIF, ND2, CZI, NDPI, Acquifer → TIFF or OME-Zarr
- Preserve spatial metadata automatically
⚡ Batch Processing
- Process entire folders with one click • 40+ processing functions • Progress tracking & quality control
📊 Complete Analysis Pipeline
- Segmentation → Tracking → Quantification → Colocalization
🚀 Quick Start
# Install napari and the plugin
mamba create -y -n napari-tmidas -c conda-forge python=3.11
mamba activate napari-tmidas
pip install "napari[all]"
pip install napari-tmidas
# Launch napari
napari
Then find napari-tmidas in the Plugins menu. Watch video tutorials →
💡 Tip: AI methods auto-install their dependencies on first use - no manual setup required!
📖 Documentation
AI-Powered Methods
| Method | Description | Documentation |
|---|---|---|
| 🎨 VisCy | Virtual staining from phase/DIC | Guide |
| 🔧 CAREamics | Noise2Void/CARE denoising | Guide |
| 🎯 Spotiflow | Spot/puncta detection | Guide |
| 🔬 Cellpose | Cell/nucleus segmentation | Guide |
| 📈 Trackastra | Cell tracking over time | Guide |
Core Workflows
- Image Conversion - Multi-format microscopy file conversion
- Batch Processing - Label operations, filters, channel splitting
- Quality Control - Visual QC with grid overlay
- Quantification - Extract measurements from labels
- Colocalization - Multi-channel ROI analysis
Advanced Features
- SAM2 Crop Anything - Interactive object cropping
- Advanced Filters - SciPy/scikit-image filters
