No project description provided
Project description
nnx
Neural Networks for JAX
nnx is a Neural Networks library for JAX that uses Pytree-based Modules and a novel
a Reference system to enable:
- Simplicity: Provides an easy-to-understand mental model and implementation.
- Shared refereces: Supports safe mutable shared state thanks to its Reference system.
- Leaf Metadata: Enables semantic splitting a Module's state (similar to Flax collections) and adding Axis Metadata.
- Stateful transformations: Seamless integration with JAX's native transformation capabilities.
NNX was designed to have the same capabilities as Flax with the simplicity of Equinox.
Status
nnx is currently a proof of concept, aimed at exploring the design space of a lightweight module
system for JAX based on references. It is not intended for production use.
Installation
To get started with nnx, first install the package using pip:
pip install nnx
To get the latest version, install directly from GitHub:
pip install git+https://github.com/cgarciae/nnx
Usage
import nnx
import jax
class Linear(nnx.Module):
w: jax.Array = nnx.param()
b: jax.Array = nnx.param()
def __init__(self, din: int, dout: int, *, ctx: nnx.Context):
key = ctx.make_rng("params")
self.w = jax.random.uniform(key, (din, dout))
self.b = jax.numpy.zeros((dout,))
def __call__(self, x):
return x @ self.w + self.b
ctx = nnx.Context(params=jax.random.PRNGKey(0))
model = Linear(din=12, dout=2, ctx=ctx)
@nnx.jit
def train_step(model, x, y):
def loss_fn(model):
y_pred = model(x)
return jax.numpy.mean((y_pred - y) ** 2)
# compute gradient
grads = nnx.grad(loss_fn, wrt="params")(model)
# SGD update
model[:] = jax.tree_map(lambda w, g: w - 0.1 * g, model["params"], grads)
# yes... there's no return :)
train_step(model, x, y)
Design
Modules
nnx has a simple and intuitive design with Modules at its core. These modules are built using simple_pytree Pytrees. To create a custom module, simply subclass nnx.Module and mark the reference fields with nnx.ref or nnx.param. These are descriptors that store a Ref instance in a separate attribute, making references transparent to the user.
Here's an example of a simple Linear module:
import nnx
import jax
class Linear(nnx.Module):
w: jax.Array = nnx.ref("params")
b: jax.Array = nnx.param() # shortcut for ref("params")
def __init__(self, din: int, dout: int, *, ctx: nnx.Context):
key = ctx.make_rng("params") # request an RNG key
self.w = jax.random.uniform(key, (din, dout))
self.b = jax.numpy.zeros((dout,))
def __call__(self, x):
return x @ self.w + self.b
ctx = nnx.Context(params=jax.random.PRNGKey(0))
model = Linear(din=12, dout=2, ctx=ctx)
nnx uses a Context object to propagate RNG and other forms of state throughout the model. Context provides a make_rng method that creates a new RNG key on demand for a given name (in this case, "params").
RefField Descriptor
nnx.ref and nnx.param are descriptors that create RefField instances. A RefField is a descriptor that stores a Ref instance in a separate {attribute_name}__ref attribute. It handles retrieving and setting the value of the reference automatically, so the user doesn't have to manipulate references directly. Additionally, RefField inherits from dataclasses.Field to ensure compatibility with nnx.dataclass when needed.
Here's a simplified version of the RefField implementation:
class RefField(dataclasses.Field):
def __set_name__(self, cls, name):
self.name = name
def __get__(self, obj, objtype=None):
ref = getattr(obj, f"{self.name}__ref")
return ref.value
def __set__(self, obj, value):
ref = getattr(obj, f"{self.name}__ref")
ref.value = value
It's important to note that Ref instances are created during the first call to __set__ if the companion {name}__ref attribute doesn't exist yet. This should only happen inside the __init__ method, otherwise, the Module will raise an error, as simple_pytree Pytrees are frozen after initialization.
GetItem and SetItem Syntactic Sugar
Module implements __getitem__ and __setitem__ to provide syntactic sugar for creating and updating Partitions. Although it may appear otherwise, __setitem__ does not modify the Module's structure. Instead, it updates the values of the references, as demonstrated in this simplified implementation:
class Module(nnx.Pytree):
...
def __getitem__(self, collection: str) -> nnx.Partition:
return nnx.get_partition(self, collection)
def __setitem__(self, _: SetItemType, value: nnx.Partition):
nnx.update_refs(self, value)
Sample usage might look like this:
# SGD update
model[:] = jax.tree_map(lambda w, g: w - 0.1 * g, model["params"], grads)
In this example, model["params"] returns a Partition that contains all the references in the params collection. grads is a Partition with the same structure as model["params"], but with gradients instead of parameters. The statement model[:] = ... updates the values of the references in model with the values of the new parameters from stochastic gradient descent (SGD) update rule.
