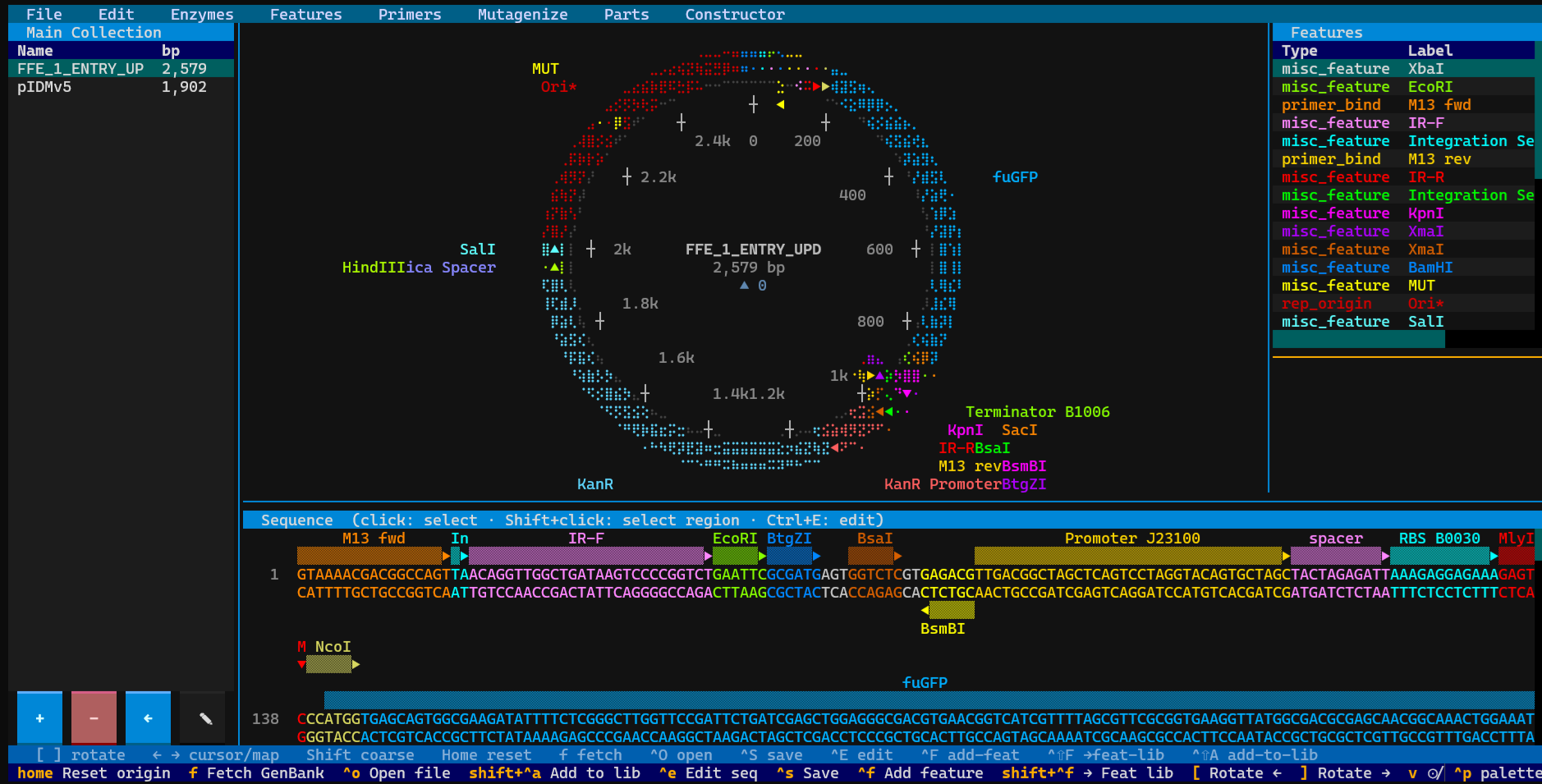
Terminal-based circular plasmid map viewer, sequence editor, and Primer3/Golden Braid primer design workbench
Project description
SpliceCraft
A terminal-based circular plasmid map viewer, sequence editor, primer design workbench, and Golden Braid parts domesticator — rendered entirely in your shell. Fetch any GenBank record by accession, load local files, organize plasmids into named collections, design diagnostic / cloning / Golden Braid primers with Primer3, run SOE-PCR site-directed mutagenesis on any CDS, and edit sequences — without ever leaving the terminal.