- Batch Label Inspection - Manual correction workflow
💻 Installation Options
Recommended (latest features):
pip install git+https://github.com/MercaderLabAnatomy/napari-tmidas.git
Stable release:
pip install napari-tmidas
With deep learning (optional):
pip install 'napari-tmidas[deep-learning]' # Includes SAM2
pip install 'napari-tmidas[all]' # Everything
Additional setup for SAM2:
mamba install -c conda-forge ffmpeg # Required for video processing
🖼️ Screenshots
File Conversion Widget
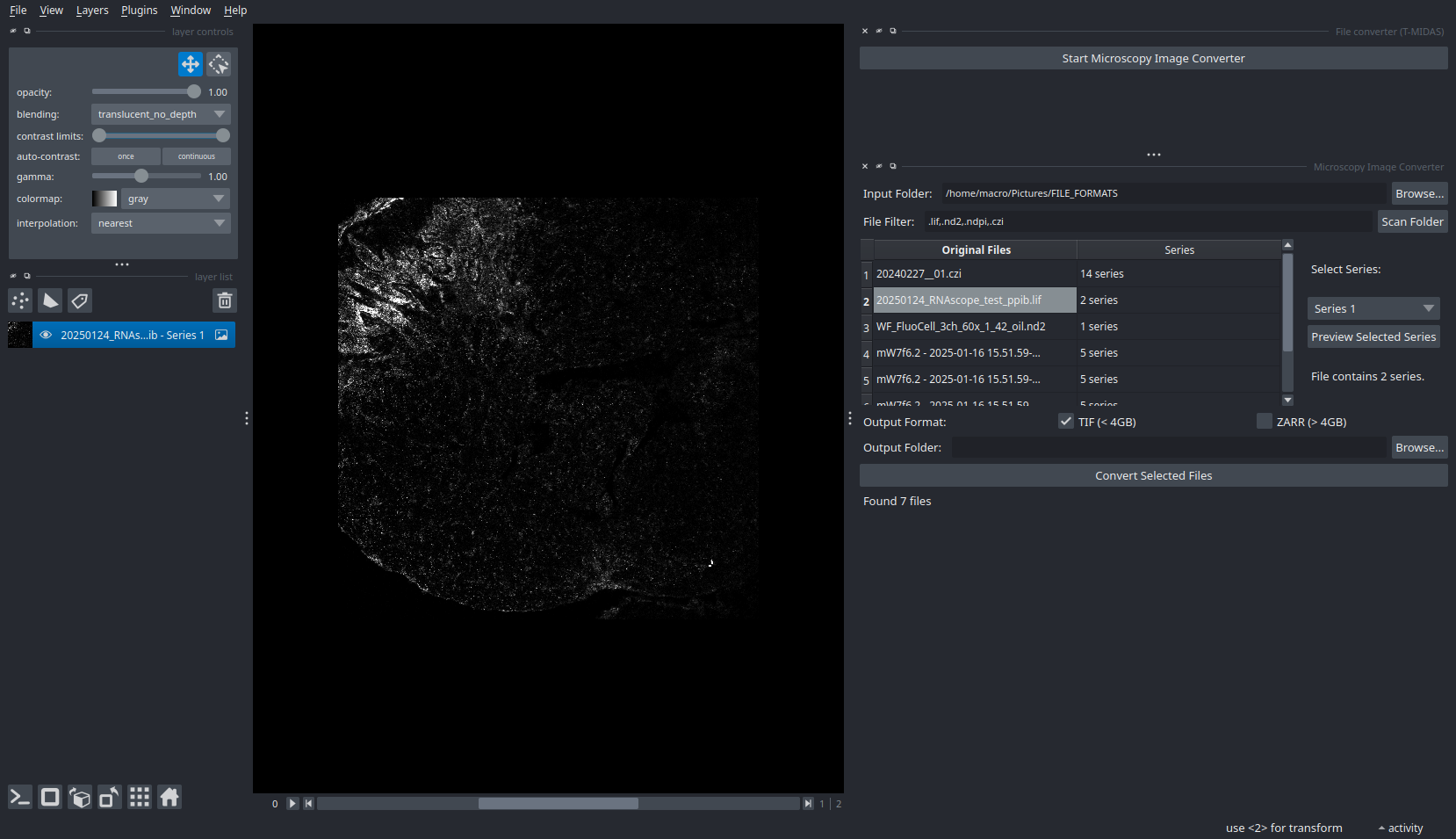
Convert proprietary formats to open standards with metadata preservation.
Batch Processing Interface
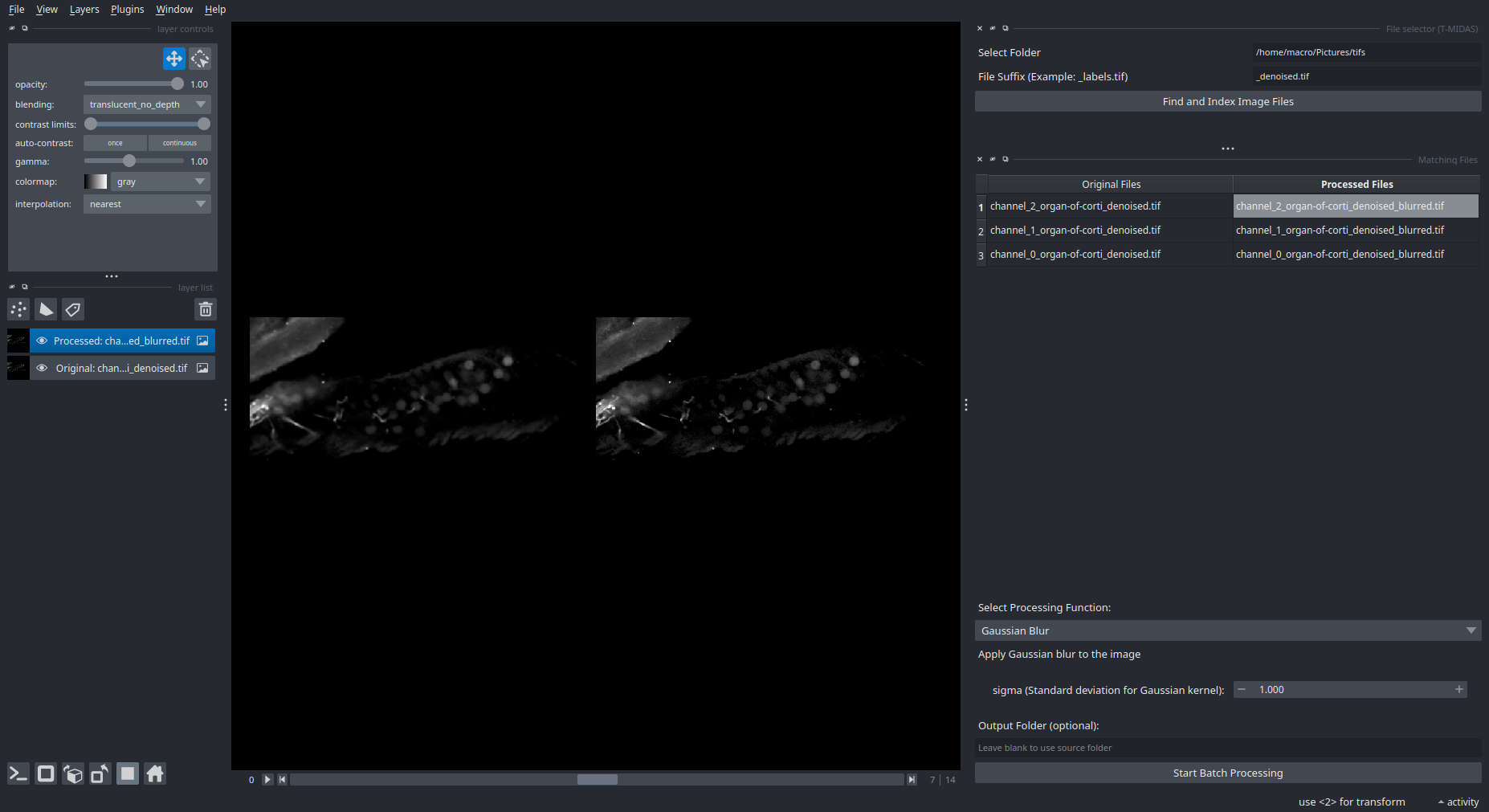
