A Python package implementing the synthetic nearest neighbors estimator for panel data causal inference.
Project description
synthnn
A Python package for panel data causal inference implementing synthetic nearest neighbors (SNN), a causal model for matrix completion that imputes treated units’ counterfactual outcomes from weighted nearest neighbors in a low-rank subspace learned from pre-treatment data..
Features
- Flexible Panel Data Support — Works with both simultaneous and staggered treatment adoption.
- Multiple Inference Methods — Jackknife, bootstrap, and Fisher-style placebo tests for uncertainty quantification.
- Built-in Visualization — Gap plots and observed vs. counterfactual comparisons.
- Customizable Imputation — Fully configurable parameters to match your data’s characteristics.
Installation
pip install synthnn
Quick Start
import pandas as pd
from synthnn import SNN
# Load your panel data
df = pd.read_csv("your_panel_data.csv")
# Initialize and fit the SNN model
model = SNN(
unit_col="Unit",
time_col="Time",
outcome_col="Y",
treat_col="W",
variance_type="bootstrap",
resamples=500,
alpha=0.05
)
model.fit(df)
model.summary()
# Visualize results
model.plot("gap") # Average treatment effect on the treated (ATT) over time
model.plot("counterfactual") # Observed vs. counterfactual
Full Example — Replicating Abadie et al. (2010)
This example reproduces the well-known California tobacco control study.
Data: prop99.csv in the demos folder.
import pandas as pd
from synthnn import SNN
# 1. Load the data from Abadie et al. (2010)
df0 = pd.read_csv("prop99.csv", low_memory=False)
df = (
df0
.query("TopicDesc == 'The Tax Burden on Tobacco' "
"and SubMeasureDesc == 'Cigarette Consumption (Pack Sales Per Capita)'")
.loc[:, ["LocationDesc", "Year", "Data_Value"]]
.rename(columns={
"LocationDesc": "Unit",
"Year": "Time",
"Data_Value": "Y"
})
)
# Drop territories & aggregate rows (keep 50 states)
bad_units = ["District of Columbia", "United States", "Guam",
"Puerto Rico", "American Samoa", "Virgin Islands"]
df = df[~df["Unit"].isin(bad_units)]
# 2. Define the treatment indicator
df["W"] = ((df["Unit"] == "California") & (df["Time"] >= 1989)).astype(int)