Quick start
⚠️ Beta software. SpliceCraft is under active development and the API, UI, and on-disk file formats may change between releases. Your data files (
plasmid_library.json,parts_bin.json,primers.json) are auto-backed up to*.bakon every save, but please keep your own off-disk copies of anything critical. Do not use SpliceCraft as your sole system of record for cloning work until it exits beta.
pipx install splicecraft
splicecraft # empty canvas
splicecraft L09137 # fetch pUC19 from NCBI
splicecraft myplasmid.gb # open a local GenBank (.gb/.gbk) or SnapGene (.dna) file
That's it. pipx creates an isolated virtual environment for SpliceCraft
and its dependencies, so it won't clash with system Python packages. If you
don't have pipx yet: sudo apt install pipx on Debian/Ubuntu/WSL2,
brew install pipx on macOS, or python -m pip install --user pipx elsewhere.
User data lives in the platform-appropriate data directory (see the
Installation section below).
Features
Core visualization & editing
- Braille dot-matrix circular map — plasmids rendered as crisp Unicode braille rings with per-strand feature arcs, directional arrowheads, and proximity-placed labels
- Linear map view — toggle with
vfor a horizontal strip layout - Dithered sequence panel — per-base DNA viewer with feature bars, restriction site overlays, and double-stranded display
- Inline amino-acid translation — every CDS shows its protein right above
(forward) or below (reverse) its DNA bar, one letter per codon midpoint, in
the feature's colour. Wrap-around CDSes translate correctly across the origin.
Click an AA letter to highlight that codon's three bases on the strand —
Ctrl+Ccopies the codon - Per-strand restriction-cut visualization — clicking a sticky cutter (EcoRI, HindIII, …) tints the upstream bases blue and downstream bases red, with the staggered overhang shown as different colours on the two strands
- Live NCBI fetch — pull any GenBank record by accession number on demand
- Local file support — open
.gb/.gbk(GenBank) or.dna(SnapGene native) files directly from disk - Free rotation — spin the origin with
[/]or arrow keys (when the map has focus) - Restriction enzyme overlay — 200+ NEB enzymes including Type IIS (BsaI, BsmBI, BbsI, SapI) with visible recognition arcs + cut markers
Libraries (all persist to JSON)
- Plasmid collections — organize plasmids into named buckets (e.g. "yeast project", "E. coli toolkit"). Library panel toggles between a collection list and the active collection's plasmids; a default "Main Collection" is created on first run, with full add / remove / rename CRUD from the panel
- Plasmid library — auto-saves on import, mirrors live into the active collection, survives restarts, rename + handslip-protected delete; unsaved edits prompt Save / Discard / Cancel before navigating away
- Library fuzzy search — type into the search box at the top of the library panel to filter the visible table by subsequence match (case-insensitive, not necessarily contiguous). Empty submit restores the full list.
- Parts Bin — Golden Braid L0 parts catalog with user-domesticated parts including sequences and primer pairs
- Primer library — all designed primers with Tm, length, date, status (Designed / Ordered / Validated), multi-select for batch operations
- Feature library — reusable feature snippets with per-entry color and strand, centralized workbench for browse / edit / rename / recolor / delete
Primer design (Primer3)
- Detection primers — diagnostic PCR; Primer3 picks the ideal pair within a selected region, 450-550 bp product by default (configurable)
- Cloning primers — RE-site tails + GCGC padding; 30+ common enzymes or type a custom recognition sequence
- Golden Braid primers — Esp3I / BsmBI domestication for all L0 positions (Promoter, 5' UTR, CDS, CDS-NS, C-tag, Terminator). Splitting L0 (Esp3I) from L1 (BsaI) lets domesticated parts survive the L1 Golden Gate reaction without re-cutting.