Select files → Choose processing function → Run on entire dataset.
Label Inspection
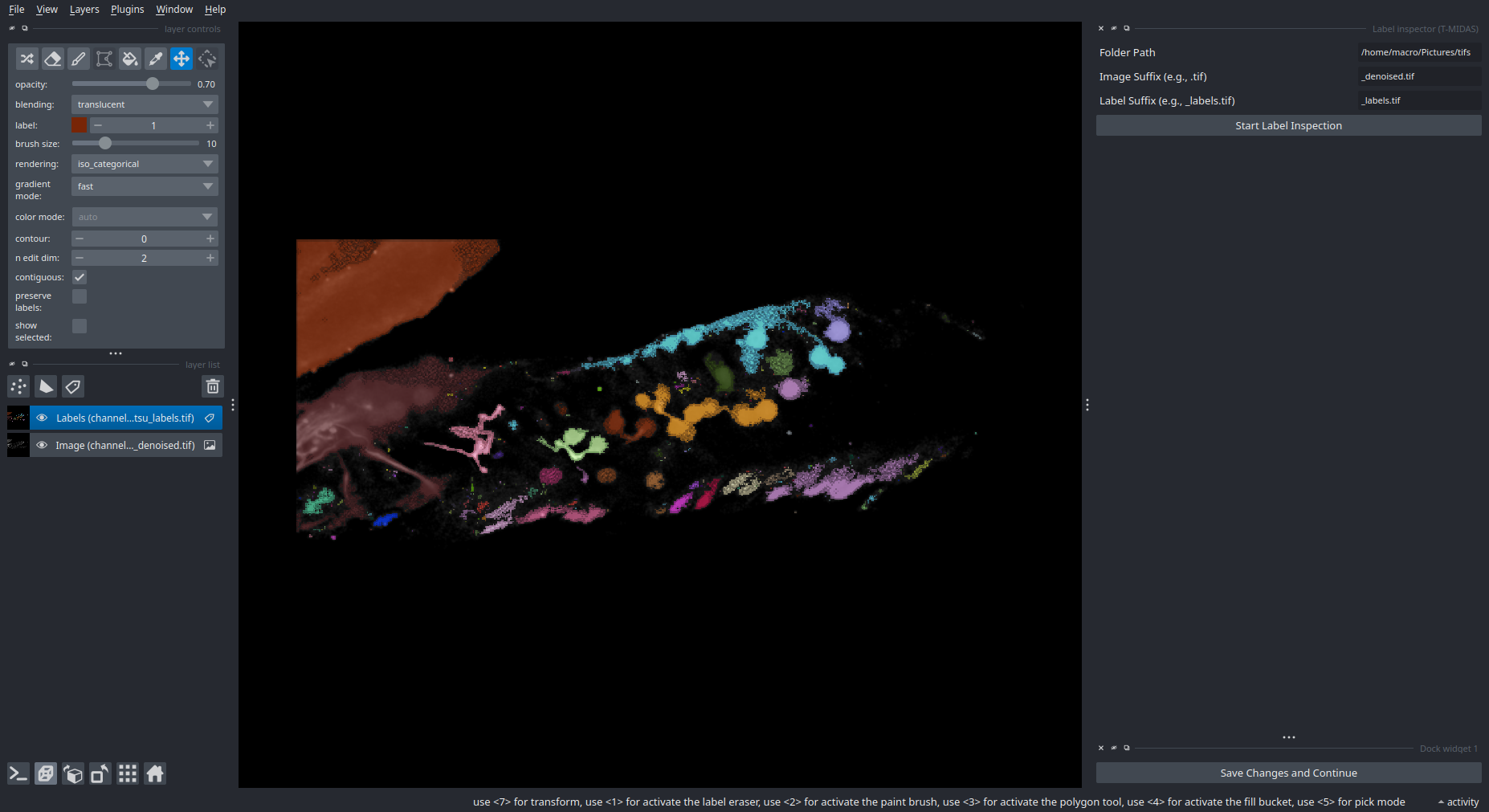
Inspect and manually correct segmentation results.
SAM2 Crop Anything
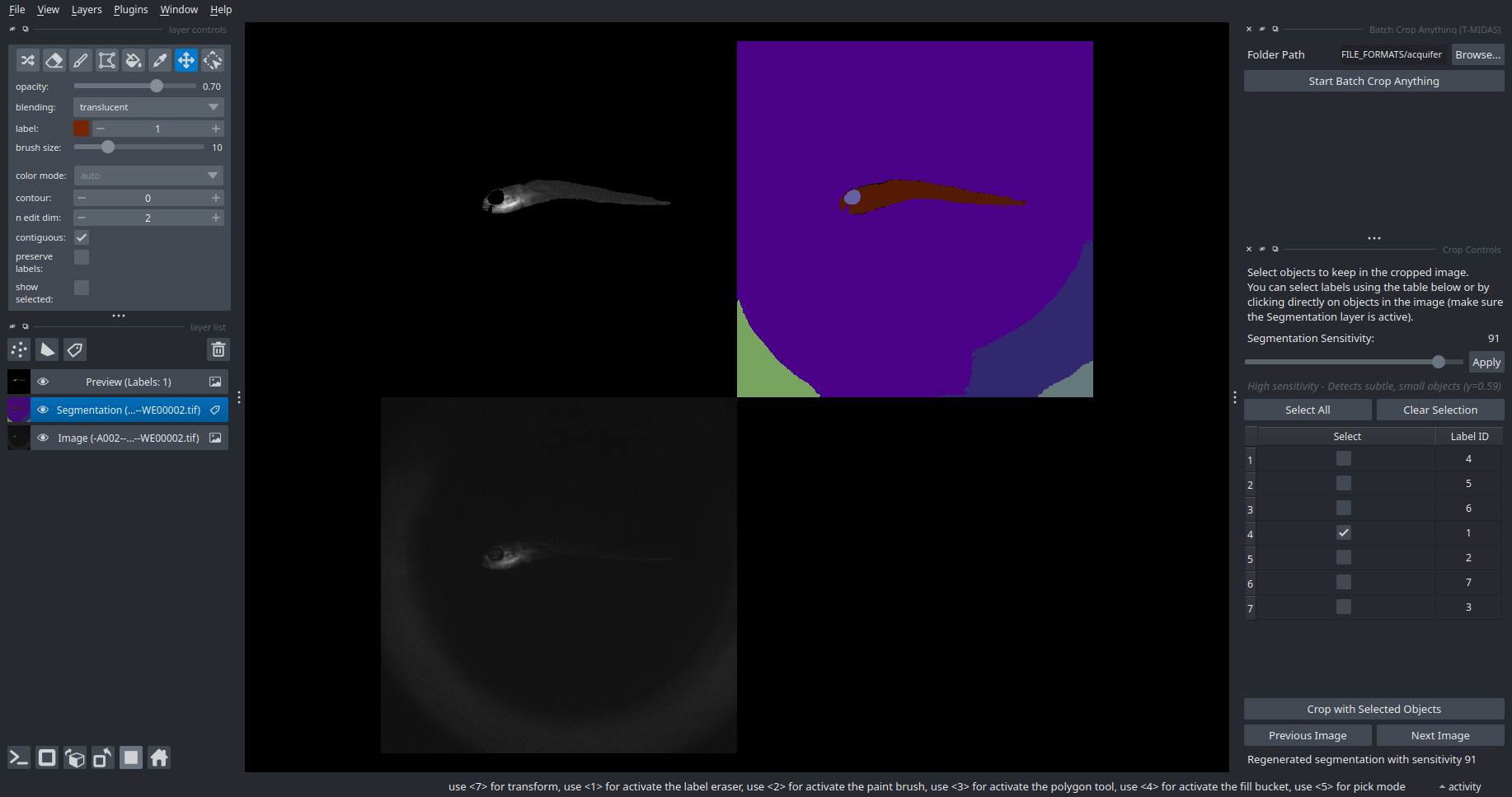
Interactive object selection and cropping with SAM2.
🤝 Contributing
Contributions are welcome! Please ensure tests pass before submitting PRs:
pip install tox
tox
📄 License
BSD-3 License - see LICENSE for details.
🐛 Issues
Found a bug or have a feature request? Open an issue
🙏 Acknowledgments
Built with napari and powered by:
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file napari_tmidas-0.3.0.tar.gz.
File metadata
- Download URL: napari_tmidas-0.3.0.tar.gz
- Upload date:
- Size: 247.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8e83b436b3c7d8cd649b6c4ea58123eac2764a53fd2e6654a2bbfb33069876d1
|
|
| MD5 |
d286067bc0ca7eb16bfb1b8c3147faf6
|
|
| BLAKE2b-256 |
a6cb1449de7969d56ed49f8b65441c4d208ed8d3b80421b6dc4a9fa12969e2d0
|
File details
Details for the file napari_tmidas-0.3.0-py3-none-any.whl.
File metadata
- Download URL: napari_tmidas-0.3.0-py3-none-any.whl
- Upload date:
- Size: 235.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dce260d5d1864a18ac8f51fe4156416bd584a5584fde12f493139d70d7a84a4d
|
|
| MD5 |
ddc5d1c4d4413b3635a6daabb08a7d1e
|
|
| BLAKE2b-256 |
8b0f2be192aa5eacd53061181394e4584072079c309734e3391c1480878e231e
|

