Transformations
Currently, NNX offers three types of transformations: stateful, filtered, and the partition API. As it is unclear which API is the best, all three will be maintained for now.
Stateful Transforms
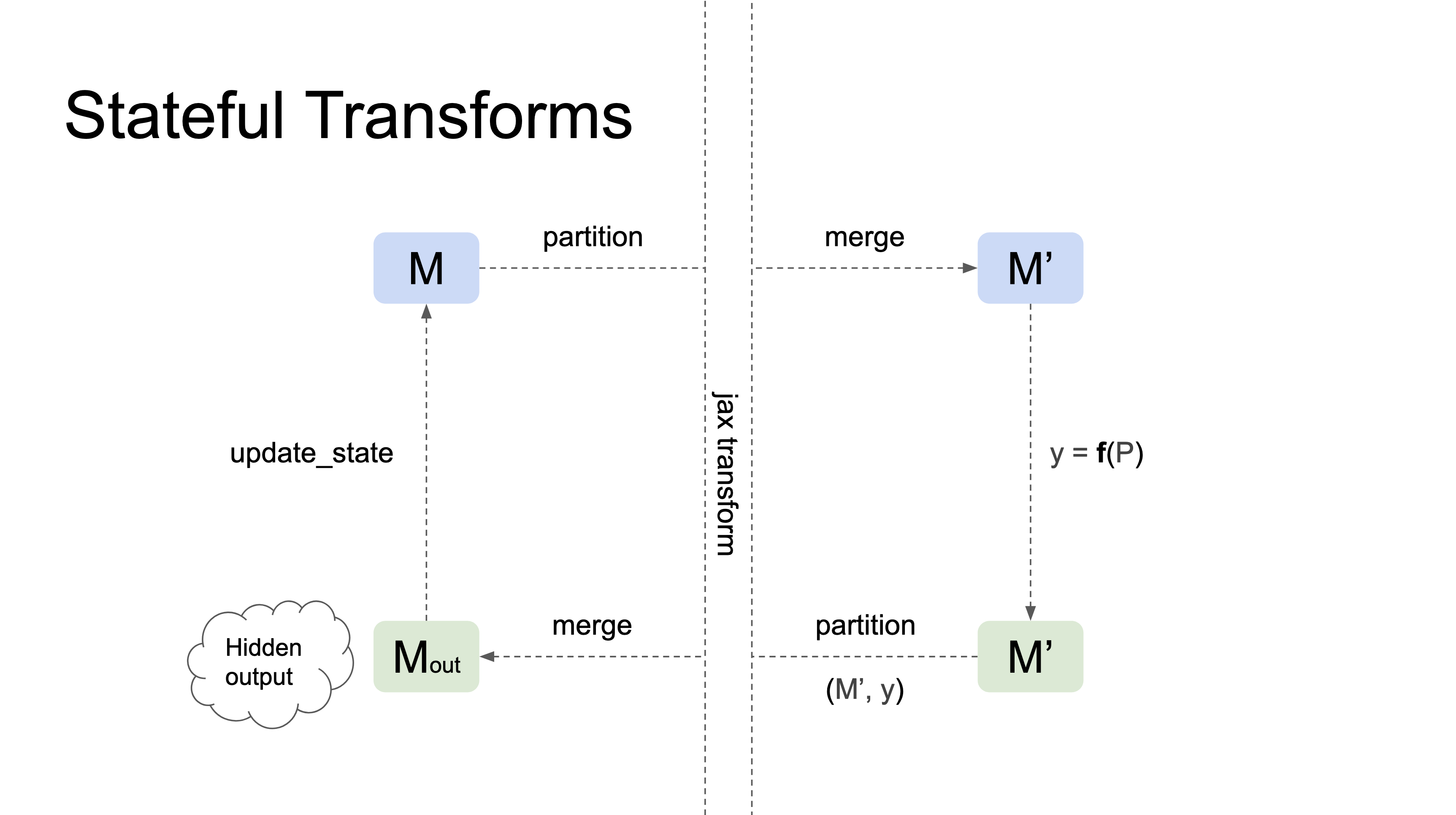
Stateful transforms take a Pytree of references (e.g., a Module) as their first argument, track changes in the state of the references that occur within the transformation, and automatically propagate those changes to the input Pytree outside the transformation. In general, they have the following properties:
- They behave as stateful functions with respect to the first argument.
- They can operate on collections and RNG streams according to the transformation's semantics, exactly like Flax's transformations.
- They handle all relevant global state, such as
nnx.scopeand Refx's trace state.
Here's a diagram illustrating how stateful transformations work:
Currently, nnx.jit and nnx.grad are the only stateful transformations.