# 3. Fit Synthetic-Nearest-Neighbors
model = SNN(
unit_col="Unit",
time_col="Time",
outcome_col="Y",
treat_col="W",
variance_type="bootstrap",
resamples=100,
alpha=0.05
)
model.fit(df)
# 4. Inspect results
model.summary()
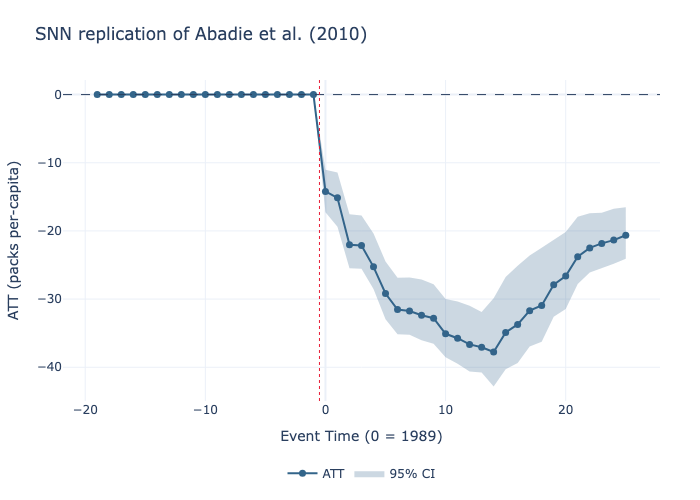
# 5. Plot the gap between treated and counterfactual
model.plot(
title="SNN replication of Abadie et al. (2010)",
xlabel="Event Time (0 = 1989)",
ylabel="ATT (packs per-capita)"
).write_image("gap.png")
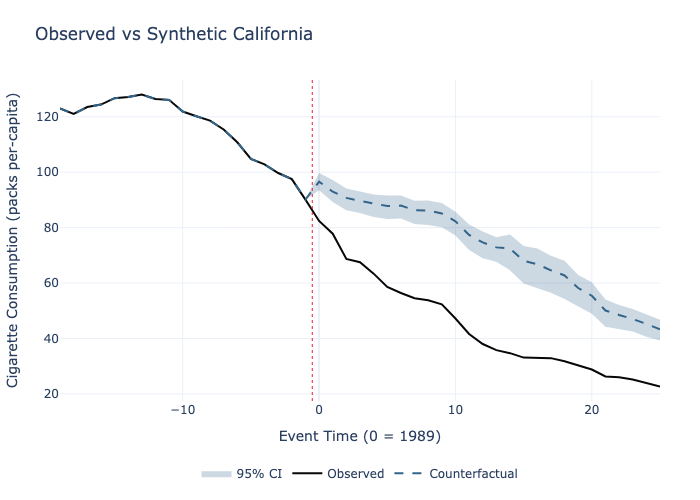
# 6. Plot observed vs counterfactual paths
model.plot(
plot_type="counterfactual",
title="Observed vs Synthetic California",
xlabel="Event Time (0 = 1989)",
ylabel="Cigarette Consumption (packs per-capita)"
).write_image("counterfactual.png")
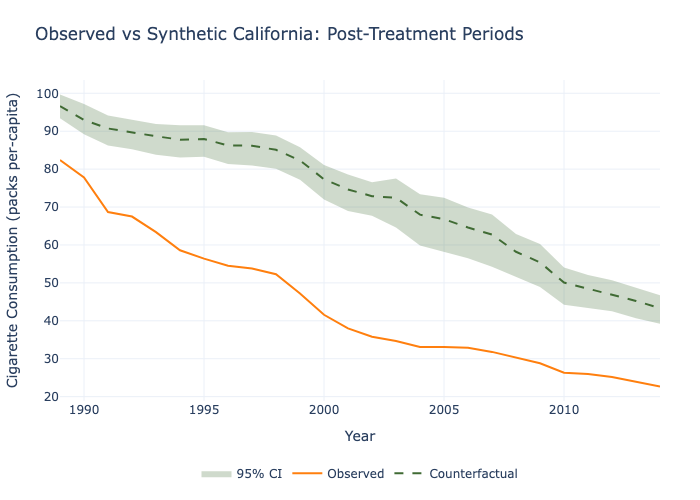
# 7. Same as before but with calendar time on the x-axis, only post-treatment periods, and custom colors
model.plot(
plot_type="counterfactual",
calendar_time=True,
xrange=(1989, 2014),
title="Observed vs Synthetic California: Post-Treatment Periods",
xlabel="Year",
ylabel="Cigarette Consumption (packs per-capita)",
counterfactual_color="#406B34", # green
observed_color="#ff7f0e" # orange
).write_image("graphics.png")
# 8. Inference using the placebo test (only works if there is exactly one treated unit)
model_pc = SNN(unit_col="Unit", time_col="Time", outcome_col="Y", treat_col="W",
variance_type="placebo", alpha=0.05)
model_pc.fit(df)
model_pc.summary()
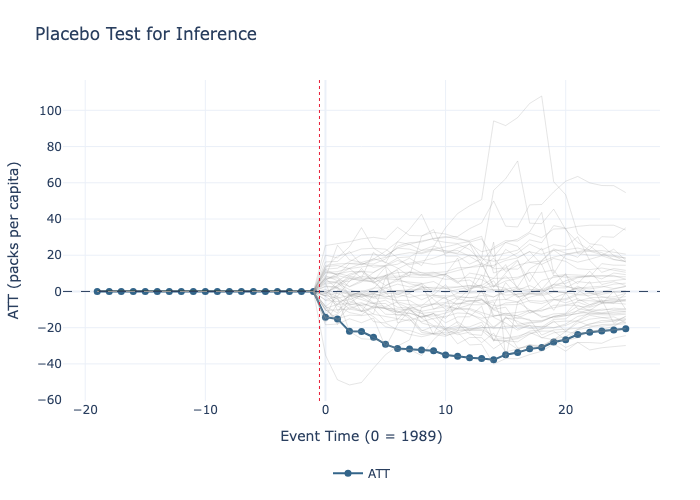
# 9. Plot the results, displaying the paths of the placebo treated units against the actual treated unit
model_pc.plot(show_placebos=True,
title="Placebo Test for Inference",
xlabel="Event Time (0 = 1989)",
ylabel="ATT (packs per capita)").write_image("placebo.png")
Output
Click to expand
============================================================
SNN Estimation Results
============================================================
--- Overall ATT ---
estimate method se p_value ci_lower ci_upper
-28.25 bootstrap 2.032 0 -32.07 -24.03
--- ATT by Event Time (Post-Treatment) ---
event_time att N_units se p_value ci_lower ci_upper method
0 -14.2 1 1.651 0 -17.06 -11.28 bootstrap
1 -15.15 1 2.077 3.015e-13 -18.75 -11.43 bootstrap
2 -22.02 1 2.089 0 -26.16 -18.22 bootstrap
3 -22.12 1 2.184 0 -26.15 -18.05 bootstrap
4 -25.27 1 1.959 0 -28.55 -21.33 bootstrap
5 -29.18 1 2.129 0 -32.97 -25 bootstrap
6 -31.54 1 2.052 0 -35.08 -27.1 bootstrap
7 -31.75 1 2.054 0 -35.6 -27.29 bootstrap
8 -32.37 1 2.207 0 -36.2 -28.41 bootstrap
9 -32.8 1 2.035 0 -36.08 -28.68 bootstrap
10 -35.09 1 2.144 0 -38.64 -31.03 bootstrap
11 -35.74 1 2.196 0 -39.74 -31.06 bootstrap
12 -36.65 1 2.301 0 -41.26 -31.28 bootstrap
13 -37.07 1 2.291 0 -41.5 -31.68 bootstrap
14 -37.75 1 3.217 0 -44.07 -31.11 bootstrap
15 -34.89 1 3.052 0 -40.54 -27.46 bootstrap
16 -33.71 1 3.303 0 -39.55 -26.32 bootstrap
17 -31.7 1 3.097 0 -37.31 -25.12 bootstrap
18 -30.94 1 3.264 0 -36.9 -23.89 bootstrap
19 -27.91 1 2.687 0 -32.99 -22.78 bootstrap
20 -26.63 1 2.583 0 -31.33 -21.51 bootstrap
21 -23.79 1 2.254 0 -27.74 -19.66 bootstrap
22 -22.49 1 2.131 0 -26.36 -18.57 bootstrap
23 -21.83 1 2.042 0 -25.58 -18.39 bootstrap
24 -21.35 1 2.044 0 -24.94 -17.73 bootstrap
25 -20.63 1 1.895 0 -24.19 -17.52 bootstrap
============================================================
============================================================
SNN Estimation Results
============================================================
--- Overall ATT ---
estimate placebo_p placebo_rank
-28.25 0.08 4
Placebo Fisher p-value: 0.08 (rank 4/50)
--- ATT by Event Time (Post-Treatment) ---
event_time att N_units placebo_p
0 -14.2 1 0.2
1 -15.15 1 0.22
2 -22.02 1 0.12
3 -22.12 1 0.12
4 -25.27 1 0.08
5 -29.18 1 0.06
6 -31.54 1 0.06
7 -31.75 1 0.06
8 -32.37 1 0.06
9 -32.8 1 0.04
10 -35.09 1 0.04
11 -35.74 1 0.04
12 -36.65 1 0.04
13 -37.07 1 0.06
14 -37.75 1 0.1
15 -34.89 1 0.12
16 -33.71 1 0.1
17 -31.7 1 0.14
18 -30.94 1 0.14
19 -27.91 1 0.14
20 -26.63 1 0.2
21 -23.79 1 0.2
22 -22.49 1 0.18
23 -21.83 1 0.18
24 -21.35 1 0.16
25 -20.63 1 0.12
============================================================
Plots
Parameters
General
-
unit_col, time_col, outcome_col, treat_col (str) — Column names for unit ID, time, outcome, and treatment indicator.
-
variance_type (str) — Inference method:
"jackknife"— Leave-one-unit-out resampling"bootstrap"(default) — Block bootstrap on units"placebo"— Fisher randomization test (only when exactly one treated unit)
-
resamples (int) — Bootstrap resamples (default: 500)
-
alpha (float) — Significance level for confidence intervals (default: 0.05)
-
snn_params (dict) — Parameters for the
SyntheticNearestNeighborsimputer.
SNN Parameters (snn_params)
- n_neighbors (int) — Number of nearest neighbors (default: 1)
- weights (str) —
'uniform'or'distance' - random_splits (bool) — Use random splits in the algorithm
- max_rank (int) — Maximum rank for low-rank approximation
- spectral_t, linear_span_eps, subspace_eps (float) — Algorithm thresholds (default: 0.1)
- min_value, max_value (float) — Bounds for imputed values
- verbose (bool) — Print progress.
Plot Parameters
- plot_type —
"gap"or"counterfactual" - calendar_time (bool) — Use calendar time (for simultaneous adoption only)
- xrange (tuple) —
(min, max)for x-axis - title, xlabel, ylabel (str) — Labels
- figsize (tuple) —
(width, height) - color, observed_color, counterfactual_color, placebo_color (str) — Plot colors
- placebo_opacity (float) — Opacity for placebo lines (default: 0.25)
Output Attributes
After fitting, the model exposes:
- overall_att_ — Overall ATT with inference statistics
- att_by_event_time_ — ATT series by event time
- att_by_time_ — ATT series by calendar time
- individual_effects_ — Unit-level effects
- counterfactual_event_df_ — Observed vs. counterfactual (event time)
- counterfactual_df_ — Observed vs. counterfactual (calendar time)
Requirements
pandas,numpy,scipy,plotly,scikit-learn
Acknowledgments
The implementation in this package adapts and builds upon the code from the syntheticNN repository by Dennis Shen.
License
This project is licensed under the MIT License - see the LICENSE file for details.
Citation
If you use this package in your research, you can cite it as below.
@software{synthnn,
author = {Lipkovitz, Rivka},
month = jun,
title = {{synthnn: a Python package for estimating treatment effects using Synthetic Nearest Neighbors}},
url = {https://github.com/rivkalipko/synthnn},
year = {2025}
}
Please also consider citing the authors of the original paper:
Agarwal, A., Dahleh, M., Shah, D., & Shen, D. (2023, July). Causal matrix completion. In The thirty sixth annual conference on learning theory (pp. 3821-3826). PMLR.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file synthnn-1.1.5.tar.gz.
File metadata
- Download URL: synthnn-1.1.5.tar.gz
- Upload date:
- Size: 27.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7961610f2a4db26b48732029fc051ab15b6b2d50b50a28a222fd77397cf02883
|
|
| MD5 |
781b80f81e576178ee69ccc53952cc7f
|
|
| BLAKE2b-256 |
d7d9813e199325751c5e83a24b2bfda8bc1250e7583065c70f411a08ab6ddd7a
|
File details
Details for the file synthnn-1.1.5-py3-none-any.whl.
File metadata
- Download URL: synthnn-1.1.5-py3-none-any.whl
- Upload date:
- Size: 23.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4497a9f52d8416392c30bb86d393f11e53a7c95d978acfbd596c6d86c6c2574a
|
|
| MD5 |
8e469d1898454faa623bbeac479b8c2d
|
|
| BLAKE2b-256 |
014f394fd39395453c3df4d1b92507d263ad4d600b638275cece42ac2dfa5bd9
|