Multi-class confusion matrix library in Python
Project description
Table of contents
- Overview
- Installation
- Usage
- Document
- Issues & Bug Reports
- Todo
- Outputs
- Dependencies
- Contribution
- References
- Cite
- Authors
- License
- Donate
- Changelog
Overview
PyCM is a multi-class confusion matrix library written in Python that supports both input data vectors and direct matrix, and a proper tool for post-classification model evaluation that supports most classes and overall statistics parameters. PyCM is the swiss-army knife of confusion matrices, targeted mainly at data scientists that need a broad array of metrics for predictive models and an accurate evaluation of large variety of classifiers.
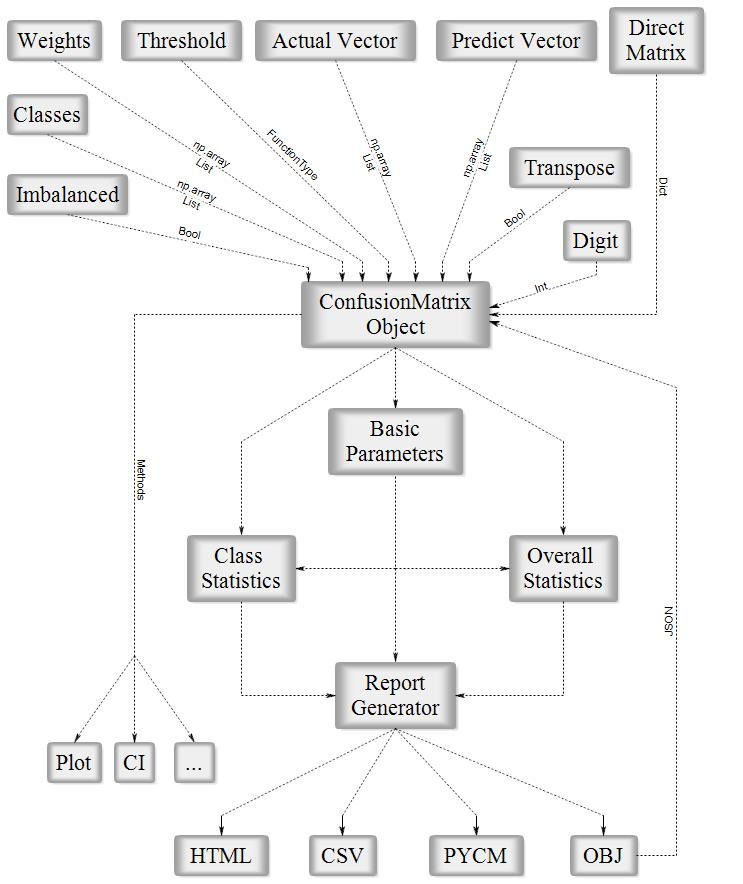
Fig1. PyCM Block Diagram
| Open Hub |  |
| PyPI Counter |  |
| Github Stars |  |
Installation
Source Code
- Download Version 1.0 or Latest Source
- Run
pip install -r requirements.txtorpip3 install -r requirements.txt(Need root access) - Run
python3 setup.py installorpython setup.py install(Need root access)
PyPI
- Check Python Packaging User Guide
- Run
pip install pycm --upgradeorpip3 install pycm --upgrade(Need root access)
Easy Install
- Run
easy_install --upgrade pycm(Need root access)
Usage
From Vector
>>> from pycm import *
>>> y_actu = [2, 0, 2, 2, 0, 1, 1, 2, 2, 0, 1, 2] # or y_actu = numpy.array([2, 0, 2, 2, 0, 1, 1, 2, 2, 0, 1, 2])
>>> y_pred = [0, 0, 2, 1, 0, 2, 1, 0, 2, 0, 2, 2] # or y_pred = numpy.array([0, 0, 2, 1, 0, 2, 1, 0, 2, 0, 2, 2])
>>> cm = ConfusionMatrix(actual_vector=y_actu, predict_vector=y_pred) # Create CM From Data
>>> cm.classes
[0, 1, 2]
>>> cm.table
{0: {0: 3, 1: 0, 2: 0}, 1: {0: 0, 1: 1, 2: 2}, 2: {0: 2, 1: 1, 2: 3}}
>>> print(cm)
Predict 0 1 2
Actual
0 3 0 0
1 0 1 2
2 2 1 3
Overall Statistics :
95% CI (0.30439,0.86228)
Bennett_S 0.375
Chi-Squared 6.6
Chi-Squared DF 4
Conditional Entropy 0.95915
Cramer_V 0.5244
Cross Entropy 1.59352
Gwet_AC1 0.38931
Hamming Loss 0.41667
Joint Entropy 2.45915
KL Divergence 0.09352
Kappa 0.35484
Kappa 95% CI (-0.07708,0.78675)
Kappa No Prevalence 0.16667
Kappa Standard Error 0.22036
Kappa Unbiased 0.34426
Lambda A 0.16667
Lambda B 0.42857
Mutual Information 0.52421
Overall_ACC 0.58333
Overall_J (1.225,0.40833)
Overall_RACC 0.35417
Overall_RACCU 0.36458
PPV_Macro 0.56667
PPV_Micro 0.58333
Phi-Squared 0.55
Reference Entropy 1.5
Response Entropy 1.48336
Scott_PI 0.34426
Standard Error 0.14232
Strength_Of_Agreement(Altman) Fair
Strength_Of_Agreement(Cicchetti) Poor
Strength_Of_Agreement(Fleiss) Poor
Strength_Of_Agreement(Landis and Koch) Fair
TPR_Macro 0.61111
TPR_Micro 0.58333
Class Statistics :
Classes 0 1 2
ACC(Accuracy) 0.83333 0.75 0.58333
BM(Informedness or bookmaker informedness) 0.77778 0.22222 0.16667
DOR(Diagnostic odds ratio) None 4.0 2.0
ERR(Error rate) 0.16667 0.25 0.41667
F0.5(F0.5 score) 0.65217 0.45455 0.57692
