Python scripting library for generating designs readable by scadnano.
Project description
scadnano-python-package
scadnano ("scriptable-cadnano") is a program for designing synthetic DNA structures such as DNA origami. The scadnano Python module source code repository here) is a library for programmatically creating and editing these nanostructures.
If you find scadnano useful in a scientific project, please cite its associated paper:
scadnano: A browser-based, scriptable tool for designing DNA nanostructures.
David Doty, Benjamin L Lee, and Tristan Stérin.
DNA 2020: Proceedings of the 26th International Conference on DNA Computing and Molecular Programming
[ paper | BibTeX ]
Overview
This module is used to write Python scripts outputting .dna files readable by scadnano, a web application useful for displaying and manually editing these structures. The purpose of this module is to help automate some of the task of creating DNA designs, as well as making large-scale changes to them that are easier to describe programmatically than to do by hand in scadnano.
Early versions of this project didn't have well-defined versions. However, we will try to announce breaking changes (and possibly new features) under the GitHub releases page. The version numbers in this Python library repo and the web interface repo won't always advance at the same time.
Following semantic versioning, version numbers are major.minor.patch, i.e., version 0.9.2 has minor version number 9. Prior to version 1.0.0, when a breaking change is made, this will increment the minor version (for example, going from 0.9.4 to 0.10.0). After version 1.0.0, breaking changes will increment the major version.
Reporting issues
Please report issues in the web interface at the scadnano web interface GitHub repository, and report issues in the Python scripting library at the scadnano Python package GitHub repository.
Installation
The scadnano Python package requires Python version 3.7 or later. If you do not have that version (or later) of Python installed, follow this link to install it. To check your current version of Python, open a command line and type
python --version
If you are using Python 3.6 and do not wish to upgrade, you can install a package providing the required features. There is a Python 3.7 module used by scadnano, which is not present in 3.6: the dataclasses module. If you have at least Python version 3.6, you can install a backported library providing this module: dataclasses backport
Once Python is installed (and the dataclasses backport if you have Python version 3.6), there are two ways you can install the scadnano Python package:
-
pip (recommended)
Use pip to install the package by executing the following at the command line:
pip install scadnanoIf your Python installation does not already have pip installed, you may have to install it. Executing this Python script should work; see also https://docs.python.org/3/installing/index.html or https://www.liquidweb.com/kb/install-pip-windows/.
First, check your version of
pipby typingpip --versionIt should say something like
pip 19.3.1 from ...lib\site-packages\pip (python 3.8)If the version of Python at the end is Python 3.7 or higher, you are good. If it is version 2.7 or lower, type
pip3 --versionIf that works and shows Python 3.7 or higher, you are good, but you should type
pip3in the subsequent instructions instead ofpip.If it shows Python 3.6, install the dataclasses backport module via
pip install dataclassesIf it shows Python 3.5 or lower, then you will need to upgrade your Python version (recommended Python 3.7 or higher).
-
download
Note: If you are reading this on the PyPI website, the links below won't work. They are relative links intended to be read on the GitHub README page.
As a simple alternative (in case you run into trouble using pip), you can download and place the following files (located in the scadnano/ subfolder) in your PYTHONPATH (e.g., in the same directory as the scripts you are running). To download them, right-click on "Raw" near the top and select (in Chrome or Firefox) "Save link as...".
- required: scadnano.py
- optional: modifications.py; This contains some common DNA modifications such as biotin and Cy3.
- optional: origami_rectangle.py; This can help create origami rectangles, but it is not necessary to use scadnano.
- optional: _version.py This ensures that the current version number is written into any
.dnafiles written by the library; otherwise it may be out of date. (Which should not matter for the most part.)
The scadnano package uses the Python package xlwt to write Excel files, so xlwt must be installed in order to call the method
Design.write_idt_plate_excel_file()to export an Excel file with DNA sequences. To install xlwt, typepip install xlwtat the command line. (If you instead use pip to install the scadnano package, xlwt will be automatically installed.)
Example
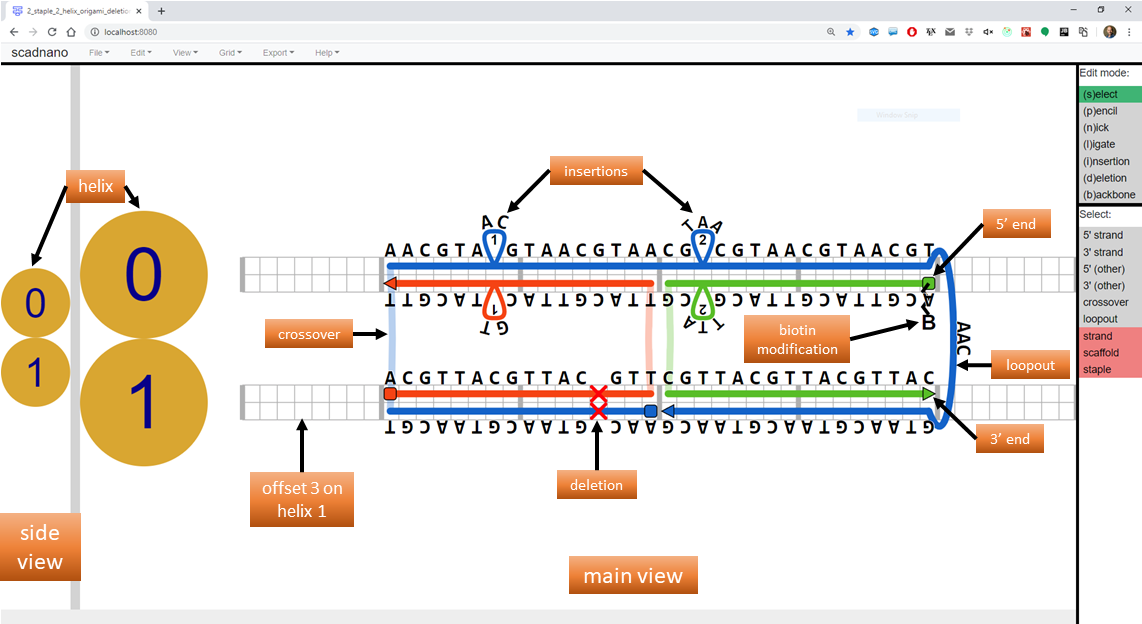
Consider the following design:
The following Python script produces this design.