The following example demonstrates how a train_step function can be implemented using nnx.jit and nnx.grad:
@nnx.jit
def train_step(model, x, y):
def loss_fn(model):
y_pred = model(x)
return jax.numpy.mean((y_pred - y) ** 2)
# compute gradient
grads = nnx.grad(loss_fn, wrt="params")(model)
# SGD update
model[:] = jax.tree_map(lambda w, g: w - 0.1 * g, model["params"], grads)
# stateful update, no return needed
train_step(model, x, y)
The most interesting aspect of this design is that the code appears very imperative, as the state is automatically propagated in and out of the transformations. However, more consideration is needed to properly support jit's in_shardings and out_shardings arguments.
Certainly! I've revised the text to improve clarity and readability. Here are the updated sections:
Filtered Transformations
Filtered transformations offer more flexibility, as they can take Pytrees of references in any of their arguments and return Pytrees of references. They simply deref and reref all their inputs and outputs to transfer the Pytrees across the transformation. In general, they have the following properties:
- They behave as pure functions.
- They don't handle any global state except for Refx's trace state.
Currently, nnx.jit_filter is the only filtered transformation.
Here's an example of how a train_step function can be implemented using nnx.jit_filter and nnx.grad:
@nnx.jit_filter
def train_step(model, x, y):
def loss_fn(model):
y_pred = model(x)
return jax.numpy.mean((y_pred - y) ** 2)
# compute gradient
grads = nnx.grad(loss_fn, wrt="params")(model)
# SGD update
model[:] = jax.tree_map(lambda w, g: w - 0.1 * g, model["params"], grads)
return model
model = train_step(model, x, y)
Filtered transformations must output any state they want to propagate but have more flexibility in how they handle it. Adding support for jit's in_shardings and out_shardings arguments is likely more straightforward than with stateful transformations.
Partition API
The partition API enables splitting a Module's state into sets of reference-less Partitions, this provides a general way of interacting with vanilla JAX transformations.
Here's an example of how a train_step function can be implemented using the partition API:
modeldef: ModuleDef[Linear]
partitions, modeldef = model.partition()
params = partitions["params"]
@jax.jit
def train_step(params, x, y):
def loss_fn(params):
model: Linear = modeldef.reref(params)
y_pred = model(x)
return jax.numpy.mean((y_pred - y) ** 2)
# compute gradient
grads = jax.grad(loss_fn)(params)
# SGD update
params = jax.tree_map(lambda w, g: w - 0.1 * g, params, grads)
return params
params = train_step(params, x, y)
The main benefit of the partition API is its compatibility with other JAX tools, as the training step can be written using regular JAX transformations. The main drawback is that it's more verbose.
Case Studies
Shared State
In NNX, you can create modules that share state between them. This is useful when designing complex neural network architectures, as it allows you to reuse certain layers and reduce the number of learnable parameters.
Here's an example of creating a module with shared state:
class Block(nnx.Module):
def __init__(self, linear: nnx.Linear):
self.linear = linear
self.bn = nnx.BatchNorm(2)
def __call__(self, x):
return nnx.relu(self.bn(self.linear(x)))
class Model(nnx.Module):
def __init__(self):
shared = nnx.Linear(2, 2)
self.block1 = Block(shared)
self.block2 = Block(shared)
def __call__(self, x):
x = self.block1(x)
x = self.block2(x)
return x
In this example, the Model module contains two instances of the Block module. Each instance shares the same nnx.Linear module. To run the model, you can use the apply method to set the use_running_average flag for all BatchNorm modules.
Here's an example of computing the loss for a Model instance:
def loss_fn(model: Model, x: jax.Array, y: jax.Array):
y_pred = model.apply(use_running_average=False)(x)
return jnp.mean((y - y_pred) ** 2)
It's important to note that the state for the shared nnx.Linear module will be kept in sync at all times on both Block instances, including during gradient updates.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file nnx-0.0.3.tar.gz.
File metadata
- Download URL: nnx-0.0.3.tar.gz
- Upload date:
- Size: 33.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.4.0 CPython/3.8.10 Linux/5.13.0-1027-gcp
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dcb3af43863baa3d9d2f365c371e48032baa84db2b9cd83025f36049950a982a
|
|
| MD5 |
2854810b29b09d4cb039888b722114ed
|
|
| BLAKE2b-256 |
e25306f423dba4035c71cd6863ec55af1036fe03987c85d555284d173b81484c
|
File details
Details for the file nnx-0.0.3-py3-none-any.whl.
File metadata
- Download URL: nnx-0.0.3-py3-none-any.whl
- Upload date:
- Size: 35.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.4.0 CPython/3.8.10 Linux/5.13.0-1027-gcp
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
082181b5a6c9235fc4782806d663c7374126622115751bebaf7e3a51968f8a47
|
|
| MD5 |
44d025c99bb5f01f328a44fa12577670
|
|
| BLAKE2b-256 |
a4c951004cbd36f08c01e6e4bc2cb492e94942de309b94e2256a551b904c2a74
|