An in-memory data analysis format for single-cell profiles alongside their corresponding images and segmentation masks.
Project description
CytoDataFrame
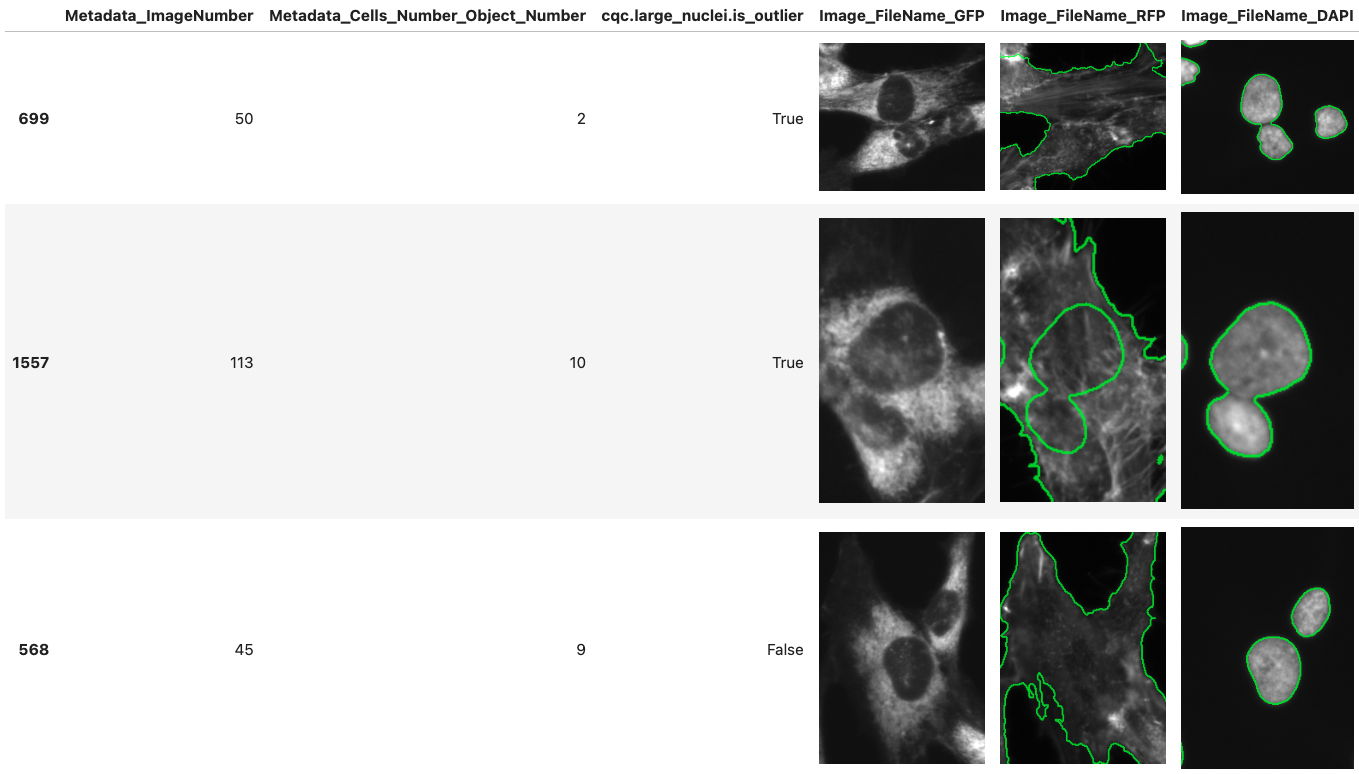
CytoDataFrame is an advanced in-memory data analysis format designed for single-cell profiling, integrating not only the data profiles but also their corresponding microscopy images and segmentation masks. Traditional single-cell profiling often excludes the associated images from analysis, limiting the scope of research. CytoDataFrame bridges this gap, offering a purpose-built solution for comprehensive analysis that incorporates both the data and images, empowering more detailed and visual insights in single-cell research.
CytoDataFrame development began within coSMicQC - a single-cell profile quality control package.
Installation
Install CytoDataFrame from source using the following:
# install from pypi
pip install cytodataframe
# or install directly from source
pip install git+https://github.com/WayScience/CytoDataFrame.git
Contributing, Development, and Testing
Please see our contributing documentation for more details on contributions, development, and testing.
References
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cytodataframe-0.0.22.tar.gz.
File metadata
- Download URL: cytodataframe-0.0.22.tar.gz
- Upload date:
- Size: 16.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e14c36d799cde03c9b98d490f6f00f3b432ba08c1241a4f6dd22caadd0767af1
|
|
| MD5 |
1557e51722e02882308caaa56a800327
|
|
| BLAKE2b-256 |
40b24b0cef12ab40d19c68409235325a9ca8cc5e5096ea6b4beab780ae3b5054
|
Provenance
The following attestation bundles were made for cytodataframe-0.0.22.tar.gz:
Publisher:
publish-pypi.yml on WayScience/CytoDataFrame
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
cytodataframe-0.0.22.tar.gz -
Subject digest:
e14c36d799cde03c9b98d490f6f00f3b432ba08c1241a4f6dd22caadd0767af1 - Sigstore transparency entry: 211532888
- Sigstore integration time:
-
Permalink:
WayScience/CytoDataFrame@922c24486164eee39dee4218c6be53a2ea4aded4 -
Branch / Tag:
refs/tags/v0.0.22 - Owner: https://github.com/WayScience
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-pypi.yml@922c24486164eee39dee4218c6be53a2ea4aded4 -
Trigger Event:
release
-
Statement type:
File details
Details for the file cytodataframe-0.0.22-py3-none-any.whl.
File metadata
- Download URL: cytodataframe-0.0.22-py3-none-any.whl
- Upload date:
- Size: 16.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3b119ea74cf6e281eb4bb36e2230f20263eca0e2f42199a131486d76305e2cdc
|
|
| MD5 |
45384869659b288f1bed42f9a96592ae
|
|
| BLAKE2b-256 |
d19d8faf99de34d1253e058fc09f2249f02431f768431b7682084f458b6b470a
|
Provenance
The following attestation bundles were made for cytodataframe-0.0.22-py3-none-any.whl:
Publisher:
publish-pypi.yml on WayScience/CytoDataFrame
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
cytodataframe-0.0.22-py3-none-any.whl -
Subject digest:
3b119ea74cf6e281eb4bb36e2230f20263eca0e2f42199a131486d76305e2cdc - Sigstore transparency entry: 211532890
- Sigstore integration time:
-
Permalink:
WayScience/CytoDataFrame@922c24486164eee39dee4218c6be53a2ea4aded4 -
Branch / Tag:
refs/tags/v0.0.22 - Owner: https://github.com/WayScience
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-pypi.yml@922c24486164eee39dee4218c6be53a2ea4aded4 -
Trigger Event:
release
-
Statement type:


