import scadnano as sc
import modifications as mod
def create_design():
# helices
helices = [sc.Helix(max_offset=48), sc.Helix(max_offset=48)]
# left staple
stap_left_domain1 = sc.Domain(helix=1, forward=True, start=8, end=24)
stap_left_domain0 = sc.Domain(helix=0, forward=False, start=8, end=24)
stap_left = sc.Strand(domains=[stap_left_domain1, stap_left_domain0])
# right staple
stap_right_domain0 = sc.Domain(helix=0, forward=False, start=24, end=40)
stap_right_domain1 = sc.Domain(helix=1, forward=True, start=24, end=40)
stap_right = sc.Strand(domains=[stap_right_domain0, stap_right_domain1])
stap_right.set_modification_5p(mod.biotin_5p)
# scaffold
scaf_domain1_left = sc.Domain(helix=1, forward=False, start=8, end=24)
scaf_domain0 = sc.Domain(helix=0, forward=True, start=8, end=40)
loopout = sc.Loopout(length=3)
scaf_domain1_right = sc.Domain(helix=1, forward=False, start=24, end=40)
scaf = sc.Strand(domains=[scaf_domain1_left, scaf_domain0, loopout, scaf_domain1_right], is_scaffold=True)
# whole design
design = sc.Design(helices=helices, strands=[scaf, stap_left, stap_right], grid=sc.square)
# deletions and insertions added to design are added to both strands on a helix
design.add_deletion(helix=1, offset=20)
design.add_insertion(helix=0, offset=14, length=1)
design.add_insertion(helix=0, offset=26, length=2)
# also assigns complement to strands other than scaf bound to it
design.assign_dna(scaf, 'AACGT' * 18)
return design
if __name__ == '__main__':
design = create_design()
design.write_scadnano_file(directory='output_designs')
Running the code above produces the .dna JSON file shown in the web interface README. That section explains many of the terms used in the code (domain, helix, loopout, insertion, etc.).
abbreviated syntax with chained methods
Instead of explicitly creating variables and objects representing each domain in each strand, there is a shorter syntax using chained method calls. Instead of the above, create only the helices first, then create the Design. Then strands can be added using a shorter syntax, to describe how to draw the strand starting at the 5' end and moving to the 3' end. The following is a modified version of the above script using these chained methods
import scadnano as sc
import modifications as mod
def create_design():
# helices
helices = [sc.Helix(max_offset=48), sc.Helix(max_offset=48)]
# whole design
design = sc.Design(helices=helices, strands=[], grid=sc.square)
# left staple
design.strand(1, 8).to(24).cross(0).to(8)
# right staple
design.strand(0, 40).to(24).cross(1).to(40).with_modification_5p(mod.biotin_5p)
# scaffold
design.strand(1, 24).to(8).cross(0).to(40).loopout(1, 3).to(24).as_scaffold()
# deletions and insertions added to design are added to both strands on a helix
design.add_deletion(helix=1, offset=20)
design.add_insertion(helix=0, offset=14, length=1)
design.add_insertion(helix=0, offset=26, length=2)
# also assigns complement to strands other than scaf bound to it
design.assign_dna(design.scaffold, 'AACGT' * 18)
return design
if __name__ == '__main__':
design = create_design()
design.write_scadnano_file(directory='output_designs')
Documentation is available in the API docs.
Documentation
Online documentation of the package API is located here: https://scadnano-python-package.readthedocs.io
Tutorial
A tutorial shows how to create a "standard" 24-helix DNA origami rectangle using the scripting library.
Other examples
Several example scripts are located in the examples/ subfolder. Their output is contained in the examples/output_designs/ subfolder.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file scadnano-0.10.0.tar.gz.
File metadata
- Download URL: scadnano-0.10.0.tar.gz
- Upload date:
- Size: 68.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/41.2.0 requests-toolbelt/0.9.1 tqdm/4.48.0 CPython/3.8.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
92e8e599417d27b7778d6393df9d1ec734e017d26a397c981989673d7046d7da
|
|
| MD5 |
7b4102ee705678fe5bce4af94118220c
|
|
| BLAKE2b-256 |
29c50656de35eea351d255fb2c58041c753c606817d1cc64230594caadff7328
|
File details
Details for the file scadnano-0.10.0-py3-none-any.whl.
File metadata
- Download URL: scadnano-0.10.0-py3-none-any.whl
- Upload date:
- Size: 66.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/41.2.0 requests-toolbelt/0.9.1 tqdm/4.48.0 CPython/3.8.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6cdefdab7d5a485589759c00c87c2b3dbc809cd16894a76d57899764adca23fb
|
|
| MD5 |
bd34cd0c2a1dfe0ccdda40b76823d3f4
|
|
| BLAKE2b-256 |
218e2a562959bf18ee9033fc9bdc26230f3523d7575836f8efc1419910002e3e
|