- Generic primers — simple binding primers, no tails
- Primers can be added to the plasmid map as
primer_bindfeatures - Scrollable
TextAreafor custom sequence input; highlighted text = target
Mutagenesis
- SOE-PCR site-directed mutagenesis — design 4-primer SOE sets for any W140F-style mutation. CDS source can be the loaded plasmid, a library entry, a domesticated part from the Parts Bin, or a free-form protein sequence (auto-harmonized to the active codon table). Edge cases (mutation within 60 nt of a CDS end) auto-fall back to a 2-primer modified-outer PCR.
Feature operations
- Feature sidebar — click a row to highlight on map; click the map to select the feature under the cursor
- Undo/redo — 50-deep snapshot stack for all sequence edits
- Delete protection — focus-aware Delete key; confirmation modal (default focus = No) for library entries
- Clipboard — OSC-52 copy works in Windows Terminal, iTerm2, modern WSL
Data safety
- Atomic saves — all JSON files written via tempfile +
os.replace - Automatic backups — every save writes
*.json.bakbefore overwriting - Corrupt-file recovery — missing files don't crash; corrupt files auto-
restore from
.bakwith a warning notification on startup
Installation
Requires Python 3.10+.
With pipx (recommended)
pipx install splicecraft
pipx installs SpliceCraft (and its Textual / Biopython / primer3-py /
platformdirs deps) into an isolated virtual environment and places the
splicecraft command on your PATH. This is the right approach on
modern Debian, Ubuntu, Fedora, and WSL2, where pip install into the
system Python is blocked by PEP 668.
If you don't already have pipx:
sudo apt install pipx # Debian / Ubuntu / WSL2
brew install pipx # macOS
python -m pip install --user pipx # everywhere else
pipx ensurepath # one-time; adds ~/.local/bin to PATH
With pip inside a venv
If you prefer a plain pip workflow, use a virtual environment:
python3 -m venv ~/.venvs/splicecraft
~/.venvs/splicecraft/bin/pip install splicecraft
~/.venvs/splicecraft/bin/splicecraft
(Plain pip install splicecraft into system Python works on older
distros and inside conda envs, but will be rejected by PEP 668 on any
recent Debian-family system — use pipx or a venv instead.)
User data location
User data (library, parts bin, primers) lives in the platform-appropriate data directory:
| Platform | Path |
|---|---|
| Linux | ~/.local/share/splicecraft/ |
| macOS | ~/Library/Application Support/splicecraft/ |
| Windows | %APPDATA%\splicecraft\ |
Override with SPLICECRAFT_DATA_DIR=/path/to/dir splicecraft.
From source
git clone https://github.com/Binomica-Labs/SpliceCraft.git
cd SpliceCraft
pip install -e . # inside a venv, or pass --break-system-packages
Usage
splicecraft # empty canvas
splicecraft L09137 # fetch pUC19 from NCBI on launch
splicecraft myplasmid.gb # open a local GenBank (.gb/.gbk) or SnapGene (.dna) file
splicecraft --version # print version
splicecraft --help # quick usage hint
If running from a git clone (pip install -e .), the same commands work;
you can also still run python3 splicecraft.py directly.