F1(F1 score - harmonic mean of precision and sensitivity) 0.75 0.4 0.54545
F2(F2 score) 0.88235 0.35714 0.51724
FDR(False discovery rate) 0.4 0.5 0.4
FN(False negative/miss/type 2 error) 0 2 3
FNR(Miss rate or false negative rate) 0.0 0.66667 0.5
FOR(False omission rate) 0.0 0.2 0.42857
FP(False positive/type 1 error/false alarm) 2 1 2
FPR(Fall-out or false positive rate) 0.22222 0.11111 0.33333
G(G-measure geometric mean of precision and sensitivity) 0.7746 0.40825 0.54772
J(Jaccard index) 0.6 0.25 0.375
LR+(Positive likelihood ratio) 4.5 3.0 1.5
LR-(Negative likelihood ratio) 0.0 0.75 0.75
MCC(Matthews correlation coefficient) 0.68313 0.2582 0.16903
MK(Markedness) 0.6 0.3 0.17143
N(Condition negative) 9 9 6
NPV(Negative predictive value) 1.0 0.8 0.57143
P(Condition positive) 3 3 6
POP(Population) 12 12 12
PPV(Precision or positive predictive value) 0.6 0.5 0.6
PRE(Prevalence) 0.25 0.25 0.5
RACC(Random accuracy) 0.10417 0.04167 0.20833
RACCU(Random accuracy unbiased) 0.11111 0.0434 0.21007
TN(True negative/correct rejection) 7 8 4
TNR(Specificity or true negative rate) 0.77778 0.88889 0.66667
TON(Test outcome negative) 7 10 7
TOP(Test outcome positive) 5 2 5
TP(True positive/hit) 3 1 3
TPR(Sensitivity, recall, hit rate, or true positive rate) 1.0 0.33333 0.5
>>> cm.matrix()
Predict 0 1 2
Actual
0 3 0 0
1 0 1 2
2 2 1 3
>>> cm.normalized_matrix()
Predict 0 1 2
Actual
0 1.0 0.0 0.0
1 0.0 0.33333 0.66667
2 0.33333 0.16667 0.5
Direct CM
>>> from pycm import *
>>> cm2 = ConfusionMatrix(matrix={"Class1": {"Class1": 1, "Class2":2}, "Class2": {"Class1": 0, "Class2": 5}}) # Create CM Directly
>>> cm2
pycm.ConfusionMatrix(classes: ['Class1', 'Class2'])
>>> print(cm2)
Predict Class1 Class2
Actual
Class1 1 2
Class2 0 5
Overall Statistics :
95% CI (0.44994,1.05006)
Bennett_S 0.5
Chi-Squared None
Chi-Squared DF 1
Conditional Entropy None
Cramer_V None
Cross Entropy 1.2454
Gwet_AC1 0.6
Hamming Loss 0.25
Joint Entropy None
KL Divergence 0.29097
Kappa 0.38462
Kappa 95% CI (-0.354,1.12323)
Kappa No Prevalence 0.5
Kappa Standard Error 0.37684
Kappa Unbiased 0.33333
Lambda A None
Lambda B None
Mutual Information None
Overall_ACC 0.75
Overall_J (1.04762,0.52381)
Overall_RACC 0.59375
Overall_RACCU 0.625
PPV_Macro 0.85714
PPV_Micro 0.75
Phi-Squared None
Reference Entropy 0.95443
Response Entropy 0.54356
Scott_PI 0.33333
Standard Error 0.15309
Strength_Of_Agreement(Altman) Fair
Strength_Of_Agreement(Cicchetti) Poor
Strength_Of_Agreement(Fleiss) Poor
Strength_Of_Agreement(Landis and Koch) Fair
TPR_Macro 0.66667
TPR_Micro 0.75
Class Statistics :
Classes Class1 Class2
ACC(Accuracy) 0.75 0.75
BM(Informedness or bookmaker informedness) 0.33333 0.33333
DOR(Diagnostic odds ratio) None None
ERR(Error rate) 0.25 0.25
F0.5(F0.5 score) 0.71429 0.75758
F1(F1 score - harmonic mean of precision and sensitivity) 0.5 0.83333
F2(F2 score) 0.38462 0.92593
