Neuron Analysis and Visualization library
Project description

NAVis is a Python 3 library for Neuron Analysis and Visualization.
Documentation
Visit our documentation here!
Features
- polyglot: work with and convert between neuron skeletons, meshes, dotprops and images
- visualize: 2D (matplotlib) and 3D (octarine, vispy, plotly or k3d) plots
- process: skeletonization, meshing, smoothing, repair, downsampling, etc.
- morphometrics: Strahler analysis, cable length, volume, tortuosity and more
- similarity: compare & cluster by morphology (e.g. NBLAST, persistence or form factor) or connectivity metrics
- transform: move data between template brains (built-in support for HDF5, CMTK, Elastix and landmark-based transforms)
- interface: load neurons directly from neuPrint, neuromorpho.org and other remote data repositories
- model neurons and networks using the NEURON simulator
- render: use Blender 3D for high quality visualizations
- R neuron libraries: interfaces with nat, rcatmaid, elmr and more
- import-export: read/write SWCs, neuroglancer's "precomputed" format, NMX/NML, NRRD, mesh-files and more
- fast: uses functions compiled in Rust under-the-hood (see fastcore)
- scalable: out-of-the-box support for multiprocessing
- extensible: build your own package on top of navis - see pymaid for example
Getting started
See the documentation for detailed installation instructions, tutorials and examples. For the impatient:
pip3 install "navis[all]"
which includes all optional extras providing features and/or performance improvements. If you encounter issues, you may want to try a minimal installation and install optional dependencies as needed. Please see the Install Instructions for a detailed explanation.
Questions?
Questions on how to use navis are best placed in discussions. Same goes for cool projects or analyses you made using navis -
we'd love to hear from you!
Changelog
A summary of changes can be found here.
NAVis & friends
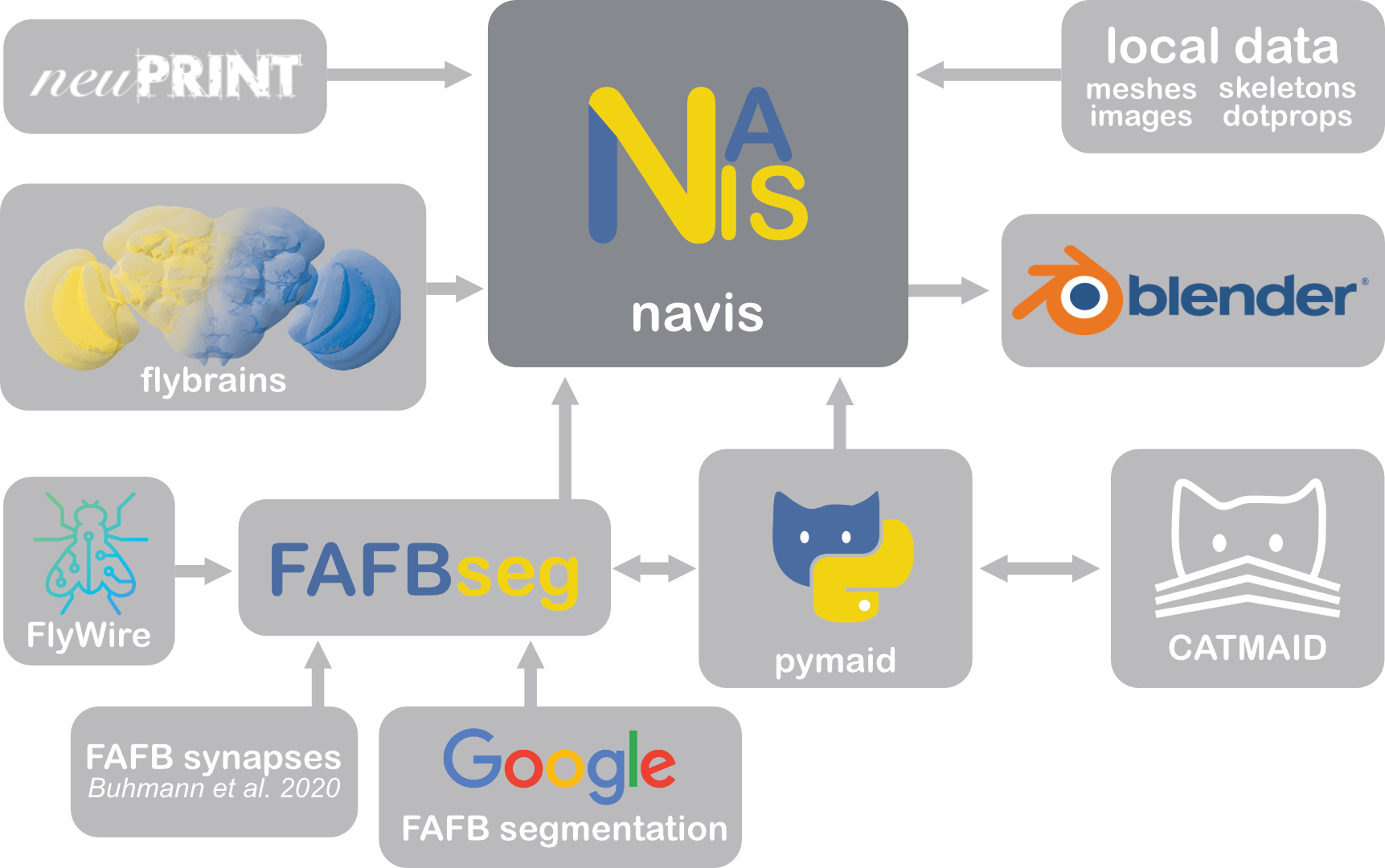
NAVis comes with batteries included but is also highly extensible. Some libraries built on top of NAVis:
- flybrains provides templates and transforms for Drosophila brains to use with navis
