Lightweight Cohort Flow Diagrams built on Matplotlib — with an interactive browser-based generator.
Project description

pycohortflow
Lightweight cohort flow diagrams built on Matplotlib.
Turn a plain Python list of cohort steps into a publication-ready vertical flow chart with a single function call. Colours, fonts, spacing, and box geometry are fully customisable via TOML configuration files.
No Python installed?
Use the Interactive Generator to build diagrams directly in your browser and export them as SVG, PNG or PDF.
Installation
pip install pycohortflow
Quick Start
from pycohortflow import plot_cohort_flow_diagram
import matplotlib.pyplot as plt
data = [
{"heading": "Registered Patients", "N": 350,
"description": "Total patients registered in database"},
{"heading": "Screening", "N": 150,
"exclusion_description": "Did not meet inclusion criteria"},
{"heading": "Eligible", "N": 120,
"exclusion_description": "Declined / Lost to follow-up"},
{"heading": "Final Analysis", "N": 115,
"exclusion_description": "Data incomplete"},
]
fig, ax = plot_cohort_flow_diagram(
data,
figure_title="Clinical Cohort Flow Diagram",
)
plt.show()
Built-in Styles
Two styles ship with the package:
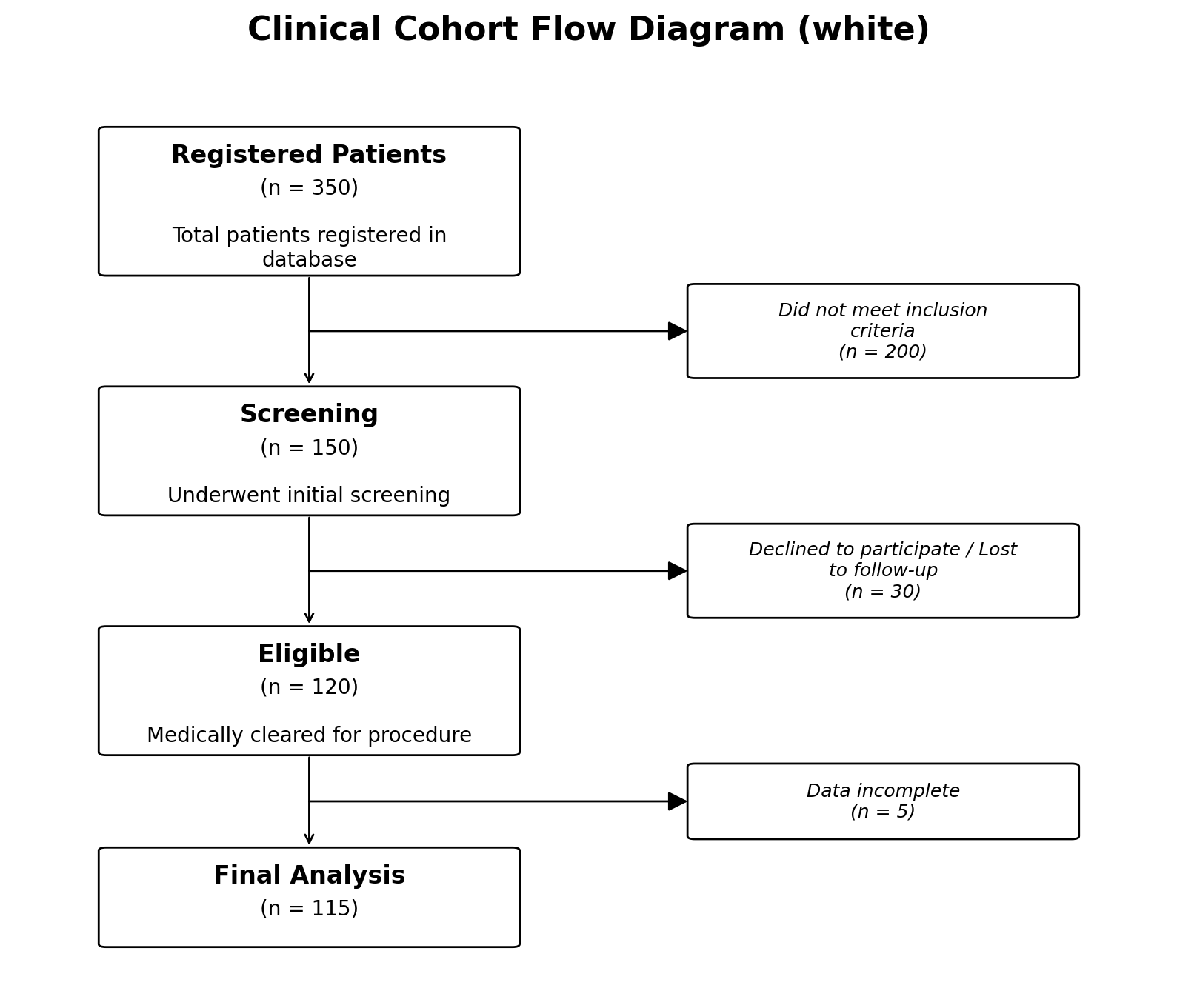
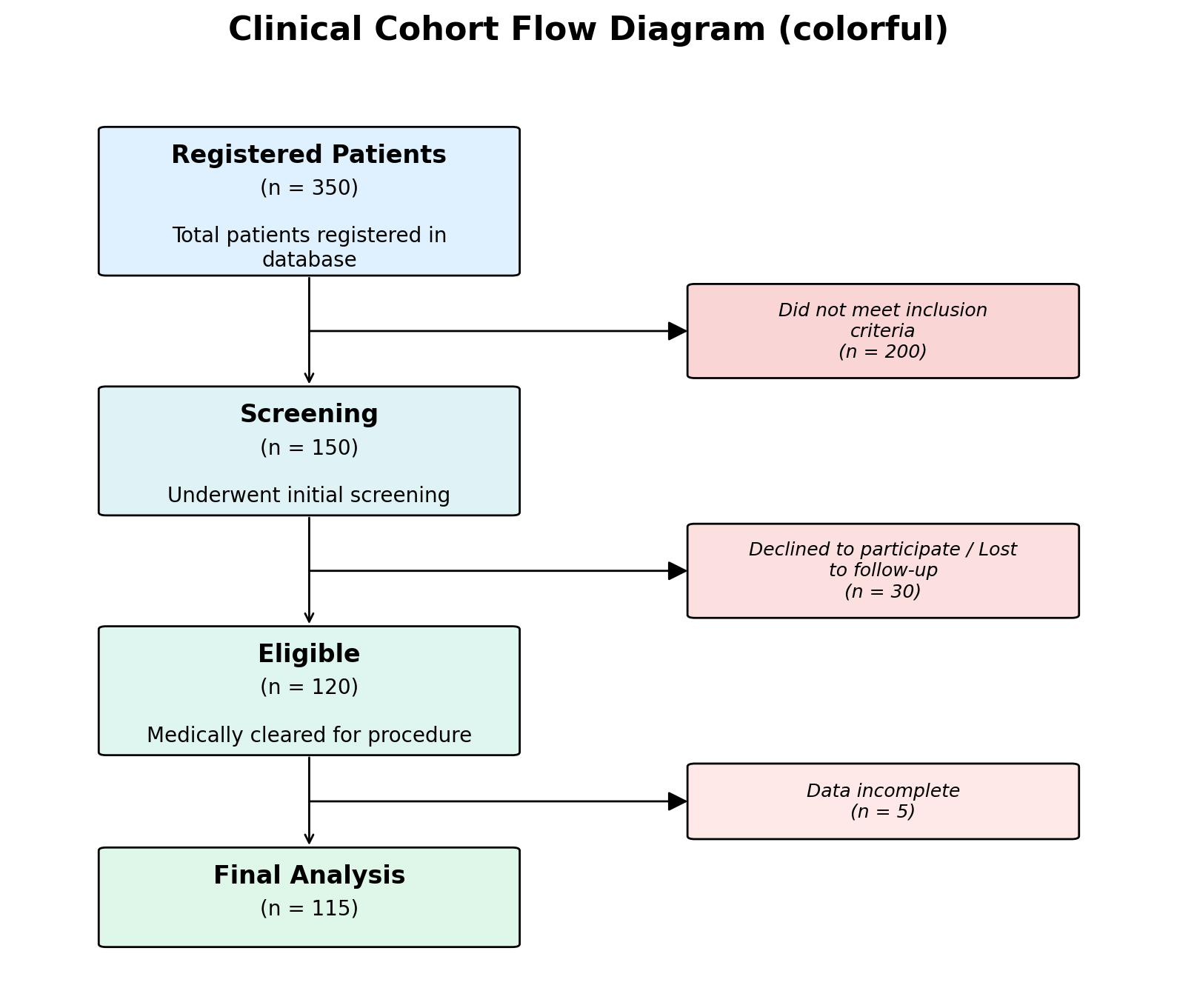
# Clean white boxes (default)
fig, ax = plot_cohort_flow_diagram(data, style="white")
# Pastel gradient backgrounds
fig, ax = plot_cohort_flow_diagram(data, style="colorful")
# Transparent figure background (for slides / posters)
fig, ax = plot_cohort_flow_diagram(data, transparent=True)
Customisation