FDR(False discovery rate) 0.0 0.28571
FN(False negative/miss/type 2 error) 2 0
FNR(Miss rate or false negative rate) 0.66667 0.0
FOR(False omission rate) 0.28571 0.0
FP(False positive/type 1 error/false alarm) 0 2
FPR(Fall-out or false positive rate) 0.0 0.66667
G(G-measure geometric mean of precision and sensitivity) 0.57735 0.84515
J(Jaccard index) 0.33333 0.71429
LR+(Positive likelihood ratio) None 1.5
LR-(Negative likelihood ratio) 0.66667 0.0
MCC(Matthews correlation coefficient) 0.48795 0.48795
MK(Markedness) 0.71429 0.71429
N(Condition negative) 5 3
NPV(Negative predictive value) 0.71429 1.0
P(Condition positive) 3 5
POP(Population) 8 8
PPV(Precision or positive predictive value) 1.0 0.71429
PRE(Prevalence) 0.375 0.625
RACC(Random accuracy) 0.04688 0.54688
RACCU(Random accuracy unbiased) 0.0625 0.5625
TN(True negative/correct rejection) 5 1
TNR(Specificity or true negative rate) 1.0 0.33333
TON(Test outcome negative) 7 1
TOP(Test outcome positive) 1 7
TP(True positive/hit) 1 5
TPR(Sensitivity, recall, hit rate, or true positive rate) 0.33333 1.0
Activation Threshold
threshold is added in Version 0.9 for real value prediction.
For more information visit Example3
Load From File
file is added in Version 0.9.5 in order to load saved confusion matrix with .obj format generated by save_obj method.
For more information visit Example4
Acceptable Data Types
actual_vector: pythonlistor numpyarrayof any stringable objectspredict_vector: pythonlistor numpyarrayof any stringable objectsmatrix:dictdigit:intthreshold:FunctionType (function or lambda)file:File object
- run
help(ConfusionMatrix)forConfusionMatrixobject details
For more information visit here
Issues & Bug Reports
Just fill an issue and describe it. We'll check it ASAP! or send an email to shaghighi@ce.sharif.edu.
Todo
- Basic
- TP
- FP
- FN
- TN
- Population
- Condition positive
- Condition negative
- Test outcome positive
- Test outcome negative
- Class Statistics
- ACC
- ERR
- BM
- DOR
- F1-Score
- FDR
- FNR
- FOR
- FPR
- LR+
- LR-
- MCC
- MK
- NPV
- PPV
- TNR
- TPR
- Prevalence
- G-measure
- RACC
- Outputs
- CSV File
- HTML File
- Output File
- Table
- Normalized Table
- Overall Statistics
- CI
- Chi-Squared
- Phi-Squared
- Cramer's V
- Kappa
- Kappa Unbiased
- Kappa No Prevalence
- Aickin's alpha
- Bennett S score
- Gwet's AC1
- Scott's pi
- Krippendorff's alpha
- Goodman and Kruskal's lambda A
- Goodman and Kruskal's lambda B
- Kullback-Liebler divergence
- Entropy
- Overall ACC
- Strength of Agreement
- Landis and Koch
- Fleiss
- Altman
- Cicchetti
- TPR Micro/Macro
- PPV Micro/Macro
- Jaccard Index
- Hamming Loss
Outputs
Dependencies
Contribution
Changes and improvements are more than welcome! ❤️ Feel free to fork and open a pull request. Please make your changes in a specific branch and request to pull into dev
Remember to write a few tests for your code before sending pull requests.
References
1- J. R. Landis, G. G. Koch, “The measurement of observer agreement for categorical data. Biometrics,” in International Biometric Society, pp. 159–174, 1977.
2- D. M. W. Powers, “Evaluation: from precision, recall and f-measure to roc, informedness, markedness & correlation,” in Journal of Machine Learning Technologies, pp.37-63, 2011.
3- C. Sammut, G. Webb, “Encyclopedia of Machine Learning” in Springer, 2011.
4- J. L. Fleiss, “Measuring nominal scale agreement among many raters,” in Psychological Bulletin, pp. 378-382.
5- D.G. Altman, “Practical Statistics for Medical Research,” in Chapman and Hall, 1990.
6- K. L. Gwet, “Computing inter-rater reliability and its variance in the presence of high agreement,” in The British Journal of Mathematical and Statistical Psychology, pp. 29–48, 2008.”
7- W. A. Scott, “Reliability of content analysis: The case of nominal scaling,” in Public Opinion Quarterly, pp. 321–325, 1955.
8- E. M. Bennett, R. Alpert, and A. C. Goldstein, “Communication through limited response questioning,” in The Public Opinion Quarterly, pp. 303–308, 1954.
9- D. V. Cicchetti, "Guidelines, criteria, and rules of thumb for evaluating normed and standardized assessment instruments in psychology," in Psychological Assessment, pp. 284–290, 1994.
10- R.B. Davies, "Algorithm AS155: The Distributions of a Linear Combination of χ2 Random Variables," in Journal of the Royal Statistical Society, pp. 323–333, 1980.
11- S. Kullback, R. A. Leibler "On information and sufficiency," in Annals of Mathematical Statistics, pp. 79–86, 1951.
12- L. A. Goodman, W. H. Kruskal, "Measures of Association for Cross Classifications, IV: Simplification of Asymptotic Variances," in Journal of the American Statistical Association, pp. 415–421, 1972.
13- L. A. Goodman, W. H. Kruskal, "Measures of Association for Cross Classifications III: Approximate Sampling Theory," in Journal of the American Statistical Association, pp. 310–364, 1963.
14- T. Byrt, J. Bishop and J. B. Carlin, “Bias, prevalence, and kappa,” in Journal of Clinical Epidemiology pp. 423-429, 1993.
15- M. Shepperd, D. Bowes, and T. Hall, “Researcher Bias: The Use of Machine Learning in Software Defect Prediction,” in IEEE Transactions on Software Engineering, pp. 603-616, 2014.
16- X. Deng, Q. Liu, Y. Deng, and S. Mahadevan, “An improved method to construct basic probability assignment based on the confusion matrix for classification problem, ” in Information Sciences, pp.250-261, 2016.
Cite
If you use PyCM in your research , please cite this JOSS paper :
Haghighi, S., Jasemi, M., Hessabi, S. and Zolanvari, A. (2018). PyCM: Multiclass confusion matrix library in Python. Journal of Open Source Software, 3(25), p.729.
@article{Haghighi2018,
doi = {10.21105/joss.00729},
url = {https://doi.org/10.21105/joss.00729},
year = {2018},
month = {may},
publisher = {The Open Journal},
volume = {3},
number = {25},
pages = {729},
author = {Sepand Haghighi and Masoomeh Jasemi and Shaahin Hessabi and Alireza Zolanvari},
title = {{PyCM}: Multiclass confusion matrix library in Python},
journal = {Journal of Open Source Software}
}
Download PyCM.bib
| JOSS |  |
| Zenodo |  |
| Researchgate |  |
License
Donate to our project
Bitcoin :
12Xm1qL4MXYWiY9sRMoa3VpfTfw6su3vNq
Payping (For Iranian citizens) :
Changelog
All notable changes to this project will be documented in this file.
The format is based on Keep a Changelog and this project adheres to Semantic Versioning.
Unreleased
1.0 - 2018-08-30
Added
- Hamming loss
Changed
README.mdmodified
0.9.5 - 2018-07-08
Added
- Obj load
- Obj save
- Example-4
Changed
README.mdmodified- Block diagram updated
0.9 - 2018-06-28
Added
- Activation Threshold
- Example-3
- Jaccard index
- Overall Jaccard index
Changed
README.mdmodifiedsetup.pymodified
0.8.6 - 2018-05-31
Added
- Example section in document
- Python 2.7 CI
- JOSS paper pdf
Changed
- Cite section
- ConfusionMatrix docstring
- round function changed to numpy.around
README.mdmodified
0.8.5 - 2018-05-21
Added
- Example-1 (Comparison of three different classifiers)
- Example-2 (How to plot via matplotlib)
- JOSS paper
- ConfusionMatrix docstring
Changed
- Table size in HTML report
- Test system
README.mdmodified
0.8.1 - 2018-03-22
Added
- Goodman and Kruskal's lambda B
- Goodman and Kruskal's lambda A
- Cross Entropy
- Conditional Entropy
- Joint Entropy
- Reference Entropy
- Response Entropy
- Kullback-Liebler divergence
- Direct ConfusionMatrix
- Kappa Unbiased
- Kappa No Prevalence
- Random Accuracy Unbiased
pycmVectorErrorclasspycmMatrixErrorclass- Mutual Information
- Support
numpyarrays
Changed
- Notebook file updated
Removed
pycmErrorclass
0.7 - 2018-02-26
Added
- Cramer's V
- 95% Confidence interval
- Chi-Squared
- Phi-Squared
- Chi-Squared DF
- Standard error
- Kappa standard error
- Kappa 95% confidence interval
- Cicchetti benchmark
Changed
- Overall statistics color in HTML report
- Parameters description link in HTML report
0.6 - 2018-02-21
Added
- CSV report
- Changelog
- Output files
digitparameter toConfusionMatrixobject
Changed
- Confusion matrix color in HTML report
- Parameters description link in HTML report
- Capitalize descriptions
0.5 - 2018-02-17
Added
- Scott's pi
- Gwet's AC1
- Bennett S score
- HTML report
0.4 - 2018-02-05
Added
- TPR Micro/Macro
- PPV Micro/Macro
- RACC overall
- ERR(Error rate)
- FBeta-Score
- F0.5
- F2
- Fleiss benchmark
- Altman benchmark
- Output file(.pycm)
Changed
- Class with zero item
- Normalized matrix
Removed
- Kappa and SOA for each class
0.3 - 2018-01-27
Added
- Kappa
- Random accuracy
- Landis and Koch benchmark
overall_stat
0.2 - 2018-01-24
Added
- Population
- Condition positive
- Condition negative
- Test outcome positive
- Test outcome negative
- Prevalence
- G-measure
- Matrix method
- Normalized matrix method
- Params method
Changed
statistic_resulttoclass_statparamstostat
0.1 - 2018-01-22
Added
- ACC
- BM
- DOR
- F1-Score
- FDR
- FNR
- FOR
- FPR
- LR+
- LR-
- MCC
- MK
- NPV
- PPV
- TNR
- TPR
- documents and
README.md
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pycm-1.0.tar.gz.
File metadata
- Download URL: pycm-1.0.tar.gz
- Upload date:
- Size: 474.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.11.0 pkginfo/1.4.2 requests/2.19.1 setuptools/40.2.0 requests-toolbelt/0.8.0 tqdm/4.23.3 CPython/3.4.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d8b566f079a501f766a89a698e525e0eef3af3621a30b10d349569387ba4de3f
|
|
| MD5 |
24eb891ba9c97ad58148614cf3635cfe
|
|
| BLAKE2b-256 |
1050907b75e6af3285f295a6cd77931ad538b5833e68d53b1f2b2b88cb65e412
|
File details
Details for the file pycm-1.0-py2.py3-none-any.whl.
File metadata
- Download URL: pycm-1.0-py2.py3-none-any.whl
- Upload date:
- Size: 28.1 kB
- Tags: Python 2, Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.11.0 pkginfo/1.4.2 requests/2.19.1 setuptools/40.2.0 requests-toolbelt/0.8.0 tqdm/4.23.3 CPython/3.4.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9f43bfae22b9ba2697bc88883374bd3e90b77af7a9e6024408921a38373e9327
|
|
| MD5 |
162263febe873cafff7701cc4936fc11
|
|
| BLAKE2b-256 |
6736880fc24bb7286bc7074ffbe2f013c602a63b121d9a5371847100187c73f6
|