- pymaid pulls and pushes data from/to CATMAID servers
- fafbseg contains tools to work with auto-segmented data for the FAFB EM dataset including FlyWire
Who uses NAVis?
NAVis has been used in a range of neurobiological publications. See here for a list.
We have implemented various published algorithms and methods:
- NBLAST: Comparison of neurons based on morphology (Costa et al., 2016)
- Vertex Similarity: Comparison of neurons based on connectivity (Jarrell et al., 2012)
- Comparison of neurons based on synapse distribution (Schlegel et al., 2016)
- Synapse flow centrality for axon-dendrite splits (Schneider-Mizell et al., 2016)
Working on your own cool new method? Consider adding it to NAVis!
Citing NAVis
We'd love to know if you found NAVis useful for your research! You can help us
spread the word by citing the DOI provided by Zenodo
License
This code is under GNU GPL V3.
Acknowledgments
NAVis is inspired by and inherits much of its design from the excellent natverse R packages by Greg Jefferis, Alex Bates, James Manton and others.
Contributing
Want to contribute? Great, here is how!
Report bugs or request features
Open an issue. For bug reports please make sure to include some code/data with a minimum example for us to reproduce the bug.
Contribute code
We're always happy for people to contribute code - be it a small bug fix, a new feature or improved documentation.
Here's how you'd do it in a nutshell:
- Fork this repository
git cloneit to your local machine- Install the full development dependencies with
pip install -r requirements.txt - Install the package in editable mode with
pip install -e ".[all]" - Create,
git add,git commit,git push, and pull request your changes.
Run the tests locally with pytest -v.
Docstrings should use the numpydoc format,
and make sure you include any relevant links and citations.
Unit tests should be doctests
and/or use pytest in the ./tests directory.
Doctests have access to the tmp_dir: pathlib.Path variable,
which should be used if any files need to be written.
Feel free to get in touch either through an issue or discussion if you need pointers or input on how to implement an idea.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters


