Create a TOML file with only the values you want to override:
[figure]
dpi = 300
[colors]
main_start = "#cce5ff"
main_end = "#d4edda"
Then pass it when plotting:
fig, ax = plot_cohort_flow_diagram(data, style_config_path="my_style.toml")
See the full documentation for all available options.
Saving Figures
fig, ax = plot_cohort_flow_diagram(
data,
save_dir="output",
img_name="flow_chart",
save_format=["png", "svg", "pdf"],
)
Requirements
- Python >= 3.9
- Matplotlib >= 3.5
License
AGPL-3.0 license
Roadmap
- Add: Python-based PRISMA2020 style generation, if necessary
- Add: Small Diagram version
- Add: Verbose option for communication of saving options
- Add: Diagrams for multiple Arms (e.g. something like CONSORT style)
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pycohortflow-0.1.2.tar.gz.
File metadata
- Download URL: pycohortflow-0.1.2.tar.gz
- Upload date:
- Size: 42.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f4cdafb0f21c844233b724b043db3ce1bfe0bd4a174885246578b054b2ffae32
|
|
| MD5 |
5ac918a1e678ca51cef0e6e929efc449
|
|
| BLAKE2b-256 |
9792622b929dcdc1033e9c2cda2ceb17c811b78235836a6fcb924c5e52c15e25
|
File details
Details for the file pycohortflow-0.1.2-py3-none-any.whl.
File metadata
- Download URL: pycohortflow-0.1.2-py3-none-any.whl
- Upload date:
- Size: 26.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
40f6c97cfe279f6aab0907413f1805650306dbb1762b935d00a84f82c18778ca
|
|
| MD5 |
4fd210f75a4127d6b99bddcd3808e14c
|
|
| BLAKE2b-256 |
eaa6d8f73547cfb848acb50bf99ceb02054cad4fd90942b26d5326002fd0be8d
|
