Key Bindings
Main screen
| Key | Description |
|---|---|
[ / ] |
Rotate map origin left / right (when map focused) |
← / → |
Same as [ / ] (when map focused) |
↑ |
Reset origin to 0 (when map focused) |
Shift+[/] |
Rotate coarse (10× step) |
Home |
Reset origin to 0 |
, / . |
Circular map aspect wider / taller |
v |
Toggle circular ↔ linear map |
l |
Toggle feature label connector lines |
r |
Toggle restriction-site overlay |
f |
Fetch a record from NCBI by accession |
Ctrl+O |
Open a .gb file from disk |
Ctrl+Shift+A |
Add current plasmid to the library |
Ctrl+E |
Enter sequence editor mode |
Ctrl+S |
Save edits to file |
Ctrl+F |
Add a new feature (from cursor or blank) |
Ctrl+Shift+F |
Capture selection / feature → Feature library |
Enter |
Highlight the feature enclosing the seq cursor |
Delete |
Context-aware delete (feature or library entry) |
Ctrl+Z |
Undo |
Ctrl+Shift+Z |
Redo |
Ctrl+C |
Copy selection (top strand 5'→3', or AA when CDS highlighted) |
Alt+C |
Copy selection (bottom strand, reverse-complement) |
q |
Quit |
Primer Design screen
| Key | Description |
|---|---|
esc |
Close primer screen |
m |
Mark / unmark primer under cursor (★) |
M |
Mark / unmark all primers |
S |
Cycle status: Designed → Ordered → Validated |
Tab |
Cycle focus between fields |
Mouse
| Action | Description |
|---|---|
| Click DNA row | Place cursor at that base |
| Click feature bar | Highlight the feature, set cursor at its 5' end |
| Click AA letter | Highlight that codon's three bases on the strand |
| Click restriction site | Highlight recognition span; tint upstream blue / downstream red per strand |
| Double-click | Select full feature span |
| Drag | Select a sequence range |
| Scroll wheel | Rotate map (when over map panel) |
| Click backbone | Clear all panel highlights |
Menus
| Menu | Items |
|---|---|
| File | Open .gb file · Fetch from NCBI · Add to Library · Save · Export GenBank · Collections · Quit |
| Edit | Edit Sequence · Undo · Redo · Add Feature · Capture → feat-lib · Delete Feature |
| Enzymes | Show RE sites · Unique cutters · 6+/4+ bp sites · Connectors |
| Features | Opens the Feature Library workbench |
| Primers | Opens the full-screen Primer Design workbench |
| Mutagenize | Opens the SOE-PCR site-directed mutagenesis designer (4-source CDS picker) |
| Parts | Opens the Parts Bin (Golden Braid L0 parts catalog) |
| Constructor | Opens the Assembly Constructor for TU building |
Requirements
| Package | Version | Purpose |
|---|---|---|
| Python | ≥ 3.10 | Runtime |
| Textual | ≥ 8.2.3 | TUI framework and rendering engine |
| Rich | ≥ 14.0 | Terminal rendering (Textual dependency) |
| Biopython | ≥ 1.87 | GenBank parsing and NCBI Entrez fetch |
| primer3-py | ≥ 2.3.0 | Primer design (Tm, thermodynamic screening) |
| platformdirs | ≥ 4.2 | Cross-platform user-data directory |
| pytest | ≥ 9.0 | Test suite (dev only) |
| pytest-asyncio | ≥ 1.3 | Async test support (dev only) |
Data files
All user data persists as human-readable JSON in the platform-appropriate user data directory:
| File | Purpose |
|---|---|
collections.json |
Named collections of plasmids; source of truth |
plasmid_library.json |
Live mirror of the active collection's plasmids |
parts_bin.json |
User-domesticated Golden Braid parts |
primers.json |
Designed primer library |
features.json |
Reusable feature snippets |
feature_colors.json |
Per-type feature color overrides |
codon_tables.json |
Cached codon-usage tables fetched from Kazusa |
cloning_grammars.json |
User-defined cloning grammars (Golden Braid / MoClo) |
settings.json |
App preferences (active collection, active grammar, ...) |
*.json.bak |
Automatic backup — written before each save |
A manual backup rotation happens on every save so accidental data loss is
always recoverable via the .bak file.
Tests
python3 -m pytest -n auto -q # full suite (1,009 tests, ~2 min on 8 cores)
python3 -m pytest tests/test_dna_sanity.py # biology correctness only (< 2s)
Codebase tour
SpliceCraft is a single-file Python app (splicecraft.py, ~17,900 lines) built on
Textual + Biopython. The single-file layout is intentional — no import puzzles,
and everything is greppable from one place.
Layout at a glance
| Region | Contents |
|---|---|
| Top of file | imports, user data dir (platformdirs), rotating session-tagged logger, _safe_save_json / _safe_load_json (atomic writes + .bak recovery) |
| Persistence | Library + parts-bin + primers + features + feature-colors + codon-tables + cloning-grammars + settings + collections (collection-driven model with active-pointer in settings.json + library mirror sync) |
| Biology core | NEB enzyme catalog, IUPAC-aware _rc, _scan_restriction_sites (palindrome- and wrap-aware), sequence panel renderer, _translate_cds |
| I/O | fetch_genbank (NCBI Entrez), load_genbank (.gb / .gbk / .dna), GenBank/FASTA export |
| Rendering | _Canvas + _BrailleCanvas (sub-cell dot matrix), PlasmidMap, SequencePanel, FeatureSidebar, LibraryPanel (two-mode: collections list ↔ active-collection plasmids), MenuBar |
| Workbenches | PrimerDesignScreen (detection / cloning / Golden Braid / generic), MutagenizeModal (SOE-PCR; 4-source CDS picker), DomesticatorModal + ConstructorModal (Golden Braid L0), FeatureLibraryScreen |
| Registries | codon-usage tables, parts bin, primer library, feature library, cloning grammars, plasmid collections — all persisted as JSON with automatic .bak backups |
| Controller | PlasmidApp — keybindings, undo/redo (50-deep snapshot stack), @work threads for NCBI / Kazusa, collection-driven startup migration in compose() |
Contributor docs
CLAUDE.md at the repo root is the agent + contributor handover document.
It covers the 10 sacred invariants (biology correctness rules with test coverage),
the error-handling convention, performance notes, and modular recipes for adding
modals, workers, menus, or persisted libraries without tripping the invariants.
Read it before touching the rendering layer, record pipeline, or primer design.
Running the suite
python3 -m pytest -n auto -q # full suite (~2 min)
python3 -m pytest tests/test_dna_sanity.py # biology only (< 2 s)
All tests run offline against synthetic SeqRecords and monkeypatched data paths;
an autouse fixture in tests/conftest.py guarantees no test can write to real
user files.
License
MIT
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file splicecraft-0.4.5.tar.gz.
File metadata
- Download URL: splicecraft-0.4.5.tar.gz
- Upload date:
- Size: 484.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f510d807451ab67c27883197071fba9699d0655e44ab8403aae762d88d54941a
|
|
| MD5 |
eb1288aa57cf487a0b60ffac861d49f4
|
|
| BLAKE2b-256 |
c70268127753c204c344a9905e414b6431b2efb53cc0855f15be013a0f9567ce
|
Provenance
The following attestation bundles were made for splicecraft-0.4.5.tar.gz:
Publisher:
publish.yml on Binomica-Labs/SpliceCraft
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
splicecraft-0.4.5.tar.gz -
Subject digest:
f510d807451ab67c27883197071fba9699d0655e44ab8403aae762d88d54941a - Sigstore transparency entry: 1409733538
- Sigstore integration time:
-
Permalink:
Binomica-Labs/SpliceCraft@2c6e4861d115281b7f614ff9a11830db546f51a7 -
Branch / Tag:
refs/tags/v0.4.5 - Owner: https://github.com/Binomica-Labs
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@2c6e4861d115281b7f614ff9a11830db546f51a7 -
Trigger Event:
push
-
Statement type:
File details
Details for the file splicecraft-0.4.5-py3-none-any.whl.
File metadata
- Download URL: splicecraft-0.4.5-py3-none-any.whl
- Upload date:
- Size: 212.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b26dc64a70f21b6c1e548bb9646de819d6a16baa311fbb776871f1c8dea223ba
|
|
| MD5 |
e4f8751fad14cb9a824aed6d35182c87
|
|
| BLAKE2b-256 |
dcf0b4552ac4bdc6a1323c5e67a0bdfbd358ffeb787032f482ef630e22055d45
|
Provenance
The following attestation bundles were made for splicecraft-0.4.5-py3-none-any.whl:
Publisher:
publish.yml on Binomica-Labs/SpliceCraft
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
splicecraft-0.4.5-py3-none-any.whl -
Subject digest:
b26dc64a70f21b6c1e548bb9646de819d6a16baa311fbb776871f1c8dea223ba - Sigstore transparency entry: 1409733572
- Sigstore integration time:
-
Permalink:
Binomica-Labs/SpliceCraft@2c6e4861d115281b7f614ff9a11830db546f51a7 -
Branch / Tag:
refs/tags/v0.4.5 - Owner: https://github.com/Binomica-Labs
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@2c6e4861d115281b7f614ff9a11830db546f51a7 -
Trigger Event:
push
-
Statement type: