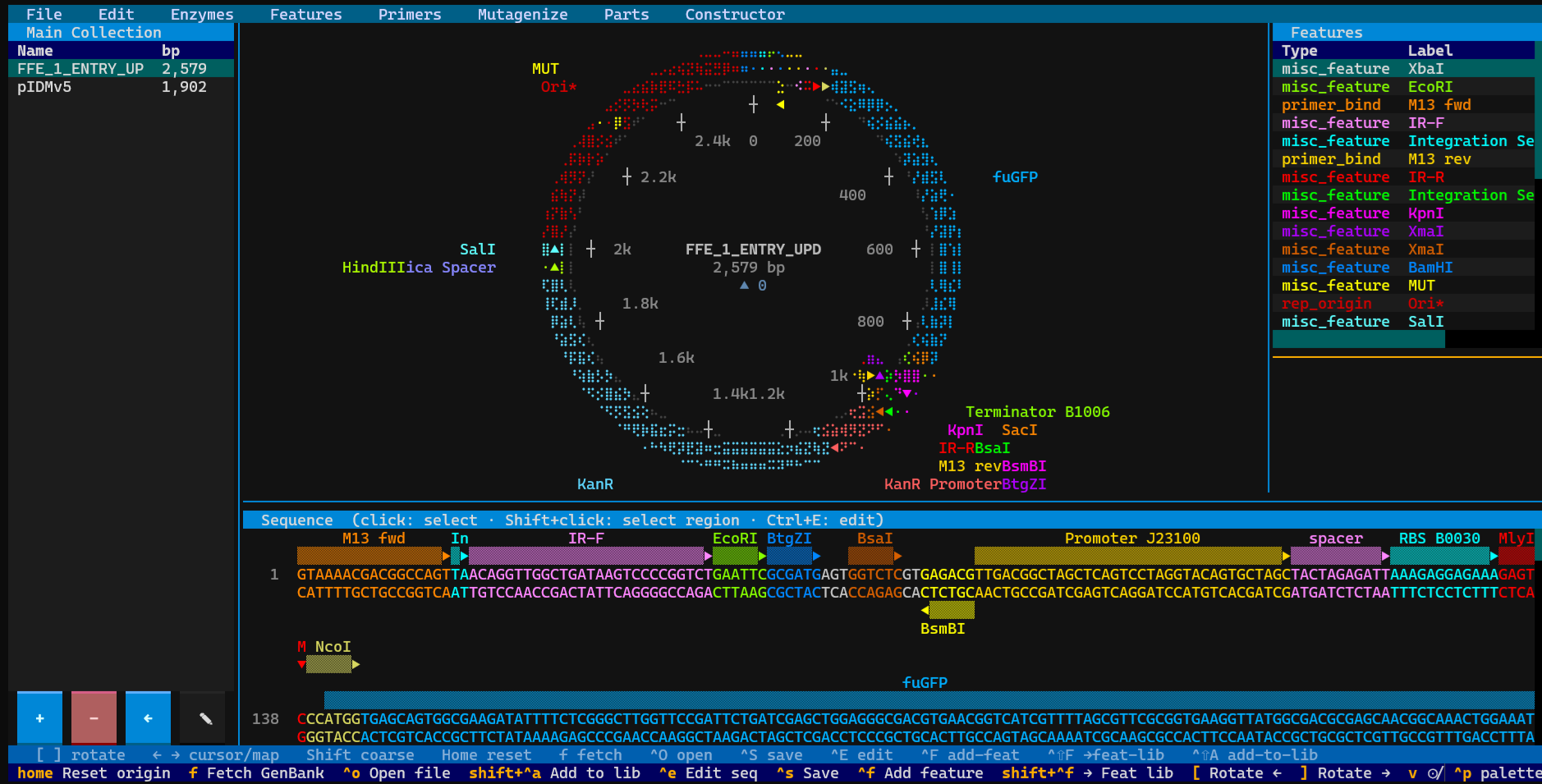
Terminal-based circular plasmid map viewer, sequence editor, and Primer3/Golden Braid primer design workbench
Project description
SpliceCraft
A plasmid workbench you live in. SpliceCraft is a terminal-native
viewer, sequence editor, primer + mutagenesis designer, Golden Braid /
MoClo cloning workbench, and in-process BLAST / HMMscan engine — all
rendered as crisp Unicode braille graphics in any modern terminal.
Fetch from NCBI, load .gb / .gbk / SnapGene .dna files (single or
in bulk), organize plasmids into named collections, design diagnostic /
cloning / Golden Braid primers via Primer3, run SOE-PCR site-directed
mutagenesis on any CDS, and search your own plasmid library by sequence
similarity — without ever leaving the shell.
Built for daily lab work. SpliceCraft is actively maintained by a practicing bioengineer who uses it as their primary day-to-day tool for plasmid design, cloning planning, and sequence triage. Bug reports come from the bench; releases ship from the bench. Every feature has a real-world job.
Robustness is a feature, not an afterthought
A workbench you trust your day to has to behave like one. SpliceCraft takes data safety and predictable behaviour as first-class design constraints:
- Atomic writes with on-disk backups. Every
*.jsonlibrary file goes throughtempfile.mkstemp+os.fsync+os.replace, with the prior version copied to*.json.bakbefore the new write lands. Mid-process crash, full-disk error, or hand-edit gone wrong — the panel re-loads from.bakon next launch with a corruption-recovery toast. - Crash-recovery autosave. Dirty edits debounce a 3-second write
to a per-record
.gbsnapshot. Power-cut your laptop mid-edit; the next launch surfaces the survivors. - 1,200+ tests (
pytest -n auto -q, ~3 min on 8 cores) anchored on ten sacred invariants for biology correctness: palindromic-enzyme scanning, reverse-strand coordinate handling, IUPAC reverse-complement including ambiguity codes, wrap-around feature math, atomic-save contract, undo deepcopy, etc. Property-based fuzzing (hypothesis) doubles up on the riskiest ones. Touching the biology primitives trips a test in under two seconds. - No external BLAST install.
pyhmmerships HMMER 3 source compiled in-wheel; BLASTN, BLASTP, and HMMscan all run in-process vianhmmer/phmmer/hmmscan, with a pure-Python ungapped fallback for queries too short for HMMER's profile builder. Zero external binaries; no PATH gymnastics. - Hardened input boundaries. Centralised sanitisers strip control
characters and Rich-markup metacharacters at every user-input boundary
(modals, agent-API endpoints, NCBI fetch, bulk-import filenames). A
hostile filename like
[red]EVIL[/red].dnaimports cleanly and renders as the literal characters in the panel and notifications. Modal-active gating keeps seq-cursor moves and selection slides from firing underneath active modals; CDS-divisibility gating prevents nonsensical AA strips on non-triple features; resite cuts carry baked per-strand offsets (no legacy schema crashes). - Refusal to start in a tiny terminal. Below 100×30, SpliceCraft prints a friendly resize-and-retry message and exits with code 2 rather than rendering a clipped, broken UI.
- Bulk-import with per-file failure isolation. The bulk SnapGene /
GenBank importer caps per-file size at 50 MB, skips zero-length
records, catches
OSError/PermissionError/struct.errorper-file, and dedups colliding ids by suffix. One bad file in a 500-plasmid archive does not abort the batch. - Drift defenses. Future-version JSON schemas load with a warning
and a
.bakrotation rather than crashing. Legacy schemas auto- rewrite on the next save. Migration code runs idempotently inApp.compose()before any child mounts so the panel always sees a consistent state.
⚠️ Beta software. SpliceCraft is under active development; the UI, on-disk file formats, and agent-API surface may evolve between releases. Your data files are auto-backed up to
*.bakon every save, but please keep your own off-disk copies of anything critical. The maintainer treats it as their primary workbench — but it should not yet be a project's sole system of record.
Quick start
pipx install splicecraft
splicecraft # empty canvas
splicecraft L09137 # fetch pUC19 from NCBI on launch
splicecraft myplasmid.gb # local GenBank (.gb/.gbk) or SnapGene (.dna) file
pipx creates an isolated virtual environment for SpliceCraft and its
dependencies, so it won't clash with system Python packages. If you
don't have pipx yet: sudo apt install pipx on Debian/Ubuntu/WSL2,
brew install pipx on macOS, or python -m pip install --user pipx
elsewhere. User data lives in the platform-appropriate data directory
(see Installation).
Press ? once running for the full keyboard-shortcut reference.
What you can do without leaving the terminal
View
- Braille dot-matrix circular maps — plasmids rendered as crisp
Unicode braille rings with per-strand feature arcs, directional
arrowheads, and proximity-placed labels.
vtoggles linear view. - Per-base sequence panel with two-strand display, wrap-aware feature lanes, restriction-site overlays, and inline AA translation (one letter per codon midpoint, in the CDS's colour, with wrap-CDS support across the origin). Click an AA letter to highlight the codon's three bases on the strand.
- Per-strand restriction-cut visualisation — clicking a sticky cutter (EcoRI, HindIII, BsaI, BsmBI, BbsI, …) tints upstream bases blue and downstream red, with the staggered overhang showing as different colours on the two strands.
- 200+ NEB enzymes including Type IIS scanners; toggle restriction
overlays with
r, filter to unique cutters / 6+ bp / connectors.
Edit
- In-place sequence edits with full undo / redo (50-deep snapshot stack, deepcopied SeqRecord). Per-plasmid undo stashes — switch records, edit, switch back, undo history is restored.
- Feature CRUD: add / merge / split / delete / rename / recolor
features; clipboard copies (top strand or reverse-complement bottom
strand). Mouse-drag selects ranges;
Enterhighlights the smallest feature enclosing the cursor. - Crash-recovery autosave writes a 3-second-debounced
.gbsnapshot to the data dir; survivors surface on next launch.
Cloning
- Cloning grammars — GB L0 (Esp3I) and MoClo Plant (BsaI) ship as
built-ins; user-defined grammars persist to
cloning_grammars.jsonand are editable inGrammarEditorModal. The active grammar parameterises the Domesticator, Parts Bin, and Constructor — change enzyme / overhang / forbidden-site set without code edits. - Domesticator — 4-source part picker (current map, library, Parts Bin, FASTA file). Auto-scrubs forbidden Type IIS sites in the CDS body via codon swap with cascade-prevention; primer tails follow the active grammar's pad / site / spacer / overhang.
- Parts Bin — domesticated parts catalog with per-grammar filtering; legacy parts default to GB L0; "Copy primed sequence" preserves the part's stored grammar.
- Constructor — assembly UI for chaining L0 parts into a TU.
- Primer design — detection / cloning / Golden Braid / generic via
Primer3; primers can be added to the map as
primer_bindfeatures or saved to the persistent primer library (Designed → Ordered → Validated lifecycle).
Mutagenesis
- SOE-PCR site-directed mutagenesis — design 4-primer SOE sets for any W140F-style point mutation. CDS source can be the loaded plasmid, a library entry, a Parts Bin part, or a free-form protein sequence (auto-optimised via the active codon table). Edge cases (mutation within 60 nt of a CDS end) auto-fall back to a 2-primer modified-outer PCR.
Search
- In-process BLAST (
Ctrl+B):- BLASTN (DNA → DNA) and BLASTP (protein → protein) via
pyhmmer.hmmer.nhmmer/phmmer(HMMER 3 in-process at C speed); pure-Python ungapped fallback for queries below the HMMER profile- builder minimum (20 bp / 6 aa). - HMMscan reads any HMMER 3
.hmm/.h3m/.h3pfile directly — point it at Pfam-A or any custom profile DB. Lazy file read so Pfam-scale (~1 GB) DBs don't pre-fetch into RAM. - DB build + search run in a
@work(thread=True)worker; UI stays responsive on a 50-plasmid index. 4-entry LRU DB cache, auto- invalidated on_save_collections.
- BLASTN (DNA → DNA) and BLASTP (protein → protein) via
- Six-frame ORF indexing (opt-in checkbox) for BLASTP against unannotated regions of plasmid backbones.
- New Plasmid modal (
Ctrl+N) — paste a sequence, optionally name- set topology, then either Create / Annotate-from-library
(substring match) / Annotate-via-BLAST (≥90% identity →
misc_feature).
- set topology, then either Create / Annotate-from-library
(substring match) / Annotate-via-BLAST (≥90% identity →
Library
- Plasmid collections — named buckets (e.g. "yeast project",
"E. coli toolkit"); the panel toggles between a collection list and
the active collection's plasmids. Atomic writes,
.bakper change. Save the loaded record withCtrl+Shift+A. - Bulk import a folder — from the collections-list view, click
+, type a name, and pick a folder via the embedded directory tree. Every.dna/.gb/.gbk/.genbankfile inside is loaded independently into a new collection; failures are isolated per file and surfaced in a notify summary. Designed for migrating a SnapGene archive in one shot. - Library fuzzy search — subsequence match (case-insensitive, non-contiguous) against the visible table.
- Feature library — reusable feature snippets (per-entry colour and strand) with a centralised browse / edit / rename / recolor / delete workbench.
Drive it from outside the GUI
- Agent API (
splicecraft --agent-api) exposes a localhost JSON API with bearer-token auth, covering every GUI action external AI agents need: get / set sequence, list features, add / update / delete features, export GenBank / FASTA, list library / collections, scan restriction sites, look up codon tables, optimize protein sequences, load files (chromosome-scale safe via the path-based loader). splicecraft-cli— stdlib-only sidecar (~50 ms cold start) that reads connection details from~/.local/share/splicecraft/agent_tokenand drives the running GUI. Intended for Claude Code, Cursor, aider, hand-rolled scripts, or any external automation.
Installation
Requires Python 3.10+ and a terminal of at least 100×30.
With pipx (recommended)
pipx install splicecraft
pipx installs SpliceCraft and its deps (Textual, Biopython,
primer3-py, platformdirs, pyhmmer) into an isolated virtual
environment and places the splicecraft command on your PATH. This
is the right approach on modern Debian, Ubuntu, Fedora, and WSL2,
where pip install into the system Python is blocked by
PEP 668.
If you don't already have pipx:
sudo apt install pipx # Debian / Ubuntu / WSL2
brew install pipx # macOS
python -m pip install --user pipx # everywhere else
pipx ensurepath # one-time; adds ~/.local/bin to PATH
With pip inside a venv
python3 -m venv ~/.venvs/splicecraft
~/.venvs/splicecraft/bin/pip install splicecraft
~/.venvs/splicecraft/bin/splicecraft
(Plain pip install splicecraft into system Python works on older
distros and inside conda envs, but is rejected by PEP 668 on any
recent Debian-family system — use pipx or a venv instead.)
From source
git clone https://github.com/Binomica-Labs/SpliceCraft.git
cd SpliceCraft
pip install -e . # inside a venv
User data location
User data (collections, library, parts, primers, features, codon tables, settings) lives in the platform-appropriate data directory:
| Platform | Path |
|---|---|
| Linux | ~/.local/share/splicecraft/ |
| macOS | ~/Library/Application Support/splicecraft/ |
| Windows | %APPDATA%\splicecraft\ |
Override with SPLICECRAFT_DATA_DIR=/path/to/dir splicecraft.
Key bindings
Press ? in-app for the full reference (rendered via Markdown so you
can drag-select a key combo to copy it).
Main screen
| Key | Description |
|---|---|
[ / ] |
Rotate map origin left / right (when map focused) |
← / → |
Same as [ / ] (when map focused) |
↑ |
Reset origin to 0 (when map focused) |
Shift+[/] |
Rotate coarse (10× step) |
, / . |
Circular map aspect wider / taller |
v |
Toggle circular ↔ linear map |
l |
Toggle feature label connector lines |
r |
Toggle restriction-site overlay |
f |
Fetch a record from NCBI by accession |
Ctrl+O |
Open a .gb / .gbk / .dna file from disk |
Ctrl+N |
New Plasmid (paste sequence + optional annotate) |
Ctrl+B |
BLAST modal (BLASTN / BLASTP / HMMscan) |
Ctrl+Shift+A |
Add current plasmid to the library |
Ctrl+A |
Select-all sequence |
Ctrl+E |
Enter sequence editor mode |
Ctrl+S |
Save edits to file |
Ctrl+F |
Add a new feature (from cursor or blank) |
Ctrl+Shift+F |
Capture selection / feature → Feature library |
Ctrl+P |
Primer Design workbench |
Enter |
Highlight the feature enclosing the seq cursor |
Delete |
Context-aware delete (feature or library entry) |
Ctrl+Z |
Undo |
Ctrl+Shift+Z / Ctrl+Y |
Redo |
Ctrl+C |
Copy selection (top strand 5'→3', or AA when CDS highlighted) |
Alt+C |
Copy selection (bottom strand, reverse-complement) |
Alt+D |
Toggle hover-status diagnostic row |
? |
Help modal |
Ctrl+Q |
Quit |
Mouse
| Action | Description |
|---|---|
| Click DNA row | Place cursor at that base |
| Click feature bar | Highlight the feature, set cursor at its 5' end |
| Click AA letter | Highlight that codon's three bases on the strand |
| Click restriction site | Highlight recognition span; tint upstream blue / downstream red per strand |
| Double-click | Select full feature span |
| Drag | Select a sequence range |
| Scroll wheel | Rotate map (when over map panel) |
| Click backbone | Clear all panel highlights |
Menus
| Menu | Items |
|---|---|
| File | Open · Fetch from NCBI · New Plasmid · Add to Library · Save · Export GenBank · Collections · Quit |
| Edit | Edit Sequence · Undo · Redo · Add Feature · Capture → feat-lib · Delete Feature |
| Enzymes | Show RE sites · Unique cutters · 6+/4+ bp sites · Connectors |
| Features | Feature Library workbench |
| Primers | Full-screen Primer Design workbench |
| Mutagenize | SOE-PCR site-directed mutagenesis designer (4-source CDS picker) |
| Parts | Parts Bin (per-grammar) |
| Constructor | Assembly Constructor for TU building |
| BLAST | BLAST / HMMscan modal |
Data files
All user data persists as human-readable JSON in the user data
directory; every save is atomic and writes a .bak first.
| File | Purpose |
|---|---|
collections.json |
Named collections of plasmids — source of truth |
plasmid_library.json |
Live mirror of the active collection's plasmids |
parts_bin.json |
User-domesticated cloning parts |
primers.json |
Designed primer library |
features.json |
Reusable feature snippets |
feature_colors.json |
Per-type feature color overrides |
codon_tables.json |
Cached codon-usage tables fetched from Kazusa |
cloning_grammars.json |
User-defined cloning grammars (Golden Braid / MoClo / custom) |
settings.json |
App preferences (active collection, active grammar, …) |
crash_recovery/*.gb |
Per-record crash-recovery autosaves |
*.json.bak |
Automatic backup — written before each save |
The schema envelope ({"_schema_version": 1, "entries": [...]})
silently accepts the legacy bare-list format (pre-0.3.1) and rewrites
it on the next save. Newer-version files load with a warning rather
than crashing.
Tests
python3 -m pytest -n auto -q # full suite (~3 min on 8 cores)
python3 -m pytest tests/test_dna_sanity.py # biology correctness only (< 2 s)
python3 -m pytest tests/test_invariants_hypothesis.py # property-based fuzzing
All tests run offline against synthetic SeqRecords and monkeypatched
data paths; an autouse fixture in tests/conftest.py guarantees no
test can write to real user files.
Codebase tour
SpliceCraft is a single-file Python app (splicecraft.py,
~23,000 lines) on Textual + Biopython. The single-file layout is
intentional — no import puzzles, everything is greppable from one
place.
grep -n "^class \|^def " splicecraft.py gives an authoritative live
map. Test files are 1:1 named after the subsystem they cover.
CLAUDE.md at the repo root is the agent + contributor handover
document: ten sacred invariants, error-handling convention, known
pitfalls. Read it before touching the rendering layer, record pipeline,
or primer design.
Maintenance
SpliceCraft is actively maintained. The maintainer is a practicing bioengineer running real cloning workflows in it daily; releases typically go out the same week a problem surfaces at the bench. Issues and PRs welcome at github.com/Binomica-Labs/SpliceCraft/issues.
License
MIT
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file splicecraft-0.5.8.1.tar.gz.
File metadata
- Download URL: splicecraft-0.5.8.1.tar.gz
- Upload date:
- Size: 1.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9a4c81e8d21bd599a6841027a214be3335564543c733abeff48c42c13383c39e
|
|
| MD5 |
cda0db6fa7295deb2e3e518f43e93039
|
|
| BLAKE2b-256 |
46524f19a6d168922ae806badd2fbda27c701ed877db0fcf72dbd6686016a5e3
|
Provenance
The following attestation bundles were made for splicecraft-0.5.8.1.tar.gz:
Publisher:
publish.yml on Binomica-Labs/SpliceCraft
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
splicecraft-0.5.8.1.tar.gz -
Subject digest:
9a4c81e8d21bd599a6841027a214be3335564543c733abeff48c42c13383c39e - Sigstore transparency entry: 1438204975
- Sigstore integration time:
-
Permalink:
Binomica-Labs/SpliceCraft@b9b0bf1c5ff1695b47d2fa5bca5fd74a8397981e -
Branch / Tag:
refs/tags/v0.5.8.1 - Owner: https://github.com/Binomica-Labs
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@b9b0bf1c5ff1695b47d2fa5bca5fd74a8397981e -
Trigger Event:
push
-
Statement type:
File details
Details for the file splicecraft-0.5.8.1-py3-none-any.whl.
File metadata
- Download URL: splicecraft-0.5.8.1-py3-none-any.whl
- Upload date:
- Size: 333.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
337ff286b77b1b8208870357d751a6a6cb72cd4e7021c869104009807b29cd5d
|
|
| MD5 |
eedd8d72f35c99e03f7cdadf15cd25e0
|
|
| BLAKE2b-256 |
ab17a69980b69cc93ad196a90f18be4bef65f63d90ad47e6c0da49d146505b97
|
Provenance
The following attestation bundles were made for splicecraft-0.5.8.1-py3-none-any.whl:
Publisher:
publish.yml on Binomica-Labs/SpliceCraft
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
splicecraft-0.5.8.1-py3-none-any.whl -
Subject digest:
337ff286b77b1b8208870357d751a6a6cb72cd4e7021c869104009807b29cd5d - Sigstore transparency entry: 1438205012
- Sigstore integration time:
-
Permalink:
Binomica-Labs/SpliceCraft@b9b0bf1c5ff1695b47d2fa5bca5fd74a8397981e -
Branch / Tag:
refs/tags/v0.5.8.1 - Owner: https://github.com/Binomica-Labs
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@b9b0bf1c5ff1695b47d2fa5bca5fd74a8397981e -
Trigger Event:
push
-
Statement type: